* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Simulating Populations of Neurons - Leeds VLE
Cognitive neuroscience wikipedia , lookup
Endocannabinoid system wikipedia , lookup
Membrane potential wikipedia , lookup
Action potential wikipedia , lookup
Binding problem wikipedia , lookup
Biochemistry of Alzheimer's disease wikipedia , lookup
Neuroeconomics wikipedia , lookup
Recurrent neural network wikipedia , lookup
Apical dendrite wikipedia , lookup
Resting potential wikipedia , lookup
Activity-dependent plasticity wikipedia , lookup
Synaptogenesis wikipedia , lookup
Axon guidance wikipedia , lookup
End-plate potential wikipedia , lookup
Theta model wikipedia , lookup
Multielectrode array wikipedia , lookup
Neural modeling fields wikipedia , lookup
Holonomic brain theory wikipedia , lookup
Neurotransmitter wikipedia , lookup
Development of the nervous system wikipedia , lookup
Artificial general intelligence wikipedia , lookup
Electrophysiology wikipedia , lookup
Nonsynaptic plasticity wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Caridoid escape reaction wikipedia , lookup
Neural oscillation wikipedia , lookup
Molecular neuroscience wikipedia , lookup
Convolutional neural network wikipedia , lookup
Metastability in the brain wikipedia , lookup
Types of artificial neural networks wikipedia , lookup
Mirror neuron wikipedia , lookup
Central pattern generator wikipedia , lookup
Chemical synapse wikipedia , lookup
Stimulus (physiology) wikipedia , lookup
Single-unit recording wikipedia , lookup
Circumventricular organs wikipedia , lookup
Premovement neuronal activity wikipedia , lookup
Neuroanatomy wikipedia , lookup
Optogenetics wikipedia , lookup
Feature detection (nervous system) wikipedia , lookup
Neural coding wikipedia , lookup
Neuropsychopharmacology wikipedia , lookup
Pre-Bötzinger complex wikipedia , lookup
Channelrhodopsin wikipedia , lookup
Biological neuron model wikipedia , lookup
Simulating Populations of Neurons
George Parish
MSc Artificial Intelligence
2012/2013
"THE CANDIDATE CONFIRMS THAT THE WORK SUBMITTED IS THEIR OWN AND THE
APPROPRIATE CREDIT HAS BEEN GIVEN WHERE REFERENCE HAS BEEN MADE TO THE
WORK OF OTHERS.
I UNDERSTAND THAT FAILURE TO ATTRIBUTE MATERIAL WHICH IS OBTAINED FROM
ANOTHER SOURCE MAY BE CONSIDERED AS PLAGIARISM."
(Signature of student)
Simulating Populations of Neurons
George Parish
2013
SUMMARY
The goal of this project was to understand cognitive neuroscience to be able to simulate
networks of neurons, as well as to use analysis techniques to evaluate the interactions between
populations of neurons. To accomplish this, a lengthy literature review was undertaken to gain an
understanding of neuroscience. This would form the basis for simulations of networks of neurons to
determine: how neurons react to the background noise of surrounding spiking neurons, whether a
layered network can transmit information through precise spike-timing, as well as what initial
conditions are required for this synchronous spiking activity.
The project aimed to understand and implement known networks and techniques, as well as
to evaluate them by creating script to analyse the dynamics of populations of neurons.
ii
Simulating Populations of Neurons
George Parish
2013
ACKNOWLEDGEMENTS
Most of all I would like to thank my supervisor Dr. Marc de Kamps for the frequent meetings
which were key to the completion of this project. I would also like to thank him for his patience in
explaining topics and motivating me to push myself harder.
Thanks also to my assessor Professor Tony Cohn for giving valuable time to give feedback
which helped guide the project.
Finally I would like to thank my parents, who made it possible to me to follow my ambitions
and study for a Masters course in the first place. Thanks also to my new German family for being a
huge positive influence and supporting me through the year.
iii
Simulating Populations of Neurons
George Parish
2013
CONTENTS
Summary ................................................................................................................................................. ii
Acknowledgements................................................................................................................................ iii
List of Figures ........................................................................................................................................ vii
1. Introduction ........................................................................................................................................ 1
1.1 Overview........................................................................................................................................ 1
1.2 Aims and Objectives ...................................................................................................................... 2
1.3 Minimum Requirements and Deliverables.................................................................................... 2
1.4 Relevance to Degree ..................................................................................................................... 2
1.5 Project Management ..................................................................................................................... 3
1.6 Research Methodology ................................................................................................................. 3
1.7 Research Questions ....................................................................................................................... 3
1.8 Project Outline .............................................................................................................................. 4
2. Literature Review ................................................................................................................................ 5
2.1 Machine Learning .......................................................................................................................... 5
2.1.1 Connectionist Models ............................................................................................................. 6
2.1 2 Non-Connectionist Models ..................................................................................................... 7
2.2 Computational Modelling.............................................................................................................. 7
2.2.1 Population Firing Rate ............................................................................................................ 8
2.2.2 Raster Plot............................................................................................................................... 8
2.2.3 Voltage Trace .......................................................................................................................... 8
2.2.4 Population Density.................................................................................................................. 8
2.3 Bio-Inspired Computing................................................................................................................. 9
2.3.1 The Brain as a Whole .............................................................................................................. 9
2.3.2 Structure of a Neuron ........................................................................................................... 12
2.3.3 The Integrate-and-Fire Model .............................................................................................. 17
2.3.4 Modelling Populations of Neurons ....................................................................................... 25
iv
Simulating Populations of Neurons
George Parish
2013
3 Experiments & Evaluation .................................................................................................................. 31
3.1 Implement a Nest Simulation ...................................................................................................... 31
3.2 Simulation of 10,000 Integrate-and-Fire Neurons ...................................................................... 32
3.2.1 Implementation .................................................................................................................... 32
3.2.2 Evaluation ............................................................................................................................. 32
3.3 The Balanced Excitation/Inhibition Model .................................................................................. 35
3.3.1 Implementation .................................................................................................................... 35
3.3.2 Evaluation ............................................................................................................................. 36
3.4 The Stable Propagation of Synchronous Spiking Model ............................................................. 38
3.4.1 Previous Research................................................................................................................. 38
3.4.2 Implementation of Background Noise .................................................................................. 39
3.4.3 Implementation of Network ................................................................................................. 42
3.4.4 Evaluation ............................................................................................................................. 42
3.4.5 Case Study : Dispersion of the Propagating Volley ............................................................... 45
3.4.6 Case Study : Synchronised Initial Volley Causing Synchronisation ....................................... 47
3.4.7 Case Study : De-Synchronised Initial Volley Causing Synchronisation ................................. 49
3.4.8 Case Study : Synchronised Initial Volley Causing Late Synchronisation ............................... 51
4 Conclusion .......................................................................................................................................... 53
4.1 Project Evaluation ....................................................................................................................... 53
4.1.1 Aim and Minimum Requirements ........................................................................................ 53
4.1.2 Exceeding Requirements ...................................................................................................... 54
4.1.3 Research Questions .............................................................................................................. 54
4.2 Challenges ................................................................................................................................... 56
4.3 Future Work ................................................................................................................................ 57
Bibliography ............................................................................................................................................. i
References .............................................................................................................................................. ii
Appendix A ............................................................................................................................................. iv
v
Simulating Populations of Neurons
George Parish
2013
Project Reflection ............................................................................................................................... iv
Appendices B:1 ....................................................................................................................................... v
Initial GANTT chart .............................................................................................................................. v
Appendices B:2 ...................................................................................................................................... vi
Final GANTT chart............................................................................................................................... vi
Appendices C:1...................................................................................................................................... vii
MAIN FUNCTION FOR ANALYSING NEURONAL DATA ....................................................................... vii
Appendices C:2...................................................................................................................................... xii
FUNCTION TO PLOT EVOLUTION OF SYNCHRONOUS SPIKING ......................................................... xii
Appendix C:3 .........................................................................................................................................xiv
NEST CODE FOR SYNCHRONOUS SPIKING MODEL ............................................................................xiv
vi
Simulating Populations of Neurons
George Parish
2013
LIST OF FIGURES
Figure 1 Waterfall Methodology ............................................................................................................. 3
Figure 2 Brodmann areas of the brain (Gazzaniga, 1998) ...................................................................... 9
Figure 3 Types of biological neurons in the nervous system (Gazzaniga, 1998) .................................. 10
Figure 4 Anatomy and Functional areas of the brain (http://catalog.nucleusinc.com) ....................... 11
Figure 5 Structure of a neuron (left) and ions traversing the cell membrane (right) .......................... 12
Figure 6 The Resistance Capacitance (RC) Circuit ................................................................................ 13
Figure 7 Excitatory and Inhibitory connections for a delta model neuron ........................................... 20
Figure 8 DC generator and single delta neuron network ..................................................................... 21
Figure 9 A single integrate-and-fire neuron with a single direct current input of 200pA (500-1000ms),
380pA (1000-1500ms) and 450pA (1500-2000ms) ............................................................................... 21
Figure 10 Spike frequency of model neuron vs value of the direct current input................................ 22
Figure 11 Alpha model analysis............................................................................................................ 24
Figure 12 Serial and Parallel chains ...................................................................................................... 25
Figure 13 Converging and Diverging chains .......................................................................................... 26
Figure 14 Poisson distribution of spike events ..................................................................................... 29
Figure 15 Network for 1 DC and 2 Poisson generators connected to a single delta neuron................ 29
Figure 16 Simulation of delta neuron with poisson and DC generators ............................................... 30
Figure 17 Network of 10,000 delta neurons connected to 800 1Hz poisson generators ..................... 32
Figure 18 Analysis of 10,000 neurons with noise ................................................................................. 33
Figure 19 Time between spikes of 10000 neurons with noise ............................................................. 34
Figure 20 Network of balanced excitation/inhibition model................................................................ 35
Figure 21 Analysis of balanced excitatory/inhibitory network ............................................................ 36
Figure 22 Example of a synchronised and de-synchronised volley ...................................................... 38
Figure 23 Network showing how noise was connected to neurons in each layer ............................... 39
Figure 24 Experimenting with the value to multiply the threshold rate by (p) .................................... 40
Figure 25 Long simulation for p=235 & p=240...................................................................................... 41
vii
Simulating Populations of Neurons
George Parish
2013
Figure 26 Network for the stable propagation of synchronous spiking model .................................... 42
Figure 27 Evolution of initial spike volleys, varying in number of neurons (a) with different
synchronisations (sigma) ...................................................................................................................... 43
Figure 28 Evolution of initial spike volleys (Diesmann, Gewaltig, & Aertsen, 1999) ............................ 44
Figure 29 Case study : Dispersion of fully synchronised initial volley (a=43 =0) ................................ 46
Figure 30 Case study : Synchronised initial volley (a=44 =0) causing synchronisation ...................... 48
Figure 31 Case study : De-synchronised initial volley (a=36 =4) causing synchronisation ................. 50
Figure 32 Case study : Synchronised initial volley (a=43.5 =0) causing late synchronisation ............ 52
viii
1. INTRODUCTION
1.1 OVERVIEW
Understanding the brain is a recent fascination in modern computing. We have come to realise that
the brain is the most advanced computational tool that we know of, to be able to replicate neuronal
processes could vastly improve current computational techniques. However, the more we
understand the more we come to realise the magnitude of this undertaking. As there are billions of
neurons separated by different classes with even more connections of differing types, a top down
approach has to be taken. We can only begin to try to replicate processes once the architecture of
the brain has been structured and different areas have been assigned to different roles. The
advances we have made over the last 100 years have allowed us to now consider processes on an
individual level and use computational techniques to be able to simulate them.
This project considers the paper Stable propagation of synchronous spiking in cortical neural
networks (Diesmann, Gewaltig, & Aertsen, 1999). The paper proposes that information can be
carried by precise spike timing and challenges the view that groups of neurons are incapable of
transmitting signals with millisecond accuracy due to noise from surrounding neurons. They go on to
prove this by creating a computational model that represents biological neurons from the cerebral
cortex, where a signal is propagated through consecutively connected groups of neurons. Successive
groups of neurons fire more and more synchronously together as the signal is propagated, proving
their proposed theory.
This project will re-create this network of neurons by creating a computational model using the
software NEST. By using computational models to analyse the outputted data from this model, it
should be possible to further examine the case in which neurons are kept ‘trigger-happy’ by
surrounding by background activity in the brain, and the effect of incoming signals of varying
strength and synchronisation on the network of neurons.
1
Simulating Populations of Neurons
George Parish
2013
1.2 AIMS AND OBJECTIVES
The overall aim of this project is to clarify the results of the proposed network (Diesmann, Gewaltig,
& Aertsen, 1999). In order to do so, an understanding of the biology of neurons and how they
interact with one another must be reached. This will be achieved by completing the following
objectives:
Practice computational methods to simulate individual and networks of neurons.
Simulate networks of neurons to understand fundamentals of computational neuroscience.
Extract data from simulations and pre-process for analysis.
Use computational methods to analyse data from simulations.
Recreate proposed network for simulation.
Analyse data for evaluation and compare results.
1.3 MINIMUM REQUIREMENTS AND DELIVERABLES
The following are the minimum requirements set for the project.
1
NEST simulations of individual alpha and delta integrate-and-fire neurons.
2
NEST simulations of 10000 delta integrate-and-fire neurons.
3
NEST simulation of synchronous spiking model.
4
MATLAB script for static and dynamic analysis of populations of neurons.
1.4 RELEVANCE TO DEGREE
This project builds on techniques and applies knowledge learnt from modules on the MSc Artificial
Intelligence in the School of Computing at the University of Leeds. The module Bio-Inspired
Computing (COMP5400M) gave the basis for the understanding and knowledge in the subject area
of computational neuroscience. This knowledge was required to implement the simulations
performed by the software NEST. The module Machine Learning (COMP5425M) was useful to give
an understanding as to how the software NEST used learning rules to simulate different models
connected in various networks. Skills and techniques learnt from the Computational Modelling
module (COMP5320M) were used to analyse and pre-process data obtained from simulations. This
project provided the challenge of further exploring an area of great interest in modern computing
and applying very useful techniques from computational analysis.
2
Simulating Populations of Neurons
George Parish
2013
1.5 PROJECT MANAGEMENT
The initial project schedule is included in Appendix B:1. This schedule changed to reflect the changes
in minimum requirements as the project progressed, most notable the drop of the use of the
software MIIND, another neural simulator tool, and the addition of more MATLAB coding for
analysis. The revised schedule can be seen in Appendix B:2.
1.6 RESEARCH METHODOLOGY
The project aims to understand currently well-defined methods and models to simulate the real
biology of networks of neurons. Because of this, a well laid plan can be followed to reach the final
goal, with little foreseen deviation. Therefore, the waterfall method will be used. Under this method,
background research will be done on each type of simulation defined in the requirements. This will
effectively be the design stage shown in Figure 1. This will be followed by simulating the network
using the software NEST, undertaken during the implementation stage in Figure 1. Following this an
analysis can be undertaken to verify the results. Maintenance will be implemented in the form of
creating and updating new MATLAB script for use in the analysis section. The waterfall approach of a
linear set of objectives is well defined in the planning stage and will be useful for this type of project.
