* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Restriction Mapping Restriction Fragment Length Polymorphism
Genetic code wikipedia , lookup
Holliday junction wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
DNA barcoding wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Human genome wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
DNA profiling wikipedia , lookup
DNA sequencing wikipedia , lookup
DNA polymerase wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Microevolution wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
DNA vaccination wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Metagenomics wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Primary transcript wikipedia , lookup
Point mutation wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Non-coding DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
Molecular cloning wikipedia , lookup
Genomic library wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome editing wikipedia , lookup
Epigenomics wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
SNP genotyping wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Helitron (biology) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
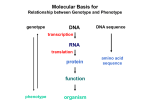
Molecular Basis for Relationship between Genotype and Phenotype genotype DNA DNA sequence transcription RNA translation protein function phenotype organism amino acid sequence Restriction Mapping DNA is restriction digested with restriction enzymes, individually (single-enzyme digest) and in combination (double digest). The restriction fragments are subjected to electrophoresis. The fragments are identified, either using UV absorbing dye or labeled probe. Double digest determines if fragment produced by one enzyme contains restriction sites for the other enzyme. Fragments are aligned by size. Enzyme 1: 8 kb, 6 kb, 3 kb or 3 kb, 6 kb, 8 kb 6 kb, 8 kb, 3 kb or 3 kb, 8 kb, 6 kb 8 kb, 3 kb, 6 kb or 6 kb, 3 kb, 8 kb Enzyme 2: 10 kb, 7 kb or 7 kb, 10 kb Double Digest: 3 kb fragment is split into 2 kb and 1 kb fragments. Restriction Fragment Length Polymorphism (RFLP) Individuals can be identified according to RFLP genotype. RFLP locus could be linked to a gene, and thus be used as a diagnostic marker. Use of Restriction Fragment Length Polymorphism I. Marker locus II. Diagnostic A. Medicine B. Forensics III. Assessment of Genetic Variation A. Within and between populations B. Within and between species Restriction Mapping versus RFLP Mapping I. Restriction Mapping A. Based on physical analysis of DNA B. Based on restriction sites with no variation C. Mostly short-range (fine-scale) maps II. RFLP Mapping A. Based on recombination analysis of matings B. Based on restriction-site variation between homologous chromosomes C. Mostly longe-range (coarse-scale) maps Other Useful Approaches 1. Single Nucleotide Polymorphisms (SNPs) Individuals differ in single nucleotides (every 11 to 300 bp in interval). 2. Simple-Sequence Length Polymorphisms (SSLPs) Very short repetitive DNA sequences are more polymorphic than RFLP sequences. These are also called Variable Number Tandem Repeats (VNTRs) - Minisatellite Markers - Microsatellite Markers Simple-Sequence Length Polymorphisms 1. Minisatellite DNA These are 1 to 5 kb in length consisting of repeats 15 to 100 nucleotides in length and are identified by Southern analysis. 2. Microsatellite DNA These are tandem repeats of dinucleotides, commonly stretches of CA. 5’ C A C A C A C A C A C A C A 3’ 3’ G T G T G T G T G T G T G T 5’ These are identified by gel electrophoresis of PCR products. Refer to Figure 4-15, Griffiths et al., 2015. Molecular Basis for Relationship between Genotype and Phenotype genotype DNA DNA sequence transcription RNA translation protein function phenotype organism amino acid sequence Dideoxy DNA Sequencing Chain-terminating (dideoxy) nucleotide Use of dideoxy nucleotide in primer extension reaction will randomly arrest DNA synthesis. Dideoxy DNA Sequencing 4 different reactions are conducted, each with a different type of dideoxy nucleotide. Fragments are separated by electrophoresis. Banding pattern is used to infer the base sequence of the original template strand. Migration 3’ Sequencing Gel Using Radioactive Primer 5’ Remember… This is base sequence of synthesized strand. Reading the DNA sequence from an automatic sequencer Oligonucleotide primers can be tagged with fluorescent dyes instead of radioactive labels. A different colored dye can be used for each of the four reactions. In Search of Potential Genes Open reading frames (ORFs) are long stretches of DNA that start with ATG and end with a stop codon. A doublestranded DNA molecule has 6 possible reading frames, 3 for each strand.