Figure 1 Waterfall Methodology
1.7 RESEARCH QUESTIONS
This project aims to answer the following questions:
How do neurons react to the background noise of other spiking neurons in the brain?
3
Simulating Populations of Neurons
George Parish
2013
Under what conditions can a network of consecutively connected groups of neurons
transmit information?
What input is required for information to be transmitted by consecutively connected
groups of neurons?
Can the implementation of such a network explain real neuronal processes?
Can this implementation be applied in other areas of computer science?
1.8 PROJECT OUTLINE
The purpose of this thesis is to understand and implement evaluation techniques used in
computational neuroscience. This understanding starts in Chapter 2 by describing the principles of
computational modelling and machine learning and how they can be used to solve and evaluate real
biological problems. The notion that everything is made up of a set of processes and can therefore
be modelled is considered.
From there, Chapter 2 continues with an overview of the history of neuroscience. This includes the
examination of the methods used to discover the inner workings of the brain, from both an
anatomical and functional point of view. The structure of individual neurons will be examined. This
will lead to the examination of the electrical properties of neurons which enables them to act as
nodes in an interconnected network. This analysis will culminate with the explanation of the
integrate-and-fire model, a commonly used computational model that aims to replicate the inner
workings of a neuron. Simulations using the software NEST will aid these descriptions.
Once the underlying operations of a single neuron have been modelled, it is necessary to examine
the interaction between many neurons. Chapter 2 goes on to discuss different methods proposed by
relevant literature in which neurons connect and transmit information to one another. Individual
neurons are subject to noise from surrounding neurons that would require the simulation of a large
portion of the brain to emulate. Methods to by-pass this are discussed and implemented via
simulations in NEST.
Chapter 3 goes on to explain how each of the 2 simulations from the requirements list, as well as
one other, are implemented. Each experiment is analysed using MATLAB script to evaluate how the
population of neurons interacts within each network. The importance of each experiment is
explained and there is a logical progression through to the final Case Study evaluations.
Chapter 4 gives a discussion on the results obtained from Chapter 4 and attempts to answer the
research questions proposed. It will also give an overview of the limitations of the project,
requirements fulfilled and proposes future research.
4
Simulating Populations of Neurons
George Parish
2013
2. LITERATURE REVIEW
The following section describes techniques used in this project by analysing the relevant literature.
Firstly, an overview of machine learning is given, followed by how it can be applied to the field of
computational neuroscience. Different types of machine learning algorithms are defined and their
usefulness predicted. This is followed by a section detailing the importance of computational
analysis of data received from machine learning algorithms. Different types of models that are used
in the field of computational neuroscience are defined with reasons for their use. Finally, there is a
detailed description of the real biology behind these models. This is important as an understanding
for the models can only be reached once the underpinning functionality is explained.
2.1 MACHINE LEARNING
Machine learning is a term to describe how learning rules can be applied for computer simulations
to ‘learn’ how to accomplish a task. Machine learning simulations effectively act as an extension of
the human brain, where a complex hierarchy of deterministic rules can sometimes outperform
human intuition, especially when vast amounts of data need to be produced or analysed.
As our methods of understanding the human brain improve, machine learning techniques have been
applied to re-create brain functions and have seen use in many disciplines. Those that study the
brain have used models to replicate brain functions and enhance our understanding of how the
brain enables the mind. Models have been inspired from these studies; artificial neural networks
(ANNs) have seen widespread use across many disciplines, often creating much more efficient
systems.
The argument of pan-computation states that everything can be defined as a set of processes. If a
process can be defined, according to machine learning it should be possible to quantify it and recreate it computationally. Therefore, according to the theory, everything can be learnt by a
computer simulation. The counter argument to this states that functions such as the weather,
digestive system and solar system, cannot be computationally explained by way of a deterministic
set of inputs and outputs, but they can be represented via a stochastic computational model to try
to predict future outcomes (Piccinini, 2007). However, this could simply be because not all
information is known. When we apply this argument to human endeavours to simulate cognition
there are two camps to choose between. Either cognition is the emergent behaviour of sets of
processes that are learnable to a machine, or it is another undefinable process that is separate from
the holistic set of processes it uses. This project assumes that pan-computation is a possibility when
5
Simulating Populations of Neurons
George Parish
2013
it comes to simulating neuronal processes; else ultimately the models proposed throughout the
project would have no justification.
An accurate computational simulation can only be accurately made if all information is known.
Often, this requires too much computational power or there is some uncertainty in experimental
literature. This means that certain assumptions are often made in order to maintain functionality.
These assumptions can be problematic and are usually refined or often redefined when more
knowledge of the process becomes available. This leads to different types of machine learning
algorithms. Some alter their assumptions to create a more functional model that may differ from the
original process. These models can claim to be inspired by the original but honed for a specific
purpose in industry. Other models attempt to replicate natural processes, thus inheriting any
computational problems from the original process. These models are especially hard to create as
they require sufficient testing under certain conditions, yet often still cannot guarantee to replicate
the original process.
When simulating these models to evaluate the brain, these models come under connectionist and
non-connectionist models. Connectionist models maintain a brain-like, network architecture where
connections are neuron-like. Models are usually developed via a connectionist learning algorithm,
assuming representations are distributed and processing is parallel. Non-connectionist models
maintain a functional architecture where the connections are not neuron-like. Models are developed
using suggested functionality from relevant literature of cognitive processes, where no assumptions
are made on representations or processing (Max, 2004).
2.1.1 CONNECTIONIST MODELS
Connectionist models are concerned with trying to replicate biological functions. These
models are made for the sole purpose of understanding the original function by simulating each of
its individual parts in as much detail as possible; because of this they are not useful for industry
purposes. The goal is to study the emergent behaviour from the sum of parts. For this reason, they
can often be computationally demanding, often requiring either a supercomputer or some form of
simplification for large simulations.
The software NEST, used in this project, uses Alpha model neurons to simulate all the known,
intricate processes of individual neurons and the way they interact as a way of simulating whole
populations of neurons. For this reason, section 2.3.2 will explain in detail these intricate processes.
6
Simulating Populations of Neurons
George Parish
2013
2.1 2 NON-CONNECTIONIST MODELS
Artificial Neural Networks (ANNs) are an attempt to create a function that is computationally
similar to a brain function. They are made up of a set of neurons which are manipulated by a
learning algorithm. The neurons are discrete, dynamic instances whose states are dependent on
their input. Dynamic connections between neurons are weighted, allowing a continuous learning
algorithm to manipulate weights, and therefore the neurons’ states. The goal of the model is to
reach an optimised solution that can take into account independent processes with multiple
constraints.
When AANs are used as a connectionist model to represent brain functions, neurons’ states bear
similarities to biological neurons, where non-linear activation rules produce spike-like behaviour.
However, the models cannot represent the full complexity of biological networks and require a
teacher to correct internal elements. It is not always clear whether ANNs represent single or sets of
biological neurons. Old information is easily lost when new information is present. All of this means
that these computational models are useful in testing known hypothesis, but less so when
generating new predictions.
Whilst the software NEST is concerned with making connectionist models to simulate real neuronal
processes, the complexity of this undertaking is such that there are still some simplified models
available. The Delta model neuron learns most of the same processes as the Alpha model neuron but
with some assumptions that limit its functionality as a connectionist model. In the context of the
software, this can be seen as a non-connectionist model. Both models are described in detail later.
2.2 COMPUTATIONAL MODELLING
Computational modelling is a method of representing data in order to show past and predict future
trends or patterns. It is a very useful technique as it can apply formula and rules to infer from large,
complex data sets. The previously described machine learning algorithms that replicate neuronal
processes output a stream of data that can be interpreted by simpler models. Computational models
offer accurate analysis that can guide evaluations of the learning algorithm. There are many
different types of model dependent on the nature of the simulation you are dealing with.
In computational modelling of the brain, there are often huge amounts of data to be processed. The
software NEST allows for the analysis of voltage data and spike data. The former gives a voltage
value (mV) for each neurons ID at every time (ms) of the simulation. The latter gives a neuron ID and
7
Simulating Populations of Neurons
George Parish
2013
a time for every spike that occurred in the simulation. These streams of data can be interpreted in
the following ways to give an analysis of single neurons or populations of neurons.
2.2.1 POPULATION FIRING RATE
The population firing rate gives a frequency in Hz of a single or group of neurons firing rate.
For example, a group of neurons firing at 12Hz will fire at 12 times a second on average. This can be
calculated by counting the number of times a neuron/groups of neurons fired in the data from the
simulation, then divide by 1000 and divided again by 1000 over the length of the simulation.
2.2.2 RASTER PLOT
Raster plots give a basic representation of spike times for individual neurons. It is essentially
a 2-dimensional plot of neuron ID against time in milliseconds. A human analysis of a raster plot is
useful in finding patterns in spike times. A raster plot can also be analysed by an algorithm in more
detail. This is a technique that will be described in more detail in Chapter 3.
2.2.3 VOLTAGE TRACE
The voltage trace is useful for analysing individual neurons. It is a simple 2-dimensional plot
of voltage value (mV) against time (ms). It shows the trace of the neurons voltage as it progresses
through time and can be useful in showing how the neurons membrane potential is affected by
eternal stimuli. These diagrams will be shown in more detail when analysing neurons later in this
project.
2.2.4 POPULATION DENSITY
The previous three approaches offer static representations of a neuron or population of
neurons as a whole. The population density of neurons is a useful, dynamic analysis technique to see
how populations react to external events in real time. It plots a frequency histogram of neurons
according to their voltage values. Voltage values are normalised between 0 (at reversal potential)
and 1 (at threshold). This method and the terms used are all explained in full in the preceding
chapters.
8
Simulating Populations of Neurons
George Parish
2013
2.3 BIO-INSPIRED COMPUTING
An accurate machine learning algorithm for use in simulation is reliant on the accuracy and extent of
the real-world information it is based upon. The information must be identified and quantified, as
well as identify parameters to enable a functional model. Parameters for a model of the brain
include inputs, outputs, the type and multitude of connections, as well as the architecture of the
space that the function operates in. All of these are defined in the relevant literature from
neuroscience, a brief account of which is given in the following section.
2.3.1 THE BRAIN AS A WHOLE
This section deals with understanding the methods used to extract the information that
machine learning algorithms in computational neuroscience are based upon. It gives a brief history
of the achievements in medical neuroscience.
ARCHITECTURE
Cytoarchitectonics is a method used to determine the different cell types in different brain
regions. This is achieved by taking tissue stains and analysing the cell structure. By this method, 52
distinct regions of the human brain were originally identified (Brodmann, 1909); this has since been
redefined as 38 by using modern techniques such as Magnetic Resonance Imaging (MRI) and
electron microscopes – which offer a resolution of about 1 cubic millimetre. This has allowed for a
visual model of the brain as a whole physical object (Figure 2), and the evaluation of the space that
brain functions operate in. The connective fibres of the brain are too dense to allow us to view
individual neurons and connections in this way, as connective highways become merged. The
BigBrain project is the latest project which analyses the state space of the brain (Amunts, et al.,
2013). This method sliced the brain into 7400 sections, stained them and scanned them into a
supercomputer to create a 3D map of the brain – offering an unparalleled resolution of 50 cubic
millimetres.
Figure 2 Brodmann areas of the brain (Gazzaniga, 1998)
9
Simulating Populations of Neurons
George Parish
2013
NEURONS
The method of impregnating neurons in a tissue stain with silver (Golgi, 1898) made it
possible to fully visualise them. This method was used to prove that neurons are independent,
unitary entities (figure 3), disproving the theory that the brain was made up of a continuous mass of
tissue that shared a common cytoplasm (Cajal, 1906). After identifying the structure of a neuron, it
was discovered that they transmitted electrical information in only one direction – a landmark
discovery that forms the basis of artificial neural-network models used today. Neuroanatomists
today use different chemicals in place of silver (Hokfelt, 1984) to create accurate, visual models of
the connections between neurons and their locations in the brain. Neurons from different locations
differ in appearance to suit their functional role. This paper considers pyramidal cells from the
cerebral cortex.
PROCESS
The most definitive method for understanding brain functions is called single-cell recording.
This method is performed on animals (Van Der Velde & De Kamps, 2001) or voluntary humans;
where an electrode is inserted into the brain and is able to record the electrical activity of a single
neuron. The cell of the neuron fires an electrical impulse when active. The goal of this method is to
experimentally manipulate conditions that cause a consistent firing of isolated cells, thus finding the
functional purpose of those cells. A functional model can then describe the interaction of neurons in
a specified region of the brain. However, brain functions enable independent processes in various
brain regions, thus the function cannot be described by the response properties of individual
neurons. However, the understanding gained from this method forms the basis of computational
models of neurons used today.
Figure 3 Types of biological neurons in the nervous system (Gazzaniga, 1998)
10
Simulating Populations of Neurons
George Parish
2013
FUNCTIONAL ROLE
Research has proved that areas of the brain with differing cell structures also represent
different functional regions of the brain. While functions are made up of independent processes
where neurons communicate across varied regions of the brain; it is possible to localise specific
functions to specific regions. The method of electrical stimulation (Penfield & Jasper, 1954)
discovered that motor control was performed by the motor and somotensory cortices; uncovering a
visual model for the motor representation of the body. MRI scanning can reveal how blood flows to
active regions of the brain, enabling a map of functionality of the brain as seen in Figure 4. This
project is concerned with the outermost layer of the brain, the cerebral cortex. It is largely
responsible for memory, attention, language and consciousness (Abeles, 1991).
Figure 4 Anatomy and Functional areas of the brain (http://catalog.nucleusinc.com)
11
Simulating Populations of Neurons
George Parish
2013
2.3.2 STRUCTURE OF A NEURON
The following section details the molecular structure of a neuron and the electrical charge that
molecules can contribute. Only by considering the smallest transitions in a single neuron can we
understand the implications of the interaction between many. This will then allow different models
to be considered in the succeeding section that attempt to replicate the processes described here.
CELLULAR STRUCTURE
The defining feature of a neuron is its ability to transmit information to surrounding cells.
This means they are particularly prevalent in the brain and nervous system, the purposes of which
are to receive and transmit signals across the body. Synapses between connected cell membranes
are the method employed by neurons to achieve their signalling purpose. A synapse is initiated by
the neurons nucleus and travels down the axon where it uses the axon terminal to connect to the
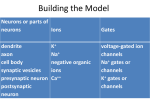
dendrites of other neurons ( Figure 5).
Like other living cells, a neuron consists of a cytoplasm surrounded by a cell membrane. The
cytoplasm is made up of different types of molecules, in particular, positive and negatively charged
ion molecules. A synapse is made possible by ion channels embedded in the cell membrane. These
act as tunnels through the impermeable membrane that allow charged ion molecules to flow in and
out, altering the overall charge of the neuron.
Figure 5 Structure of a neuron (left) and ions traversing the cell membrane (right)
12
Simulating Populations of Neurons
George Parish
2013
As there are different types of ion molecules, different types of ion channels exist that will only allow
passage to a certain type of ion molecule. If the neuron is to operate properly, it is important to
maintain differences in the concentration of ions inside and outside of the cell. This is performed by
ion pumps that expend energy to maintain this equilibrium.
MEMBRANE POTENTIAL
By definition, a neuron must be able to receive and transmit signals. This is done by
undertaking an on/off approach, where the neuron is either active or inactive. By default, a neuron
is inactive – where it’s internal charge from the collection of ion molecules is negative. This overall
charge is called the membrane potential and can be changed by ion channels opening or closing. A
membrane potential can have a charge in the range of about -90 to +50 mV. A neuron rests with a
membrane potential of about 24-27 mV in warm and cold blooded animals. Thus, the scale for
membrane potentials causing a neuron to be active or inactive ranges from about +2 to -3 times the
resting charge respectively. Such a state change caused by the membrane potential is termed an
action potential – and are the cause of synapses.
ELECTRICAL PROPERTIES
Thus far we have described how the membrane potential influences the flow of ion
molecules within a neuron to directly change its state. But how is the membrane potential altered in
order to bring about an action potential? In order to answer this question, we must consider the
electrical properties of the neuron. Figure 6 shows the RC circuit, which works in much the same way
as a model neuron, where a voltage source sends energy through the resistor to be stored in the
capacitor. Once the capacitor’s limit is reached, an excess charge is released.
Figure 6 The Resistance Capacitance (RC) Circuit
13
Simulating Populations of Neurons
George Parish
2013
As previously mentioned, a neurons typical internal charge from its collection of ion molecules is
negative. This means that there has to be a counter balance of positively charged ion molecules just
on the other side of the neuron’s cell membrane. Because of this, the membrane creates a
capacitance (Cm), where the voltage across the membrane (V) and the amount of excess charge (Q)
form the standard equation for a capacitor;
Q = CmV
All neurons have roughly the same capacitance per unit area of membrane of about 10 nF/mm 2. The
membrane capacitance is proportional to the surface area of the neuron. Neuronal surface areas
tend to be in the range 0.01 to 0.1 mm2, so the membrane capacitance for a whole neuron is
typically in the range 0.1 to 1 nF.
Determining the membrane capacitance is essential when it comes to calculating how much current
is required over time to change the membrane potential. This is given by the formulae;
Cm
=
where dQ/dt represents the cells incoming current and Cm dV/dt gives the amount of current
required to change the membrane potential.
REACTION TO EXTERNAL STIMULI
Now that we understand how the voltage of a membrane potential can be changed, the next
step in causing an action potential is to hold the membrane potential steady at a value that is not its
resting one. This requires constant current (Ie) to enforce a change in voltage ( V). According to
Ohm’s law;
V = IeRm
there has to be a resistance to input. In this case, it is called the membrane resistance. This
resistance is proportionate on the surface area of the neuron. For neurons with an area in the range
0.01 to 0.1 mm2, the membrane resistance is usually between 10 and 100 M . For example, to
cause an action potential of 50 mV where the resting value of a membrane potential is 25 mV and
the membrane resistance is 50 M , we need a constant current of 0.5 nA to hold the membrane
potential the required 25 mV from resting value.
14
Simulating Populations of Neurons
George Parish
2013
Changes in the membrane potential have to occur on a constant time scale, called the membrane
time constant. This is the membrane resistance times the membrane capacitance, typically between
10 and 100 ms.
m
= RmCm
INTRACELLULAR ACTIVITY
When measuring the membrane potential in different parts of a neuron, they often give
different values. These different charges across the neuron cause ions to flow within it in order to
equalize the differences. However, the neuron produces a resistance to this flow of ions in order to
maintain itself – this is called the intracellular medium. This is particularly high in long narrow parts
of the neuron – for example dendritic or axonal cable. In such parts, the longitudinal resistance can
be calculated as inversely proportional to the cross-sectional area of the neuron segment. The
degree of this proportionality between resistance and area is called the intracellular resistivity.
All of this means that it is possible to calculate how much voltage is required to force n amount of
current down a neuronal segment of any size. This is important when it comes to predicting the
ramifications of voltage from an action potential forcing current through the neuron and causing it
to interact with other neurons. The intracellular resistance to current flow means that different parts
of the neuron will measure different voltages at the time of an action potential. Because smaller
neurons with less axonal or dendritic cable have less intracellular resistance, they are termed
electrotonically compact and can be described by a single membrane potential as opposed to many.
THE ROLE OF DIFFUSION
The electrical properties of neurons are not the only force responsible for the flow of ions in
the ion channels. As mentioned previously, the concentration of ions on the inside and outside of
the cell membrane is maintained by ion pumps. These operate by way of diffusion. Thus, there is a
balance between the probability that an ion has sufficient energy to overcome the membrane
potential and enter or exit a neuron, and the ion traversing through an ion pump due to the
diffusion gradients. In other words, there must be a balance between electric and diffusive forces.
This occurs because if the former were solely responsible, there would be no control over the
proportion of ions inside and outside of the neuron – potentially leading to instability.
15
Simulating Populations of Neurons
George Parish
2013
FINDING AN EQUILIBRIUM
The diffusive properties of the ion pumps compensate for this by controlling the rate at
which ions flow in and out of the neuron. This can be given by the Nernst equation;
E=
(
ln
)
Where E denotes the potential of an ion channel required to enforce an equilibrium between the
two forces, VT is the voltage of the membrane potential, z is the charge of the ion and [outside] and
[inside] denote the concentration of ions both sides of the neurons cell membrane.
The value of E lies within different ranges dependent on the type of ion, for example potassium (K+)
ion channel potentials lie between -70 and -90 mV, whilst sodium (Na+) and calcium (Ca+) ion
channel potentials are 50 and 150 mV or higher respectively. As mentioned previously, this shows
how ion channels can be highly selective. We can approximate for E over the different ion channels
using the Goldman equation. E can then be termed a reversal potential, as when the membrane
potential changes value to greater than or less than E, the flow of ions in that channel reverses.
A neurons reversal potential determines what type of neuron it is. For example, a depolarized
neuron (more positive membrane potential) is when there are many sodium or calcium ion channels
active in a neuron. A hyperpolarized neuron (more negative membrane potential) is when there are
more active potassium ion channels. As there are two types of neuron because of the reversal
potential, there are also two different types of synapses. Inhibitory synapses occur when a nearby
hyperpolarized neuron spikes, lowering the neurons membrane potential. Excitatory synapses occur
when a nearby depolarized neuron spikes, raising the neurons membrane potential.
THE MEMBRANE CURRENT
Lastly, the membrane current is simply the total ion current flowing across a membrane.
When ions leave a neuron, the current is positive, when ions enter, the current is negative. As
neurons are of different size, the membrane current per unit area is more useful – denoted by im.
The equation for the membrane current is as follows;
im = ∑i gi (V - Ei)
16
Simulating Populations of Neurons
George Parish
2013
The reverse potentials of different types of ion current are denoted as Ei where i is an ion type. The
difference between the membrane potential (V) and the reverse potential (Ei) is called the driving
force, and the factor gi is the conductance per unit area.
At this point we can see that certain factors attributing to the membrane current do not fluctuate
much and remain somewhat constant. Therefore we can make a simplifying assumption that they
will always remain constant and lump them together to form a single term;
ḡL(V – EL)
This is called the leakage or passive conductance. Because of the assumptions about these values
seemingly constant nature, it is important not to use this formula as the basis for others, but to use
it to manually adjust the resting potential of the model neuron to that of the real neuron.
2.3.3 THE INTEGRATE-AND-FIRE MODEL
The integrate-and-fire model is the most commonly accepted model used when simulating
neurons. It is a simple model that allows any external inputs to alter the membrane potential (V) of
the neuron measured in millivolts. The model specifies a resting potential (E) which is the normal
resting value of the membrane potential. There is also a threshold (Vth) value and a reset (VR) value.
The threshold value specifies at what value an action potential occurs and the reset value gives the
value to which the neuron is reset to after an action potential. The integrate-and-fire model can
simulate many different types of neurons by allowing these attributes to change.
The software NEST allows for different types of integrate-and-fire neurons dependent on how much
simplicity is required. Two such models will be considered in this project, the alpha model and the
delta model. The alpha model works more like a real neuron, where the conductance of different ion
channels has more of a direct effect on the overall voltage of the membrane potential. The delta
model makes the assumption that this conductance based approach has minimal effect on the
neuron, and so uses a leakage term (as described previously) that assumes there is no variation in
the overall conductance of the membrane. This means that external events (i.e. the spiking of
neighbouring neurons) have a constant effect on a delta integrate-and-fire neuron, but a varying
effect on an alpha integrate-and-fire neuron – dependent on the conductance of ion channels at the
time of the event.
17
Simulating Populations of Neurons
George Parish
2013
SOLUTION TO THE INTEGRATE-AND-FIRE MODEL
The standard equation for an integrate-and-fire model can be given as a homogenous or
non-homogenous equation, dependent on what is required. The homogenous equation (below left)
is the formula for the membrane potential with no external input, the non-homogenous (below
right) allows for external inputs such as constant currents or membrane conductance.
The homogenous equation (left) dictates that the change of voltage over time (dv/dt) multiplied by
the membrane time constant ( ) is equal to negative membrane potential (-V). It is worth noting that
the value V here represents the driving force between the resting potential and the membrane
potential (E-V), and the varying conductance of the membrane potential due to the opening and
closing of different ion channels (gi) is assumed constant and ignored. The non-homogenous
equation in its simplest form (right) gives the same formula but with an added input of a constant
current (I). Solving these equations allows the calculation of the neuron’s membrane potential at any
time.
The solution to the homogenous equation for the integrate-and-fire model (left) shows how the
voltage at time t (V(t)) is given by the voltage at time zero (V(0)) times the exponential (e) to the
power of minus time t divided by the time constant tau (-t/ ). We call this solution the ‘general
solution.’ We then find any solution to the non-homogenous solution and call it the ‘particular
solution.’ A solution in this case is we assume V is constant and then derive V=I as a solution. The
‘most general’ solution can then be found by adding the ‘general’ and ‘particular’ solutions together.
This gives us
where k is solved to give the solution to the non-homogenous
equation on the right.
18
Simulating Populations of Neurons
George Parish
2013
CALCULATE TIME BETWEEN SPIKES
We can use the solution to the non-homogenous equation (above right) to find the time
between spike events of the integrate-and-fire model, generated because of the constant current
input I. This is done by looking at when this solution is equal to the threshold value of the model ( )
and solving for t.
Where time (t) is equal to the time constant ( ) times the natural log of the voltage at t=0 minus the
input current value (I) divided by the threshold value ( ) minus the input current value (I). This gives
the time steps in between spike events assuming that the current stays constant and there are no
other external events.
REFRACTORY PERIOD
The integrate-and-fire model makes some assumptions that deviate from the original
biological function. This includes using a simplified model to initiate an action potential as well using
a linear approximation for the membrane current. These assumptions mean that the model will
likely not perform in the same way as its biological counterpart, meaning that the model has to be
manually tweaked if it is to replicate the original function with a certain degree of accuracy. In this
case, within the model, an action potential could theoretically be caused instantly after the previous
action potential. This we know could not be true, as a biological neuron requires more time for
particular ions to traverse its membrane in order to change the overall conductance of the
membrane potential and cause a second action potential.
This problem is solved by implementing a refractory period. After an action potential, the neuron is
clamped to its resting potential for a specified time, causing an absolute refractory period. This
means that the neuron will ignore any external inputs during this time. Immediately after the
specified clamp time, a relative refractory period is introduced, where all external inputs are
weighted so they have a smaller effect on the neuron. This means that a second action potential is
inhibited but not impossible within this time. After the refractory period has finished the model
neuron continues as before.
19
Simulating Populations of Neurons
George Parish
2013
DELTA MODEL
The delta model is the implantation of the leaky-integrate-and-fire model described earlier,
where the membrane potential jumps on each spike arrival. The event where the threshold is
crossed is followed by an absolute refractory period where the membrane potential is clamped to
the resting potential. Any spikes that arrive during the refractory period are discarded. This can be
considered as a non-connectionist model in context of the software NEST, as it makes some
simplifying assumptions.
POST-SYNAPTIC SPIKE
Figure 7 is the NEST implementation of a single delta model neuron receiving a single spike
from a neighbouring neuron. The amount the potential goes up depends on the weight of the
connection between the two neurons. The higher the weight of the connection, the more a spike
from one neuron will influence the other. As the neuron is a delta model any effect from external
events is constant, and so the effect of the spike on its membrane potential is proportional to the
weight. This can be seen in the excitatory connection diagram from Figure 7 where weights of 5, 10
and 15 were used. If a single external spike had enough effect on the neuron to immediately raise
the potential to its threshold of 55mV, then it would also immediately spike, thus re-setting it to its
reversal potential of -70mV and enabling the refractory period – somewhat unrealistically causing no
change in potential overall.
The inhibitory connection diagram of Figure 7 shows how the neurons potential goes down when
there is a negative connection between the two neurons. This is to simulate how inhibitory synapses
from biological neurons with negatively charged ions affect their neighbours. Once the potential of
the delta model neuron has been changed by the excitatory or inhibitory spike, there is a gradual
change back to the reversal potential of -70mV. This simulates how the neuron’s membrane
gradually releases the charged ions that influenced the charge of its membrane potential.
Figure 7 Excitatory and Inhibitory connections for a delta model neuron
20
Simulating Populations of Neurons
George Parish
2013
CONSTANT INPUT
Figure 8 shows how a DC generator can be connected to a single neuron. The voltage
generated by the DC generator is transferred to the membrane potential of the delta model neuron
(d1) via a positive connection.
Figure 8 DC generator and single delta neuron network
Figure 9 shows a single integrate-and-fire neuron with a varying direct current input over time. It can
be seen how the current needs to be of a sufficient amount to cause the membrane potential to rise
to the threshold value (here 55mv). Once it hits threshold, the potential is reset to -70mv before
rising again due to the current. The strength of the input current determines how fast the
membrane potential rises, as can be seen in the period 1500-2000ms where the firing rate is much
faster due to a higher current.
The method of supplying the model neuron with a direct current input is used to test the
capabilities of the model and is not meant to replicate real processes. Realistically, a neuron’s inputs
will be from surrounding neurons that spike irregularly, causing an almost unpredictable and
irregular stream of inputs that can alter the membrane potential.
Figure 9 A single integrate-and-fire neuron with a single direct current input of 200pA (500-1000ms), 380pA (1000-1500ms) and 450pA
(1500-2000ms)
21
Simulating Populations of Neurons
George Parish
2013
Figure 10 is a graph showing the spike frequency of the model neuron against the value of the direct
current input. This graph shows how there needs to be a sufficient current in order to cause spiking
in the neuron, shown by spike frequency only rising past the 380pA mark. There is initially a sharper
rise in spike frequency due to a stronger current, but as more current is applied, this rise is less
dramatic. This is due to the spike frequency rising and encroaching on the territory of both the
relative refractory period and the absolute refractory period causing the input to have less of an
effect to having no effect at all. If there were no refractory period modelled, the spike frequency
would rise in a linear fashion.
Figure 10 Spike frequency of model neuron vs value of the direct current input
ALPHA MODEL
As mentioned previously, alpha models aim to be a more realistic representation of a
biological neuron. The effect of external stimuli on the alpha model neuron is not constant as they
depend on the conductance of the ion channels at that point in time. Ion channels operate by
opening and closing a swinging gate. As there are different types of ion channels, dependent on
which one was open, the neurons membrane potential will take on different characteristics. For
example, if potassium channels are open then the neurons potential will be more hyperpolarised
(more negative) and if sodium or calcium channels are open then the potential will be more
depolarised (more positive). This is considered a connectionist model in the context of the software
NEST as it does make any assumptions in its model.
22
Simulating Populations of Neurons
George Parish
2013
SOLUTION
In order to calculate the effect of an input on the model neuron, we much calculate the
probability that a certain ion channel gate is open.
The above equation describes the gating equation, where the probability of the gate being open
over time (dn/dt) is equal to the opening rate of the gate (
the gate is closed (1-n). The closing rate of the gate (
) multiplied by the probability that
) times the probability of the gate being
open (n) is then taken from this. The opening and closing rates can be described as follows:
(
)
(
)
The two formulas for the opening rate and closing rate are given by two exponential functions.
These functions depict the energy required for the gate to open and close, where
&
represents the amount of charge being moved as well as the distance it travels and
show
&
the energy required to close or open the gate.
POST-SYNAPTIC SPIKE
Both of these exponential functions can be seen in action in Figure 11, where a single alpha
model neuron receives one spike from a neighbouring neuron. The Alpha model neuron from NEST is
supplied via connections modelled in the same way as from the balanced excitation/inhibition model
from the documentation with NEST (Balanced random network model). The rise in membrane
potential due to the spike is modelled by the exponential for the opening rate, and the fall in
membrane potential after the spike is modelled by the exponential for the closing rate.
We can calculate the time that the voltage takes to reach the peak of an alpha synapse by deriving
the equation of the alpha model at t=0. The maximum value of the alpha synapse is called the Peak
Amplitude and can be found by obtaining the voltage value (mV) at t=0. The influence of an alpha
synapse on another neuron can be found by computing the area under the curve.
23
Simulating Populations of Neurons
George Parish
2013
The weight of the connection between the alpha model neuron and its neighbouring neuron
determines how much the membrane potential of the alpha model neuron will rise by. The effect of
changing this weight can be seen in the first graph of Figure 11, where a higher weight allows the
neighbouring neuron’s spike to influence our alpha model neuron the most. By changing the risetime of the alpha model neuron, we can influence the length of time that the potential will rise for.
The longer the time, the longer the incoming spike can influence the membrane potential, causing it
to rise to a higher value. This is shown by the second graph of Figure 11. The final graph of Figure 11
shows the effect of changing both the weight and rise-time. The dotted lines show that by increasing
the time-to-peak we magnify the effect of a spike with both a high and a low weight. We can
conclude that by increasing the weight or time-to-peak will allow alpha synapses to have a greater
influence on other neurons.
Figure 11 Alpha model analysis
24
Simulating Populations of Neurons
George Parish
2013
2.3.4 MODELLING POPULATIONS OF NEURONS
When modelling populations of neurons it is important to consider a few important steps.
The most important of these is how these neurons are connected. As neurons do little more than
essentially turn on and off, it is the connections between neurons that allow for vast networks to
communicate information. Connections indicate how likely a signal from one neuron will affect many
other neurons. Another aspect to be considered is how a background population of neurons with
many connections will affect the study of a single or group of neurons. This background noise plays a
key role in the state of neurons, oftentimes keeping whole groups of neurons poised just below their
threshold for action potential, ready for a push from a more direct stimulus to activate them.
CHAINS OF NEURONS
Neurons transmit information by sending signals that must traverse networks containing
large numbers of neurons. In order to do this, a signal must cross many neurons simultaneously. This
has been hypothesised to work in a few ways; a popular way of visualising such a transmission of
information is through a serial or parallel chain. This method will be shown as inefficient when
compared with converging and diverging connections between neurons.
SERIAL/PARALLEL CHAINS
A serial chain is where neurons are connected in a sequential linear chain, where a signal
goes through each neuron in turn. The initial action potential that causes spiking through this type of
network would need to very strong if the signal is to be transmitted through to the end node. This is
due to dispersion in the signal as it is transmitted from node to node. The single connection between
each node doesn’t give the signal much chance of successfully traversing the network.
Figure 12 Serial and Parallel chains
25
Simulating Populations of Neurons
George Parish
2013
A series of serial chains could work in parallel, making it more probable that a signal would reach its
destination. In this set-up there are w amounts of serial chains of length n, as seen in Figure 12. It
has been found that this type of formation is very unlikely in the cortex (Abeles, 1991). The reason
being that it does not offer a very flexible solution to the very complex problem of successfully
transmitting a signal across such a vast network. Signals would have to waste time and energy crisscrossing through a series of road-like parallel chains, allowing congestion and dispersion to hamper
them. It might be much faster for neurons to have direct links to many different areas, allowing
them to act as an airport-like hub where signals converge from and diverge to many different
locations.
DIVERGING/CONVERGING CHAINS
In a chain of diverging and converging connections, each neuron in a group sends excitation
to various neurons in the next group, and each neuron in this next group is excited by several
neurons of the previous group. Figure 13 shows how several neurons outputs can converge onto
one, as well as how one neuron can diverge and send signals to many other neurons. By considering
each neurons capability of having multiple incoming converging and outgoing diverging connections,
we can form a chain of such neurons as seen at the bottom of Figure 13. This limited view shows
how neuron a(0,1) receives input from some other neurons before sending output to the other
neurons in the chain, who in turn also receive input from some other neurons.
Figure 13 Converging and Diverging chains
26
Simulating Populations of Neurons
George Parish
2013
A group can be made up of neurons that are physically in the same region of the brain; however,
they can also be made up of neurons from various different areas. In this structure it is important to
view such a chain more as a time series rather than a physical chain, where a neuron is excited after
another because of their connection and not because it is physically nearby. This structure makes it
possible for a strong signal to activate many different groups of neurons in very different parts of the
brain, perhaps explaining how a smell can evoke images or feelings, utilising different areas of the
brain simultaneously.
COMPLETE & INCOMPLETE CHAINS
In order to better explain and simulate these chains we often limit our consideration to
chains where the width (w) of each chain is constant, so that for all neurons the average number of
converging connections is equal to the average number of diverging connections. We can consider
the number of neurons in each group as the width (w) of the chain, and the degree of
divergence/convergence as the multiplicity (m) of connections between groups. The case where
w=m is termed Griffith’s complete chain (Griffith, 1963), and represents how each neuron in one
group will excite every neuron in the next group. It has been proven that in the cerebral cortex we
are most likely to find complete chains of up to width 5, but these chains would not be functional as
the synapse strength would need to be higher than is thought probable for cortical neurons (Abeles,
1991).
Alternatively, it is possible to find incomplete chains of neurons, where w m. This means that the
amount of connections from and to each neuron (m) is not equal to the total number of neurons
(w). This arrangement is much more likely to be found in the brain as connections can be formed
spontaneously and it would be inconvenient for a neurons number of outgoing connections to have
to equal its incoming connections. The software NEST allows for a choice between diverging,
converging and regular connections. Any neuron can connect to any other without the limitation of
being part of a complete chain.
FEEDBACK NETWORK
What has been described so far is a feed-forward network, where a group of neurons will
excite the next group and so on. A feed-back network can be implemented when the output from a
group of neurons feeds back as its own input. This means the state of this group will be constantly
changing until it reaches equilibrium, making this type of network efficient at solving more complex
problems. If the output of the final group of neurons in a feed-forward network feeds back into the
27
Simulating Populations of Neurons
George Parish
2013
first group of neurons then a feed-forward network can also become a large feed-back network of a
sort. This project considers the use of feed-forward networks and if they encourage synchronous
spiking between groups of neurons, so networks of a more feedback architecture are not covered in
extensive detail.
GENERATING EXTERNAL EVENTS
So far, we have limited our discussion to complete groups of interconnected neurons. Whilst this is
true for the brain as a whole, it is currently impossible to have knowledge of and simulate every
neuron and their connections. When considering a group of neurons in simulation, each neuron will
be connected to any number of surrounding neurons. Whilst it isn’t feasible to simulate them all, it
isn’t sensible to ignore them either.
POISSON DISTRIBUTION
The best approach is to attempt to model the influence of the spiking of surrounding
neurons. This is done by using a poisson distribution. It is a discrete probability distribution that
takes a given number of events and gives a probability of these events occurring independently
within a fixed time interval. Events can be distributed by the following equation:
Where a the probability of a discrete event X occurring at time interval k is equal to some constant λ
to the power k times the exponential to the power minus λ, all divided by k factorial. The index k in
this case represents the time between events, where an event X has a probability of occurring k
milliseconds after the previous event. The constant lambda (λ) is equal to the expected value of X,
and will have a profound effect on the distribution. For a poisson distribution for neural simulators, a
lambda is chosen such that the exponential fit to the data is as can be seen in Figure 14, where an
event X is most likely to occur very shortly after the previous event.
The software NEST allows the user to create a ‘poisson generator,’ that generates events at a
specified rate. For example, the effect of the 20,000 surrounding neurons of the cerebral cortex can
be modelled by creating poisson generators, 88% of which fire at 2Hz connected with excitatory
connections, and 12% of which fire at 12Hz connected with inhibitory connections.
28
Simulating Populations of Neurons
George Parish
2013
Figure 14 Poisson distribution of spike events
INDIVIDUAL NEURON WITH POISSON EVENTS
Figure 15 is a representation of how poisson generators (p1 &p2) are connected to a neuron
(d1). By taking a positive connection (p1), any events generated by a poisson generator will have an
excitatory effect on the neuron’s membrane potential. With a negative connection (p2), the events
will have an inhibitory effect. In this particular simulation, a DC generator is also constantly pumping
in positive voltage to the delta model neuron.
Figure 15 Network for 1 DC and 2 Poisson generators connected to a single delta neuron
Figure 16 shows this simulation in action. During the first 500ms, the rate at which excitatory events
come in directly raises the neurons potential without the need for a DC generator. This rise of the
potential is halted due to the rate at which inhibitory events come in, holding the potential at a
steady state. This means that the membrane potential neither rises nor falls too far from a certain
value, and we can see these fluctuations in Figure 16. Because of these extra additions to the
29
Simulating Populations of Neurons
George Parish
2013
potential, we can see that the DC generator has to provide much less amplitude compared to Figure
9 to cause the neuron to fire – 90pA compared to 380pA. Between 1000-1500ms we can see that the
noise generated by the poisson generators sometimes exceeds the threshold, causing the neuron to
spike and the potential to be reset. This firing is irregular due to the nature of a poisson distribution
and is a good simulation of how neurons are often kept ‘trigger happy’ (just below threshold) and
will fire if the firing rates of neighbouring neurons provide enough stimulus.
Figure 16 Simulation of delta neuron with poisson and DC generators
30
Simulating Populations of Neurons
George Parish
2013
3 EXPERIMENTS & EVALUATION
The following chapter details the results and evaluations from the simulations detailed in the
minimum requirements, as well as some supporting simulations.
3.1 IMPLEMENT A NEST SIMULATION
This section gives the code for the first simulation as an example of how NEST was used in this
project. The first step is to create a set of nodes for the simulation. There are many different types of
models available in NEST, however, this project focuses on just the Alpha and Delta model neurons.
A poisson or DC generator is also created as a node. We also have to create a spike detecting node
and a voltmeter node to record information from our simulation. For example we create a network
of 10000 neurons with a = 50ms as follows:
neurons = nest.Create(‘iaf_psc_delta’,10000,{’tau_m’:50.0})
noise = nest.Create(’poisson_generator’,1,{‘rate’:800})
voltmeter = nest.Create(‘voltmeter’)
spikeDetector = nest.Create(‘spike_detector’)
nest.setStatus(voltmeter,{‘to_file’:true})
nest.setStatus(spikeDetector,{‘to_file’:true})
The poisson generator we have created fires at 800Hz. This is the equivalent of 800 generators firing
at 1Hz. We set the status of voltmeter and spikeDetector to write to file, this creates a file in the
current directory of nest for each node. Voltmeter data gives the neuron ID, time (ms) and voltage
(mV), whilst the spike detector data gives neuron ID and time (ms). We then connect the noise to
the neuron via an excitatory connection so that noise adds to the membrane potential.
nest.divergentConnect(Noise, Neurons,0.45,1.0)
nest.convergentConnect(neurons,spikeDetector)
nest.divergentConnect(voltmeter,neurons)
This command uses a divergent connect to connect the noise node to every node in neurons. A
weight of 0.45 was used to indicate a spike from the poisson generator influences the potential of
neurons by a factor of 0.45. A delay of 1.0 has been used to indicate a millisecond delay from the
event occurring to it effecting the potential of neurons. This is to simulate the length of time it takes
for signals to travel through connections, in real biology this value would likely be dependent on the
distance between the connected neurons. Each neuron signals to the spike detector when a spike
occurs via a convergent connection, and the voltmeter is connected to every neuron via a divergent
connection. We can then simulate the network for 1 second (1000ms) with the command:
nest.Simulate(1000)
31
Simulating Populations of Neurons
George Parish
2013
3.2 SIMULATION OF 10,000 INTEGRATE-AND-FIRE NEURONS
Now that we can simulate the effect of neighbouring neurons on a single neuron, we can start
building different networks of neurons. The following is the implementation of a group of 10,000
neurons each with input from 800 surrounding neurons that fire at a rate of once a second (1Hz).
3.2.1 IMPLEMENTATION
Figure 17 shows how such a network is implemented. A group of 10,000 delta neurons (d1d10000) were given input from 800 poisson generators (p1-p800) firing at 1Hz. Each poisson
generator provides input to every neuron, and each neuron receives input from every poisson
generator.
As the neurons are delta models, the poisson generated events will have a constant effect on each
neuron’s membrane potential. The connections are all excitatory, meaning that we can expect two
situations. The potential will either continually rise and reach threshold due to the events, or the
potential will fluctuate around a certain value if the rate of the generators is not high enough.
Figure 17 Network of 10,000 delta neurons connected to 800 1Hz poisson generators
3.2.2 EVALUATION
There are several methods to analyse networks of neurons. Some are shown in Figure 18,
which analyses the performance of the network. A raster plot can be seen in the top left of Figure
18, this plots each neuron ID against time, where a blue dot indicates that a neuron has reached
threshold (-55mV) and fired. Below the raster plot is the population firing rate which plots the
average firing rate of every neuron against time.
32
Simulating Populations of Neurons
George Parish
2013
The analyses of both diagrams show that that there are no firings for the first 35ms. This is due to
the fact that the initial membrane potential started at the reversal potential of the neuron (-70mV).
The poisson events cause the membrane potential to rise and we see that all of the 10,000 neurons
begin firing between 50-100ms. This is indicated by a thick band of blue dots on the raster plot and a
peak of 20Hz in the population firing rate at 80ms.
Once a neuron fires a refractory period is enforced where the potential is clamped to its reversal
potential. As most of the neurons fire together, there is a lull in the firing rate of the population at
about 120ms due to the time it takes for the poisson generators to raise the potential back to
threshold. This is supported by the period 110-140ms on the raster plot, where a less dense band is
clearly visible. Due to the nature of the poisson distribution, it takes neurons different lengths of
time to once again reach threshold. This causes the population to reach a steady firing rate of 12Hz.
Whilst the raster plot and population firing rate diagrams offer a static way of analysing populations
of neurons, we can analyse more dynamically by showing the population density through time. This
method takes the voltages of each neuron and plots it against frequency. The values for voltage (xaxis) range from 0 (reversal potential) to 1 (threshold). This method allows us to see where the bulk
of the population lies at and see developing patterns. The population density for different times can
be seen in Figure 18a-c, where each time is marked by a-c in the bottom left time diagram.
Figure 18 Analysis of 10,000 neurons with noise
33
Simulating Populations of Neurons
George Parish
2013
Figure 18a shows the population at 35ms. This diagram shows how the poisson distribution is clearly
in effect as the neurons move up together from 0 to 1 in an evenly distributed blob. Figure 18b
shows the density at 70ms, during the densest band of the raster plot and just before the peak of
12Hz in the firing rate. The density diagram shows that most neurons are in two locations. There are
many neurons at 0 (in refractory) which have just fired and many neurons just about to fire bulking
up towards 1. Neurons coming out of refractory tend to bunch together due to the clamping effect;
this can be seen in the tall bars until 0.2 on the voltage scale. Figure 18c is a density diagram at
about 250ms and shows how the bulk on the right has shallowed in a wave-like effect. The fact that
there are less neurons at 0 and the population density is more evenly distributed indicates that the
population has reached a steady firing rate, supporting the analyses of the population firing rate and
raster plot diagrams at this time.
TIME BETWEEN SPIKES
Figure 19 plots a histogram of the time between spike events for the whole population of
10,000 neurons. At first glance we can see that it resembles the poisson distribution shown in Figure
14 as a spike most often occurs just a short time after a previous spike. The difference is that the
refractory period enforced on neurons combined with the time it takes to re-reach threshold means
it is impossible for them to spike immediately after a previous spike. However, Figure 19 shows that
the spiking of neurons subject to background firing of neighbouring neurons is poisson in nature.
This helps understand how a population reacts to noise – which was the purpose of this simulation.
Figure 19 Time between spikes of 10000 neurons with noise
34
Simulating Populations of Neurons
George Parish
2013
3.3 THE BALANCED EXCITATION/INHIBITION MODEL
The purpose of the previous simulation was to see how background noise causes a group of neurons
to react. The objective of this next simulation is to see how different groups of neurons can affect
each other.
The balanced excitation/inhibition model is an example network given in the
documentation for the software NEST. The aim is to establish a balance between excitatory and
inhibitory populations of neurons that interact with each other to cause a consistent spiking pattern.
3.3.1 IMPLEMENTATION
Figure 20 shows how the balanced network was implemented. A group of excitatory neurons
(E1) and a group of inhibitory neurons (I1) were both connected to another group of excitatory
neurons. The inhibitory neurons were connected via a negative connection to reflect how
neighbouring neurons high in potassium (K) lower the membrane potential of the connected neuron.
The excitatory neurons were connected via a positive weight to reflect how neighbouring neurons
high in calcium (Ca) or sodium (Na) raise the membrane potential of the connected neuron. All
neurons were modelled using the Alpha model neuron in NEST. Connections were based on an
alpha-function to reflect the realistic time dependent connections of neurons.
All neurons were connected to a poisson generator (p) to reflect the goings on of surrounding
activity. The poisson generator (p) was modelled assuming that background noise of neurons
generally keeps neurons ‘trigger happy.’ It calculates how many excitatory spikes are required for
the population of neurons to be kept just below threshold and feeds this to all groups of neurons.
The intention of this network is for the excitatory population E2 to receive a balance of excitatory
and inhibitory spikes that cause the population to spike in a regular pattern.
Figure 20 Network of balanced excitation/inhibition model
35
Simulating Populations of Neurons
George Parish
2013
3.3.2 EVALUATION
Figure 21 shows the analysis of the balanced network. The raster plot shows how a
consistent firing pattern was achieved. Groups of neurons would fire together before being clamped
via a refractory period. Their potentials would then rise together to fire the neuron again. We can
see this by the repeated bands of blue dots followed by white space. The bands are not perfectly
straight and in fact follow a wave pattern looking down the y-axis (neuron ID). This shows how
different groups of neurons in the network synchronise together, yet each group’s firing pattern is
similar to the firing patterns of its surrounding groups. This could be because of fluctuations caused
by noise at the start of the simulation are exaggerated as time progresses. This is supported by the
fact that all neurons initial spike was at the same time, causing the first band on the raster plot to be
fairly linear. As time progresses, the bands become more wave-like.
The population firing rate of Figure 21 reflects the findings of the raster plot. The firing rate is at its
highest when all neurons fire together at the start of the simulation. It then falls back to almost zero
as all neurons are in refractory together. This pattern is repeated, however, each time the
population spikes the firing rate is raised to a lower peak and falls to a higher trough. This reflects
how the population as a whole spikes with less synchronisation as time goes on. After about 200ms
a steady firing rate is established. Minor fluctuations in this rate show the same pattern of the firing
Figure 21 Analysis of balanced excitatory/inhibitory network
36
Simulating Populations of Neurons
George Parish
2013
rate reaching a peak before falling to a trough, indicating that the population has retained some
synchronicity in its spiking pattern. This supports the analysis of the raster plot where bands of blue
spikes and white space are clearly visible throughout the simulation.
The pattern for synchronous spiking in the balanced network can best be seen in the population
density diagrams of Figure 21a-c. Figure 21a is taken as the population moves together out of
refractory towards threshold just before the populations 3rd spike. This shows how even after 2
spikes, the population is still very much synchronised together. In Figure 21b we can see how the
population is much more spread out. Whilst neurons are spread between being in refractory and
spiking, we can still vaguely see a bulk of neurons between 0.4 and 0.8 on the voltage scale. This bulk
is represented in the population firing rate diagram, where at 90ms the rate has yet to reach a
steady state.
Figure 21c shows the population density once the firing rate has reached a steady state and depicts
the true nature of the balanced network. Neurons are equally spread out between the reversal
potential and threshold. Neurons move up through the voltage scale in packets, supporting the
analysis of the raster plot where neurons maintain a local synchronous spiking pattern, yet fire at an
asynchronous steady state as a population as a whole. As neurons are constantly spiking, there are
always many in refractory, reflected by the large bar at 0 in Figure 21c.
Previously we have shown how a steady population firing rate can be achieved via the constant
spiking of neurons in a noisy fashion. Here we have shown how a population can seem to be working
in a similar fashion, but the underlying synchronisation between groups of neurons shows how this
balance can also be achieved through organisation. It is also more realistic as it takes into account
groups of both excitatory and inhibitory neurons, whereas the previous simulation only considered a
population of excitatory neurons.
37
Simulating Populations of Neurons
George Parish
2013
3.4 THE STABLE PROPAGATION OF SYNCHRONOUS SPIKING MODEL
In this model, a volley is propagated through consecutively connected groups of neurons, all under
the influence of the repetitive firing of surrounding neurons. The model proves that, because all the
neurons are kept ‘trigger-happy’ by surrounding neurons, consecutive groups of neurons are pushed
over the edge by the first part of the incoming signal. This means that the amount of neurons firing
in each layer increases and the variance in firing times decreases as the volley propagates - causing
synchronisation. However, experimentation showed that the initial volley must be over a certain
number of numbers firing over a certain variance for the signal to propagate.
Figure 22 Example of a synchronised and de-synchronised volley
Figure 22 gives an example of how the initial volley is strong enough to be propagated through
layers of neurons. Synchronisation occurs when neurons fire together within a shorter time period.
De-synchronisation occurs when the initial volley is not strong enough to be propagated. Whilst
some neurons in the subsequent layers fire, there are not enough to cause major activity through
the connected layers.
3.4.1 PREVIOUS RESEARCH
Previous research (Diesmann, Gewaltig, & Aertsen, 1999) has shown that the initial volley
must be over 50 spikes and fully synchronised (no variance in time of firing,
=0), or over 90 spikes
with less synchronisation ( =3). Anything less and the volley will become de-synchronised as shown
in Figure 22. As the volley propagates, there is an attractor point where the final synchronised layers
of neurons contain about 90 spikes with a low variance ( =0.5). There is also a saddle point that
both synchronous and de-synchronous volleys avoid, revolving around a=60 and
=1.2. It seems
38
Simulating Populations of Neurons
George Parish
2013
that if the volley has not reached these values after propagating through 2 or 3 layers then the signal
will become unstable and disperse.
Synchronous activity is capable of transmitting all kinds of information. From causing visual input to
correspond with visual cortical areas in the brain (Ayzenshtat, Meirovithz, Edelman, Werner-Reiss,
Bienenstock, & Abeles, 2010), to quick responses to visual attention via the movement of a joystick
(Shmiel, Drori, Shmiel, Ben-Shaul, Nadasdy, & Shemesh, 2006).
The purpose in this next simulation will be to reproduce a network of synchronous activity that
produces precise spike-timing, and also to confirm the conditions for the initial spike volley that are
required for synchronous spiking to occur through the network.
3.4.2 IMPLEMENTATION OF BACKGROUND NOISE
Figure 23 is a representation for how the poisson generator that generated noise was
attached to each alpha model neuron in every layer. Poisson generator p was connected to each
neuron (a1-a100). This was repeated for all layers 1-10 so that every neuron received the same
amount of noise. Previous research used conditions likely to be found in the cerebral cortex
(Diesmann, Gewaltig, & Aertsen, 1999). This simulation assumes that these conditions were such
that neurons were kept just below threshold, therefore the threshold rate will be calculated instead
to keep neurons in this ‘trigger happy’ state.
Figure 23 Network showing how noise was connected to neurons in each layer
The balanced excitation/inhibition network calculated the threshold rate to keep neurons trigger
happy. This was then multiplied by 1000 to fit that specific network. However, this value was found
to produce far too much noise when implemented into this particular network. Therefore
experimentation was done to determine what value (p) to multiply the threshold rate by in order to
produce just enough noise to keep the neurons just below threshold.
39
Simulating Populations of Neurons
George Parish
2013
Figure 24a-d show the effect of choosing successively higher values of p for one layer of 100
neurons. The population density was used as an analysis method as this effectively shows exactly
where in the scale between the reversal potential (0) and threshold (1) the population of neurons
were. Figure 24a takes p as 220 and we can see the population is kept below threshold. However,
the conditions in previous research were such that there was some sporadic firing of neurons due to
background noise. Therefore Figure 24b takes p as 235 in the hope of re-producing this effect. We
can see that the familiar triangle shape of the population is higher up the voltage scale, the median
value being around 0.85 as opposed to 0.75 in Figure 24a. This means that some neurons at the far
right edge of the population density triangle have reached threshold and fire. It is possible to see 3
such neurons recovering from refractory to re-join the other neurons.
Further experimentation in Figure 24c (p=240) shows that a slight increase in p causes the same
effect. The bulk density of neurons is now centred about 0.9 – virtually on the edge of threshold.
This causes several more neurons to fire. Figure 24d takes p as 260 and we can see about 1/3
neurons surpass threshold and fire due to this amount of noise. Judging from these population
density diagrams, taking p as 235 or 240 in Figure 24b & 23c would give just enough background
noise to keep the group of neurons just below threshold with some sporadic firing.
Figure 24 Experimenting with the value to multiply the threshold rate by (p)
40
Simulating Populations of Neurons
George Parish
2013
After establishing potential values for p, the next step is to construct a long simulation using these
values in consecutively connected groups of 100 neurons with no other input. This is important as
background noise will directly affect the population and skewer results when experimenting with the
strength of the initial spike volley. Figure 25a-b shows a long simulation for p=235 and p=240. It is
possible to instantly see that Figure 25a is our preferred choice. We can see that there are sporadic
spikes spread out amongst the layers. Figure 25b shows how the background noise builds up as it
gets propagated through each layer. This is apparent as there are many more spikes in the top right
quadrant of the diagram. Whilst this is far too much noise for our requirements, it is interesting how
seemingly random noise can be enough to trigger a volley in layers 8-10. This effect shall be
simulated and discussed in more detail later in this section.
MATLAB code used for the analysis of all the following networks can be found in Appendices C1-2.
Figure 25 Long simulation for p=235 & p=240
41
Simulating Populations of Neurons
George Parish
2013
3.4.3 IMPLEMENTATION OF NETWORK
Figure 26 shows how the network for this simulation was assembled. Neurons were
modelled using alpha model neurons supplied in NEST. Connections were modelled based on the
alpha function synapses as described in the balanced excitation/inhibition model. A peak-amplitude
of 0.1mV and a time-to-peak of 1.5ms were chosen for connections.
In Figure 26 we can see how layer a(1,n) connects via divergent connections to layer a(2,n), meaning
that ever y neuron in layer 2 receives excitatory input from each neuron in layer 1. This is repeated
through each layer, where each layer receives input from the layer preceding it and passes output to
the layer succeeding it. NEST code for this model can be seen in Appendix B:3.
Figure 26 Network for the stable propagation of synchronous spiking model
3.4.4 EVALUATION
Figure 27 shows the evolution of initial spike volleys as they get propagated through the
network. Volleys were started fully synchronised ( =0), somewhat synchronised ( =1.5) and less
synchronised ( =4). The effect of using different numbers of neurons in each volley can be seen
where lines start at different values of a. Each arrow indicates the number of neurons (a) and spread
( ) at each layer in the propagated volley. Blue arrows indicate a successfully propagated fully
synchronised initial volley. Green arrows indicate a successfully propagated volley with a low . Red
arrows indicate spike volleys that failed to cause synchronised spiking.
To create a fully synchronised initial volley of a spikes, a poisson generator was created at the
required rate for 0.1ms. To create a volley of =1.5 and =4, poisson generators were in action for
the first 5ms and 13ms respectively. This meant that the standard deviation of a spikes spaced
between 0 and t ms gave the underlying pulse density ( ) of the initial volley. To calculate the
42
Simulating Populations of Neurons
George Parish
required rate r of a poisson generator for a spikes lasting t ms:
2013
. A volley was
defined as any spikes occurring within 6ms of the time of the median spike in the layer. This design
decision was made so that noise wouldn’t have an impact in the calculation of a at each layer.
Similarly, a volley with less than 5 spikes was given an arbitrarily large
to signify that it wasn’t
strong enough to be considered a volley.
The results from Figure 27 are somewhat similar to that of previous research shown in Figure 28
(Diesmann, Gewaltig, & Aertsen, 1999), where synchronised volleys are attracted to a value of 99
spikes and about 0.5 . This indicates that the simulation was successful in defining a fully
synchronised volley, as previous research produced an attractor point of 90 spikes and 0.5 . The
fact there are more spikes in this simulation could be due to the fact that there was more
background noise used in previous research, causing some of the 100 neurons to spike at irregular
times and leaving them unable to synchronise
Previous research showed that there is a saddle point of about a=60 and
nd
=1.2 (figure 28), where if
rd
a volley has not reached these values by its 2 or 3 layer, it will likely disperse. Results from this
simulation show otherwise. When the initial volley contained 44 spikes it became very dispersed by
the 2nd layer (a=17, =3.4), following the pattern of other volleys that became unstable and
dispersed (initial a<40,
=0). Yet it provoked a volley in the next layer with a similar
50 spikes. This subsequently caused 99 neurons to fire in the next layer and
but containing
to gradually decrease
from 1.6 to the attractor point after two more layers.
Figure 27 Evolution of initial spike volleys, varying in number of neurons (a) with different synchronisations (sigma)
43
Simulating Populations of Neurons
George Parish
This appears to be a common feat, where a volley has an unchanging or deteriorating
2013
yet produces
many more spikes than the previous layer. This is likely because neurons are kept trigger happy and
will be caused to jump past threshold by just the first part of the incoming volley from the preceding
layer. This would mean that there would be many more neurons firing but as the incoming
low, the outgoing
could also be low. It seems that the fall in
may be
in subsequent layers is dependent
on the rise of a. This is supported by the fact that failed attempts (red arrows) fail to rise above
a=10.
Perhaps the most notable deviation from the original research is the case where an initial volley with
high
can become synchronous despite having less spikes than the lowest possible a of a fully
synchronised volley. This is possibly because the poisson generator produces noise for a longer
period of time as opposed to a short burst. This could result in the situation in Figure 25b where
noise can be propagated to cause a volley. This is unlikely, however, as by the 2nd layer this volley
has already achieved a lower
and higher a than a fully synchronised volley of 44 spikes. It is more
likely to be an error in the calculation of , which would question the results from all green arrows.
The following sections evaluate particularly interesting cases from Figure 27 by looking at the raster
plot, firing rate and evolution of the volley as a whole. The population densities of each layer and the
population as a whole at an interesting time t will also be considered. The case in which the volley
disperses is evaluated first so as to get an understanding as to what is required for successful
propagation. Three different cases of successful propagation will then be considered, where the
initial volley is synchronised and desynchronised, as well as a novel case of late synchronisation
caused by the propagation of background noise.
Figure 28 Evolution of initial spike volleys (Diesmann, Gewaltig, & Aertsen, 1999)
44
Simulating Populations of Neurons
George Parish
2013
3.4.5 CASE STUDY : DISPERSION OF THE PROPAGATING VOLLEY
Figure 29 shows the analysis of the dispersion of an initially synchronised volley containing 43 spikes.
The evolution of the synchronised spike volley diagram highlights which case from Figure 27 we are
studying. It shows how the first layer achieved a volley with
before completely dispersing. This
can be seen on the raster plot between 5-10ms, but this volley wasn’t strong enough to produce a
volley in the subsequent layer.
The raster plot also shows how noise gradually builds up in the simulation due to the initial spike
volley. This is confirmed when we see that there are less spikes in the simulation of just background
noise from Figure 25a when compared to the raster plot in Figure 29. The fact that there are more
spikes in the right half means that the level of background noise could have a larger effect if the
simulation ran for a longer time. The fact that there are more spikes in the upper right quadrant
than the upper left quadrant means that spikes are gradually pushed through each layer, perhaps
even some from the initial volley.
The firing rate diagram of Figure 29 supports how noise gets propagated through the network. There
is a brief pulse at the start where the initial volley causes a secondary volley. This is followed by a
steady stream of intermittent spikes as the first few layers start to push through a few spikes. After
50ms the population firing rate shows the level of activity increases as each layer starts to spike
more frequently.
The population density diagrams taken at t=80ms tell a similar story. In all layers, we can see the
population is steady just below threshold, making these diagrams very similar to the population
density diagrams depicting noise from Figure 24. This is because each layer hasn’t received enough
input for them to be pushed past threshold. The layer 1 density shows spikes from the secondary
volley recovering from refractory. From layer 2 onwards, we can see a gradual increase in spike
activity. As the general population of each consecutive layer makes a slight move towards threshold
(centre
0.875 in layer 2 to 0.925 in layer 10) we can see a few more spikes occurring in each layer.
These spikes then act as an input for the next layer which explains the populations slight more
towards threshold, causing more spikes.
The population density diagram of all layers in the bottom left of Figure 29 confirms how the
population as a whole is steady below threshold with a steady stream of neurons that have spiked
moving up the voltage scale to rejoin the main population. This diagram shows how there is no one
point in time where there were more spikes and re-iterates how the firing rate remains steady
throughout the simulation.
45
Figure 29 Case study : Dispersion of fully synchronised initial volley (a=43 =0)
46
3.4.6 CASE STUDY : SYNCHRONISED INITIAL VOLLEY CAUSING SYNCHRONISATION
Figure 30 shows the case where a fully synchronised volley of 44 spikes causes synchronisation. This
case is particularly interesting as the propagated volley initially follows the trend of desynchronisation, as described in the previous case, before becoming almost fully synchronised by
layer 4. This is highlighted by the evolution of the synchronous spike volley diagram, which also
shows exactly which case from Figure 27 we are discussing.
By examining the raster plot of Figure 30 we see how the initial volley caused a secondary volley in
layer 1 between 5-10ms, similar to our previous study of Figure 29. This suggests a fully synchronised
initial volley of any number of spikes will more quickly affect the subsequent layer. The next two
volleys in layer 2 & layer 3 show a small number of spikes spread over a long time frame. The way
the algorithm decides which spikes are part of the volley means that spikes after 25ms in layer 2 and
35ms in layer 3 are disregarded. The population density diagrams of layers 2 & 3 show how half the
neurons were recruited by the volley by showing a split in the voltages of the neurons at t=40ms.
Where half are still hovering at threshold and the other half are recovering from refractory.
When the volley reaches layer 4 it has recruited all of the neurons in that layer. This is shown by the
population density diagram of layer 4 where all the neurons have surpassed threshold and are
currently recovering from refractory at t=40ms. The raster plot confirms this as all of the spikes are
now focused in the vicinity of the volley rather than scattered throughout the simulation. It also
shows a rather large spread for the spikes in layer 4, supporting the 5th arrow in the evolution
diagram where 99 neurons are recruited with a spread of =1.6. The population density diagram of
the 5th layer is similar to that of the 4th layer. The key difference is that we can now see that the
underlying pulse density
is smaller as the group of neurons is spread over 0.3 of the voltage
scale in layer 5 as opposed to 0.45 in layer 4. The raster plot also shows this tightening in neuron
spike times at layer 5, and the evolution diagram stresses the change of
from 1.6 in layer 4 to 0.7
in layer 5. This defines how a volley synchronises after recruiting all neurons in a layer.
The key messages of Figure 30 are that whichever neurons are not recruited into the propagating
volley will intermittently spike throughout the simulation. This is shown best in the raster plot where
there is no noise after layer 5. Another key point is that the fewer spiking neurons in a volley and the
higher their spread, the longer it takes for the subsequent layer to react. This is supported by the
raster plot analysis, where the first 3 layers take 30ms to propagate the volley and the final 5 achieve
this in just 20ms. Perhaps the most important point of this simulation is that if the spread
reaches 3.5 and there is no increase in the volley’s number of spikes (a) the signal will likely disperse.
47
Figure 30 Case study : Synchronised initial volley (a=44 =0) causing synchronisation
48
3.4.7 CASE STUDY : DE-SYNCHRONISED INITIAL VOLLEY CAUSING SYNCHRONISATION
Figure 31 shows the case where a de-synchronised initial volley causes the propagating volley to
synchronise. This is interesting because it appears that a de-synchronised initial volley requires
fewer spikes than a synchronised initial volley to propagate the signal, a=36 is the cut-off point for
the former and a=44 for the latter. This could be because a de-synchronised initial volley goes on for
a longer time frame (5ms as opposed to 0.1ms). We already know that a population that is on the
verge of threshold only needs a small stimulus to be pushed over. Results here indicate that the first
part of a de-synchronised signal achieves this whilst the rest of the signal then propagates through
to encourage the just fired population to recover faster. This would give the volley more chance to
recover as a whole compared to if its only input was background noise. The next layer then receives
a more synchronised input as opposed to a string of intermittent spikes. The population density
diagrams of Figure 31 show exactly this when compared to those from Figure 30. We can see a
group of recovering neurons in the density diagrams of layers 1 & 2 that are clearly synchronised
together. When looking at the same groups of just fired neurons in layers 1 & 2 of Figure 30, we see
that they are spread over much of the voltage scale. This theory is also supported by the evolution
diagram of Figure 31, where layers 1, 2 & 3 revolve around a spread
of about 2.5. The same
spreads in Figure 30 start at 1.6 because of a fully synchronised input – but go up to 3.5 because of
the lack of a more sustained initial input.
What is also interesting about this simulation is that both cases of Figure 30 & Figure 31 achieve the
same level of synchronisation in layer 4 at t=40. Due to the de-synchronised initial input in Figure 31,
the population in layer 1 does not react until 15ms. This can be seen best on the firing rate diagram,
where the rate gradually increases from 15-35ms, at which point fully synchronised volleys start
occurring. In contrast, the firing rate of Figure 30 shows an initial reaction in layer 1 at 5-10ms and
no sustained activity until after 30ms, at which point volleys become synchronised. This highlights
how a more sustained initial volley containing fewer spikes can be just as effective as a stronger,
fully synchronised volley. This is also true for any volley during the simulation, where in Figure 30 we
can see a de-synchronised
=3.5) volley of just 17 spikes encourage a volley of 50 spikes in layer 3.
This leads us to conclude that synchronisation in Figure 30 occurs because of the sustained noise
generated from the initial synchronised volley rather than the volley itself. This could be because
connections between layers are weighted too strong, meaning that a few spikes can affect a large
population more easily.
49
Figure 31 Case study : De-synchronised initial volley (a=36 =4) causing synchronisation
50
3.4.8 CASE STUDY : SYNCHRONISED INITIAL VOLLEY CAUSING LATE SYNCHRONISATION
Figure 32 shows another interesting case that is not shown on the evolution diagram in Figure 27.
This case is the midpoint between the cases showcased by Figure 30 & Figure 29, where a
synchronised initial volley has some dispersion before propagating synchronised activity and
synchronised input completely disperses, respectively. This is because Figure 32 shows the case
where a fully synchronised initial volley
contains the equivalent of 43.5 spikes, as opposed
to 43 spikes in Figure 29 and 44 spikes in Figure 30.
The most interesting thing about this case is that completely de-synchronised noise can encourage
synchronised volleys. The raster plot of Figure 32 shows how the initial volley is completely
dispersed by the time synchronised activity occurs in layer 6. However, if we compare this raster plot
to that of Figure 25, it is clear that the initial volley caused just enough activity to propagate a
slightly higher amount of noise through the connected layers and cause synchronised volleys.
We can see how noise causes synchronisation by looking at the population density diagrams at t=87,
where the final layer 10 has just spiked. The centre of each population cluster in layers 1-5 moves
slightly higher on the voltage scale, meaning each layer gets closer to threshold. As described in how
noise gets propagated in Figure 29, this causes each layer to send an intermittent stream of an
increasing number of spikes (a). By layer 6, we can see that this value of a has increased to almost all
of the population for that layer (a 80 for layer 6 on evolution diagram). This emphasises the point
that a lengthy volley containing just a few spikes can eventually cause synchronous activity.
Supporting this is the evolution diagram of Figure 32, where volleys containing a small number of
spikes gradually encourage volleys of increasing numbers of spikes. Once enough neurons are
recruited by the propagated volley, the spread of firing times
reduces layer by layer as the final
step of stable and synchronised propagation. This is supported by the density diagrams where we
can see that in layers 1-5, a increases with barely any synchronisation. In layers 6-10 we begin to see
increasing once all neurons are firing.
The population density diagram for all layers in this network as a whole looks rather like that of
Figure 18, where 10000 neurons were fed continuous noise. Most neurons are on the edge of
threshold and there are often some in refractory that have just spiked. However, this density
diagram acts more like the balanced network density diagram of Figure 21, where packets of
neurons from each layer move together in pulses from refractory to threshold. This suggests that the
network implemented here would be fairly balanced if there were an infinite number of layers or if
the final layer fed back into the first layer.
51
Figure 32 Case study : Synchronised initial volley (a=43.5 =0) causing late synchronisation
52
4 CONCLUSION
This chapter concludes the project by evaluating whether the minimum requirements have been
met, if the research questions can be answered and what extra requirements have been
implemented. There is also a section detailing the limitations of the project and any future work.
4.1 PROJECT EVALUATION
This section deals with evaluating whether the project has been successful in the way it attempted
to answer the research questions posed and fulfil the minimum requirements.
4.1.1 AIM AND MINIMUM REQUIREMENTS
1
NEST simulations of individual alpha and delta integrate-and-fire neurons.
These simulations were developed as part of the Chapter 2: Literature Review. Their
intention was to give the writer a better understanding of how individual neurons operated
so that they could be explained throughout Chapter 2. Where necessary, voltage traces of
these simulations have been included, for example Figure 9 was one such simulation.
2
NEST simulations of 10000 delta integrate-and-fire neurons.
This simulation was achieved in section 3.2. As an extra, a diagram describing the layout of
the network was made (Figure 17) to more easily explain the dynamics of the network. This
simulation was effectively evaluated and provided insight into how background noise can
effect populations of neurons; this was useful going forward in the next simulations.
3
NEST simulation of synchronous spiking model.
The objective here was to cause and evaluate synchronous spiking through one NEST
simulation. To achieve this, a NEST framework for the simulation was achieved in section
3.4. This framework then allowed for a further 16 simulations with varying strength and
synchronicity in the initial pulse volley (Figure 27). This allowed for an analysis on a similar
level to the original paper (Diesmann, Gewaltig, & Aertsen, 1999).
4
MATLAB script for static and dynamic analysis of populations of neurons.
The aim here was to write script that would allow for the evaluation of population densities
for the aforementioned NEST simulations. This was achieved in the evaluation sections of
sections 3.2-3.4. Where the initial requirement was to do this analysis on the whole
population, it was also achieved for each layer in the synchronous spiking model. This
allowed for a more in-depth evaluation, giving different insights from the original research.
53
Simulating Populations of Neurons
George Parish
2013
4.1.2 EXCEEDING REQUIREMENTS
1. MATLAB script to plot population analyses for any NEST simulation
The MATLAB functions used are such that they can handle any data from a NEST simulation
in the form (voltmeterData, spikeData, layers) where ‘layers’ denotes how many layers were
simulated. It can then show the raster plot, firing rate, population densities of each layer as
well as the population density as a whole. It also shows the evolution of the spike volley,
which is specific to the Synchronous Spiking model. Figures 29-32 show screenshots of the
outputted MATLAB function and Figures 18 & 21 are edited screenshots of the function.
2. Analysis of initial volley of varying strength and synchronisation
This was made possible by creating a framework in NEST code that could be implemented as
many times as needed. The initial requirement was to cause synchronous spiking in a NEST
simulation and evaluate results. The additional requirement was being able to do this
several times to evaluate the effect of changing the strength of the initial volley.
3. Evaluation of balanced excitation/inhibition network from NEST examples
This was made possible by the MATLAB function described above, where data was simply
inputted into the function for the analysis seen in Figure 21. This added to the project as an
example of balanced background noise, a very useful simulation going forward.
4.1.3 RESEARCH QUESTIONS
The following questions were set to be answered upon completion of the project.
How do neurons react to the background noise of other spiking neurons in the brain?
This depends on the level of noise and the way it is connected to neurons.
Figure 18 shows noise that is connected by excitatory connections with no inhibition. This
causes the population to move towards threshold at a steady pace. Eventually we see
equilibrium where neurons are evenly spread between refractory and threshold and there is
a steady firing rate.
Figure 21 shows both excitatory and inhibitory noise making a balanced network. This is a
more likely case for neurons in the brain and we can see the effect on the population of
neurons in the analysis of Figure 21, where pulses of neurons move together from refractory
to threshold.
If we are to believe that neurons are connected via a series of layers defined in section 3.4,
then we would see most neurons balancing on the edge of threshold with pulses of neurons
moving from refractory to threshold much like in the balanced network.
54
Simulating Populations of Neurons
George Parish
2013
Under what conditions can a network of consecutively connected groups of neurons
transmit information?
We have found that information can be carried by precise spike timing – just as the original
research concluded (Diesmann, Gewaltig, & Aertsen, 1999). Figure 25 showed how if there
were too much background noise, then an incoming signal could be lost as the layers would
be inadvertently spiking due to this high level of background noise.
What input is required for information to be transmitted by consecutively connected
groups of neurons?
Within the confines of this simulation, a fully synchronised initial volley must be made up of
at least 44 spikes and a less synchronised volley can be made up of 36. This is contrary to
results published in the original research and is probably due to the different method of
creating noise used here or differing methods used to initiate the initial volley. Assuming
these simulations are correct, the interesting conclusion could be made that a desynchronised volley could sometimes be more effective than a synchronised volley due to
the length of time is was active for and the nature of layered networks.
Can the implementation of such a network explain real neuronal processes?
We could compare a fully synchronised strong initial volley to a strong suggestion (eg sight
of an incoming ball) which causes immediate synchronisation through the layers, leading to
an immediate reaction (eg movement). Another example could be a strong suggestion of a
sure smell, where the initial volley is strong and synchronised and immediate
synchronisation causes other layers responsible for memories or reactions to be activated.
A less strong, de-synchronised initial volley is a weaker suggestion (unsure smell/sight)
where less neurons firing at a larger spread (ms) don’t immediately cause recognition or
reaction. However, research here has shown that a de-synchronised initial volley can
synchronise after a longer amount of time (Figure 31). This could explain why it takes longer
to recognise unsure smells or sights. Research here also shows that even after complete desynchronisation layers will synchronise later on (Figure 32). This is reminiscent of times
when the answer to a question will pop into the mind (synchronise through relevant layers)
sometime after the initial thought (volley).
Can this implementation be applied in other areas of computer science?
The use of simulations such as this could be useful when studying the brain for medical
purposes. The role of synchronous activity in consciousness and brain disorders is becoming
an increasingly researched field. Computational learning mechanisms have always proven
useful when used in medical research, particularly so for the brain because of the vast
quantities of data to be sifted.
55
Simulating Populations of Neurons
George Parish
2013
4.2 CHALLENGES
Change of minimum requirements
There was a change of minimum requirements during the project where research using the
software MIIND was no longer necessary. This was due to the fact that it took longer than
foreseen to understand the nature of computational neuroscience and how to implement
models using the software NEST.
Implementation of background noise in final simulation
The original research used poisson generators to replicate the goings on in the cerebral
cortex. When these generators were implemented, noise became an issue and it seemed the
correct values were giving too much noise. This was why the firing rate was used from the
balanced network model. This is a probable cause for variation in results between this and
previous research.
Use of alpha models and alpha synapses in final simulation
An understanding of the alpha model neuron and alpha synapses wasn’t reached until the
implementation stages for the final simulation. As progress had already been made using the
alpha synapse described in the balanced network, rather than the alpha synapse described
in previous research, it was decided to continue forward with the former. This is also a
probable cause for differing results as spikes would affect neurons differently.
MATLAB script working in every case
The MATLAB script submitted is capable of simulating any voltmeter data and spike data in
any number of layers up to 10. There are previous versions that work better with single
layers, as the evolution of the spike volley is specific to layered synchronous spiking.
Skewed evaluation of synchronous spiking model
Due to the variations in the parameters of the simulation, the evaluation of the synchronous
spiking model may not be comparable to that of the original simulation. This would limit its
use in describing the actual goings on in the cerebral cortex, as the conditions used were
dissimilar to those found in this region of the brain. The way in which the initial volleys
spread (sigma) was calculated and initiated by a poisson generator could potentially be
incorrect, further misinforming the analysis.
56
Simulating Populations of Neurons
George Parish
2013
4.3 FUTURE WORK
Research into the role of synchronous activity in the cerebral cortex has seen recent attention in
neuroscience, with some theories pointing to it being responsible for directing conscious cognition.
Studies into the cause of brain disorders have often found that a lack of synchronous activity can be
the cause. The deregulation in the path of synchronous activity can lead to epileptic seizures
(Sabolek, et al., 2012), as paths change with each activation along the network, comparable to the
dispersion of synchronous activity at each layer seen in Figure 29. Additionally, neural synchrony is
abnormal in brain disorders such as schizophrenia and autism (Uhlhass, Pipa, & Singer, 2009), where
a state of consciousness is arguably flawed. This is evidence in itself that neural synchrony is
responsible for conscious cognition, supporting the previous discussion of how a strong or weak
suggestion can cause attention in various areas of the brain via synchronous activity.
57
BIBLIOGRAPHY
Section 2.1 took elements from the Computational Modelling of the Brain coursework submitted as
part of the Computational Modelling module (COMP5320M).
Section 2.3.1 & Section 2.3.2 were inspired by and borrowed definitions from the following:
Dayan, P. (2005). Theoretical Neuroscience. MIT Press.
Gazzaniga, M. (1998). Cognitive Neuroscience: The biology of the mind. New York:
Norton & Company.
Section 2.3.4 borrowed research from:
Abeles, M. (1991). Corticonics - Neural circuits of the cerebral cortex . Cambridge
University Press.
i
REFERENCES
Abeles, M. (1991). Corticonics - Neural circuits of the cerebral cortex . Cambridge University Press.
Amunts, K., Lepage, C., Borgeat, L., Mohlberg, H., Dickscheid, T., Rosseau, M., et al. (2013). BigBrain:
An Ultrahigh-Resolution 3D Human Brain Model. Science, 340(6139), 1472-1475.
Ayzenshtat, I., Meirovithz, E., Edelman, H., Werner-Reiss, U., Bienenstock, E., & Abeles, M. (2010).
Precise spatioremporal patterns among visual cortical areas and their relation to visual
stimulus processing. Journal of Neuroscience, 30, 11232-11245.
Balanced random network model. (n.d.). Retrieved from NEST Initiative: http://www.nestinitiative.org/index.php/The_balanced_random_network_model
Brodmann, K. (1909). On the Comparitive Localization of the Cortex.
Cajal, S. (1906). The Structure and Connexions of Neurons. Nobel Lecture, 220-253.
Dayan, P. (2005). Theoretical Neuroscience. MIT Press.
Diesmann, M., Gewaltig, M., & Aertsen, A. (1999). Stable propagation of synchronous spiking in
cortical neural networks. Nature, 402, 529-532.
Gazzaniga, M. (1998). Cognitive Neuroscience: The biology of the mind. New York: Norton &
Company.
Golgi, C. (1898). On the structure of nerve cells. Journal of Microscopy, 155.
Griffith, J. (1963). On the stability of brain-like structures. Biophysical Journal, 3, 299-308.
Hokfelt, T. (1984). Chemical anatomy of the brain. Science, 225(4668), 1326-1334.
http://catalog.nucleusinc.com. (n.d.). Retrieved from Nucleus Medical Art.
Max, C. (2004). Brain Imaging, Connectionionism, and Cognitive Neuropsychology. Cognitive
Neuroscience, 21-25.
Penfield, W., & Jasper, H. (1954). Epilepsy and the functional anatomy of the human brain. Boston:
Little and Brown.
Piccinini, G. (2007). Computational modelling vs. Computational explanation: Is everything a Turing
Machine, and does it matter to the philosophy of mind?. Australasian Journal of Philosophy,
93-115.
Sabolek, H., Waldemar, S., Lillis, K., Cash, S., Huberfeld, G., Zhao, G., et al. (2012). A canditate
mechanism underlying the variance of interictal spike propagation. Journal of Neuroscience,
32(9), 3009-3021.
ii
Simulating Populations of Neurons
George Parish
2013
Shmiel, T., Drori, R., Shmiel, O., Ben-Shaul, Y., Nadasdy, Z., & Shemesh, M. (2006). Temporally
precise cortical firing patterns are associated with distinct action segments. Journal of
Neurophysiology, 96, 2645-3652.
Uhlhass, P., Pipa, G., & Singer, W. (2009). Neural Synchrony in Cortical Networks: History, Concept
and Current Status. Frontiers in Integrative Neuroscience, 3(17).
Van Der Velde, F., & De Kamps, M. (2001). From knowing what to knowing where: Modeling objectbased attention with feedback disinhibition of activation. Journal of Cognitive Neuroscience.
iii
Simulating Populations of Neurons
George Parish
2013
APPENDIX A
PROJECT REFLECTION
The main challenge of this project was the amount of background research required to get an
understanding of computational neuroscience. Whilst the module Bio-Inspired Computing touched
on some of these areas, there wasn’t much background knowledge I could take into the project. This
was to be expected, however, and was one of the main reasons why I chose to do a project in this
field. The problem was overcome by keeping the topic in mind and doing some reading every day.
As the simulation software is used by a relatively small, experienced community, there isn’t a lot of
advice apart from basic tutorials. Whilst there is a very helpful online community, most questions I
had required an in-depth discussion to be able to understand the answer. I often had to wait until
the next supervisor meeting and ask a barrage of questions to better understand the topic at hand.
This was where frequent (bi-weekly) meetings with my supervisor, Dr. Marc De Kamps, were
incredibly useful and I usually came out of each meeting with a new understanding for the project.
These meetings were key to the completion of the project and I am grateful for them.
Because of my lack of understanding in the field, I felt there were moments when time was wasted
as I simply didn’t know where to look for help in some aspects of simulation. This caused moments
of desperation during the project, especially when trying to implement the final simulation in NEST.
In the end, the main problem was getting all the smalls things correctly implemented rather than
one big swooping change. As with any programming problem faced throughout the MSc course, all it
took was a bit of calm and patience to solve the big problems – so keeping a level head is advised!
I didn’t start research for the project until after the exam period. Whilst I do not believe it was
possible to start much earlier due to other work, it would have been easier going forward.
I have always found that writing the project report whilst reading and experimenting is the best
method. There is never a better time to write about something than when you are currently doing it,
as this is when you have the best understanding for it. This is useful as you accumulate a log of
simulations and accompanying notes as you go, making the final write up much easier and less
stressful and leaving you more time and energy to start fulfilling extra requirements.
I thoroughly enjoyed this project and I am extremely pleased I took the opportunity to study this
field whilst at Leeds University. I would advise future students to follow suit and choose a project
that interests them, no matter how hard it may seem.
iv
Simulating Populations of Neurons
George Parish
2013
APPENDICES B:1
INITIAL GANTT CHART
v
Simulating Populations of Neurons
George Parish
2013
APPENDICES B:2
FINAL GANTT CHART
vi
Simulating Populations of Neurons
George Parish
2013
APPENDICES C:1
MAIN FUNCTION FOR ANALYSING NEURONAL DATA
function [ PFRAll, s_in, a_in] = populationDensity( voltmeterData, spikeData, layers, inputSpikes,
inputSpreadms)
% Plots PFR, raster plot and evolution of spike volley and simulates population density. Takes
voltmeter data, spike data, number of layers, initial volley strength (a) and spread (ms)
% pre-process data
voltmeterVolts = voltmeterData(:,3);
reversalPotential = min(voltmeterVolts);
threshold = max(voltmeterVolts);
% normalise voltage
voltmeterVolts = (voltmeterVolts-reversalPotential)./(threshold-reversalPotential);
voltmeterTime = voltmeterData(:,2);
spikeDataID = spikeData(:,1);
spikeDataTimes = spikeData(:,2);
c = unique(voltmeterTime); % max(c) = length of simulation
n = unique(voltmeterData(:,1)); % max(n) = number of neurons
perLayer = round(max(n)/layers);
layers = round(max(n)/perLayer); % number of layers
stepms = 0.5; % step size for pop firing rate
voltmeterLayered = cell(layers,1);
spikesLayered = cell(layers,1);
lsC = [];
lold = [];
binSize = 100; % bin size of pop density histograms
v = [];
speed = 2; % speed of simulation
p = speed/100;
time = linspace(0,1,100);
hists = cell(max(c),layers);
histCentres = cell(max(c),layers);
top=0;
PFRAll = cell(layers,1);
% seperate voltmeter data by layers and calculate histogram data
for i = 1:max(c)
if(layers>1)
vii
Simulating Populations of Neurons
George Parish
2013
for j=1:layers
i2 = layers*perLayer*(i-1);
j2 = (j-1)*perLayer;
takeFrom = i2+j2+1;
takeTo = i2+j*perLayer;
voltmeterLayered{j} = [voltmeterLayered{j}; voltmeterVolts(takeFrom:takeTo)
ones(perLayer,1)*i];
[f, x] = hist(voltmeterVolts(takeFrom:takeTo),binSize);
hists{i,j} = f;
histCentres{i,j} = x;
end
else
v = voltmeterVolts(voltmeterVolts(voltmeterTime==i));
[f, x] = hist(v,binSize);
hists{i,1} = f;
histCentres{i,1} = x;
end
end
% set figure size and position
hFig = figure(1);
set(hFig, 'Position', [1450 600 1800 900])
if(layers>3)
gridy=layers;
if(layers>5)
gridy=5;
end
else
gridy=3;
end
if(layers>5)
gridx=3;
else
gridx=2;
end
% seperate spike data by layers
for i = 1:layers
ls = spikeDataID(spikeDataID<(perLayer*i)+1);
if(i>1)
lsC(i) = length(ls(:,1))-sum(lsC);
ls = ls(sum(lsC)-lsC(i)+1:sum(lsC));
ls(:,2) = spikeDataTimes(sum(lsC)-lsC(i)+1:sum(lsC));
viii
Simulating Populations of Neurons
George Parish
2013
else
lsC(i) = length(ls(:,1));
ls(:,2) = spikeDataTimes(1:length(ls(:,1)));
end
spikesLayered{i} = ls;
end
% calculate population firing rates
for j = 1:layers
e = [];
PFR = [];
layerSpikes = spikesLayered{j};
for i=1:(max(c)+1)/stepms
spikes = layerSpikes(layerSpikes(:,2)<i*stepms+0.1);
lnew = length(spikes);
if(i>1)
spikes = spikes(lold+1:end);
end
lold = lnew;
PFR = [PFR; i*stepms length(spikes)/max(n)/(stepms/1000)];
end
for i = 1:length(PFR)
e = [e; sqrt(PFR(i,2))];
end
PFRAll{j} = PFR;
% Plot Firing Rate
[xData, yData] = prepareCurveData( PFR(:,1), PFR(:,2) );
% Set up fittype and options.
ft = fittype( 'pchipinterp' );
opts = fitoptions( ft );
opts.Normalize = 'on';
% Fit model to data.
[fitresult] = fit( xData, yData, ft, opts );
subplot(gridy,gridx,gridx+1)
plot( fitresult, xData, yData );
hold all
legend off
%errorbar(PFR(:,1),PFR(:,2),e)
xlim([0 max(c)+1])
if(max(PFR(:,2))+sqrt(max(PFR(:,2)))>top)
top = max(PFR(:,2))+sqrt(max(PFR(:,2)));
ix
Simulating Populations of Neurons
George Parish
2013
end
ylim([0 top])
title('Population Firing Rate')
xlabel('Time')
ylabel('Frequency (Hz)')
end
% Plot Raster Plot
subplot(gridy,gridx,1)
scatter(spikeData(:,2),spikeData(:,1),'b.');
xlim([0 max(c)+1])
title('Raster Plot')
xlabel('Time')
ylabel('Neuron ID')
% Plot evolution
[s_in, a_in] = evolution(spikeData,inputSpikes,inputSpreadms);
subplot(gridy,gridx,10)
plot(s_in,a_in,'g<')
hold on
plot(s_in,a_in,'g')
ylim([0 100])
xlim([0 4])
title('Evolution of synchronous spike volley')
xlabel('sigma (ms)')
ylabel('a (spikes)')
% main loop
for t=1:length(c)-1
for i = 1:layers
% plot time
subplot(gridy,gridx,gridx*2+1)
plot(t,time,'r')
xlim([0 length(c)])
ylim([0 1])
xlabel('Time')
% Plot Population Density for each layer
if(i<6)
subplot(gridy,gridx,i*gridx-(gridx-2))
else
subplot(gridy,gridx,(i-5)*gridx)
x
Simulating Populations of Neurons
George Parish
2013
end
bar(x,f/trapz(f)*10);
bar(histCentres{t,i},hists{t,i}/trapz(hists{t,i})*10);
ylim([0 1])
xlim([0 1])
title('Population Density')
xlabel('Voltage')
ylabel('Frequency')
% Plot Population Density for all layers
if(layers>1)
if(layers==2)
subplot(gridy,gridx,6)
else
subplot(gridy,gridx,gridx*gridy-2)
end
[fAll, xAll] = hist(voltmeterVolts((t-1)*perLayer*layers+1:t*perLayer*layers),binSize);
bar(xAll,fAll/trapz(fAll)*10);
ylim([0 1])
xlim([0 1])
title('Population Density All Layers')
xlabel('Voltage')
ylabel('Frequency')
end
end
% pause and step through option
pause(p);
%k = waitforbuttonpress;
if(p == 0)
display(t)
end
end
end
xi
Simulating Populations of Neurons
George Parish
2013
APPENDICES C:2
FUNCTION TO PLOT EVOLUTION OF SYNCHRONOUS SPIKING
function [ s_in, a_in ] = evolution( spikeData, inputSpikes, inputSpread )
%S Summary of this function goes here
% Calculates evolution of spike volley. Takes spike data, initial volley strength(a) and
spread(ms)
a_out = [];
a_in = [];
s_in = [];
s_out = [];
for i = 1:10
s=[];
% Split layers
for j = 1:length(spikeData)
if(spikeData(j,1)>(i-1)*100)
if(spikeData(j,1)<i*100)
s = [s;spikeData(j,:)];
end
end
end
% Exclude spikes not in volley
m = median(s(:,2));
for j=1:length(s)
if(s(j,2)>m+6)
s(j,2)=0;
end
if(s(j,2)<m-6)
s(j,2)=0;
end
end
s=s(s(:,2)>0,:);
a_out(i,1) = length(s);
if(length(s)<10)
if(i>1)
s_out(i,1) = 10;
else
s_out(i,1) = std(s(:,2));
end
else
s_out(i,1) = std(s(:,2));
xii
Simulating Populations of Neurons
George Parish
2013
end
if(i>1)
a_in(i,1) = a_out(i-1);
s_in(i,1) = s_out(i-1);
else
a_in(i,1) = inputSpikes;
s_in(i,1) = std(linspace(0,inputSpread,inputSpikes));
end
end
end
xiii
Simulating Populations of Neurons
George Parish
2013
APPENDIX C:3
NEST CODE FOR SYNCHRONOUS SPIKING MODEL
import nest
import nest.raster_plot
import numpy
from numpy import exp
import time
def LambertWm1(x):
nest.sli_push(x); nest.sli_run('LambertWm1'); y=nest.sli_pop()
return y
# Computer Post-synaptic potential
def ComputePSPnorm(tauMem, CMem, tauSyn):
"""Compute the maximum of postsynaptic potential
for a synaptic input current of unit amplitude
(1 pA)"""
a = (tauMem / tauSyn)
b = (1.0 / tauSyn - 1.0 / tauMem)
t_max = 1.0/b * ( -LambertWm1(-exp(-1.0/a)/a) - 1.0/a )
return exp(1.0)/(tauSyn*CMem*b) * ((exp(-t_max/tauMem) - exp(t_max/tauSyn)) / b - t_max*exp(-t_max/tauSyn))
nest.ResetKernel()
nest.SetKernelStatus({"overwrite_files": True})
startbuild= time.time()
# Declarations
dt
= 0.1
simtime = 80.0
delay
= 1.5
g
= 1.0
eta
= 1.0
epsilon = 0.1
order
NE
N_neurons
N_rec
=
=
=
=
50
2*order
NE
NE
CE
= epsilon*NE
C_tot = int(CE)
tauSyn
CMem =
tauMem
theta
J
= 1.5
240.0
= 20.0
= -55.0
= 0.1
xiv
Simulating Populations of Neurons
George Parish
2013
# Calculate weight of connections
J_ex = J / ComputePSPnorm(tauMem, CMem, tauSyn)
J_in = -g*J_ex
# Calculate rate of background noise to keep neurons just below
threshold
nu_th = (-theta * CMem) / (J_ex*CE*numpy.exp(1)*tauMem*tauSyn)
nu_ex = eta*nu_th
p_rate = 235.0*nu_ex*CE
nest.SetKernelStatus({"resolution": dt, "print_time": True})
print "Building network"
# Parameters for Alpha model neuron
neuron_params= {"C_m"
: CMem,
"tau_m"
: tauMem,
"tau_syn_ex": tauSyn,
"tau_syn_in": tauSyn,
"t_ref"
: 1.0,
"E_L"
: -70.0,
"V_reset"
: -70.0,
"V_m"
: -58.0,
"V_th"
: theta}
# Create layers of Neurons
nest.SetDefaults("iaf_psc_alpha", neuron_params)
layer1=nest.Create("iaf_psc_alpha",NE)
layer2=nest.Create("iaf_psc_alpha",NE)
layer3=nest.Create("iaf_psc_alpha",NE)
layer4=nest.Create("iaf_psc_alpha",NE)
layer5=nest.Create("iaf_psc_alpha",NE)
layer6=nest.Create("iaf_psc_alpha",NE)
layer7=nest.Create("iaf_psc_alpha",NE)
layer8=nest.Create("iaf_psc_alpha",NE)
layer9=nest.Create("iaf_psc_alpha",NE)
layer10=nest.Create("iaf_psc_alpha",NE)
# Create and connect spike detector and voltmeter
spikesAll=nest.Create("spike_detector")
nest.SetStatus(spikesAll,[{"label": "spikesAll", "withtime": True,
"withgid": True, "to_file":True}])
voltmeterAll = nest.Create("voltmeter")
nest.SetStatus(voltmeterAll,[{"label": "voltmeterAll", "withtime":
True, "withgid": True, "to_file":True}])
nest.DivergentConnect(voltmeterAll,layer1)
nest.ConvergentConnect(layer1,spikesAll)
nest.DivergentConnect(voltmeterAll,layer2)
nest.ConvergentConnect(layer2,spikesAll)
nest.DivergentConnect(voltmeterAll,layer3)
nest.ConvergentConnect(layer3,spikesAll)
nest.DivergentConnect(voltmeterAll,layer4)
xv
Simulating Populations of Neurons
George Parish
2013
nest.ConvergentConnect(layer4,spikesAll)
nest.DivergentConnect(voltmeterAll,layer5)
nest.ConvergentConnect(layer5,spikesAll)
nest.DivergentConnect(voltmeterAll,layer6)
nest.ConvergentConnect(layer6,spikesAll)
nest.DivergentConnect(voltmeterAll,layer7)
nest.ConvergentConnect(layer7,spikesAll)
nest.DivergentConnect(voltmeterAll,layer8)
nest.ConvergentConnect(layer8,spikesAll)
nest.DivergentConnect(voltmeterAll,layer9)
nest.ConvergentConnect(layer9,spikesAll)
nest.DivergentConnect(voltmeterAll,layer10)
nest.ConvergentConnect(layer10,spikesAll)
# Define excitatory and inhibitory connections
nest.CopyModel("static_synapse","excitatory",{"weight":J_ex,
"delay":delay})
nest.CopyModel("static_synapse","inhibitory",{"weight":J_in,
"delay":delay})
# Create cerebral cortex noise
nest.SetDefaults("poisson_generator",{"rate":p_rate})
noiseE=nest.Create("poisson_generator",1)
#noiseI=nest.Create("poisson_generator",12)
#nest.SetStatus(noiseE,{"rate": 1760.0})
#nest.SetStatus(noiseI,{"rate": 250.0})
# Initial Pulse Volley
volley = nest.Create("poisson_generator",1,{"rate":2800.0,
"start":0.0,"stop":13.0})
nest.DivergentConnect(volley,layer1, model="excitatory")
print "Connecting devices."
# Connect background noise to each layer
nest.ConvergentConnect(noiseE, layer1, model="excitatory")
#nest.ConvergentConnect(noiseI, layer1, model="inhibitory")
nest.ConvergentConnect(noiseE, layer2, model="excitatory")
#nest.ConvergentConnect(noiseI, layer2, model="inhibitory")
nest.ConvergentConnect(noiseE, layer3, model="excitatory")
#nest.ConvergentConnect(noiseI, layer3, model="inhibitory")
nest.ConvergentConnect(noiseE, layer4, model="excitatory")
#nest.ConvergentConnect(noiseI, layer4, model="inhibitory")
nest.ConvergentConnect(noiseE, layer5, model="excitatory")
#nest.ConvergentConnect(noiseI, layer5, model="inhibitory")
nest.ConvergentConnect(noiseE, layer6, model="excitatory")
#nest.ConvergentConnect(noiseI, layer6, model="inhibitory")
nest.ConvergentConnect(noiseE, layer7, model="excitatory")
xvi
Simulating Populations of Neurons
George Parish
2013
#nest.ConvergentConnect(noiseI, layer7, model="inhibitory")
nest.ConvergentConnect(noiseE, layer8, model="excitatory")
#nest.ConvergentConnect(noiseI, layer8, model="inhibitory")
nest.ConvergentConnect(noiseE, layer9, model="excitatory")
#nest.ConvergentConnect(noiseI, layer9, model="inhibitory")
nest.ConvergentConnect(noiseE, layer10, model="excitatory")
#nest.ConvergentConnect(noiseI, layer10, model="inhibitory")
# Connect layers of neurons together
nest.ConvergentConnect(layer1, layer2,
nest.ConvergentConnect(layer2, layer3,
nest.ConvergentConnect(layer3, layer4,
nest.ConvergentConnect(layer4, layer5,
nest.ConvergentConnect(layer5,
nest.ConvergentConnect(layer6,
nest.ConvergentConnect(layer7,
nest.ConvergentConnect(layer8,
nest.ConvergentConnect(layer9,
print "Connecting network."
model="excitatory")
model="excitatory")
model="excitatory")
model="excitatory")
layer6, model="excitatory")
layer7, model="excitatory")
layer8, model="excitatory")
layer9, model="excitatory")
layer10, model="excitatory")
endbuild=time.time()
print "Simulating."
nest.Simulate(simtime)
endsimulate= time.time()
events_ex = nest.GetStatus(spikesAll,"n_events")[0]
rate_ex
= events_ex/simtime*1000.0/N_rec
build_time = endbuild-startbuild
sim_time
= endsimulate-endbuild
print
print
print
print
print
print
"Brunel network simulation (Python)"
"Number of neurons :", N_neurons
"
Exitatory :", int(CE*N_neurons)+N_neurons
"Excitatory rate
: %.2f Hz" % rate_ex
"Building time
: %.2f s" % build_time
"Simulation time
: %.2f s" % sim_time
nest.raster_plot.from_device(spikesAll, hist=True)
nest.raster_plot.show()
xvii