* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Human genome wikipedia , lookup
DNA profiling wikipedia , lookup
DNA sequencing wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
DNA polymerase wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
Metagenomics wikipedia , lookup
Genealogical DNA test wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Primary transcript wikipedia , lookup
Non-coding DNA wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
DNA vaccination wikipedia , lookup
SNP genotyping wikipedia , lookup
DNA supercoil wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Point mutation wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Epigenomics wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Molecular cloning wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome editing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Helitron (biology) wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
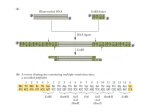
Zoo/Bot 3333 Genetics 4/22/05 Quiz 4 For answers to the quiz, click here: 1. The restriction enzyme, AvaI has the six base recognition sequence CPyCGPuG, where Py stands for a pyrimidine, and Pu stands for a purine). On average, this enzyme might be expected to cut a random DNA sample every ____________bp. a) 64; b) 256; c) 1024; d) 1296; e) 4096. 2. The recognition sites for the restriction enzymes BamHI, XbaI and BglII are 5’-GGATCC-3’, 5’-TCTAGA-3’ and 5’-AGATCT-3’, respectively, where the arrows represent the cut locations for each strand of the palindromic site. Which enzymes leave compatible ends that will facilitate ligation? a) All of these enzymes leave ends that are compatible with ends generated by the others; b) None of the enzymes produce compatible ends; c) Only BamHI and BglII fragments are compatible; d) Only BamHI and XbaI fragments are compatible; e) only BglII and XbaI fragments are compatible. 3. True or false. One useful property of plasmid vectors used in molecular cloning is their ability to integrate into the E. coli chromosome. Questions 4-5 pertain to the following. DNA was prepared from white blood cells isolated from a large human population. These DNAs were individually digested with EcoRI, subjected to electrophoresis and Southern blotting, and the blot was probed with a radioactively labeled cloned DNA sequence. Ten * samples. The figure below indicates the different different restriction patterns were observed among these patterns observed in human populations. 4. In the following diagrams, the vertical lines represent EcoRI restriction sites. An asterisk over the site represents a polymorphism (presence or absence of the site in individuals) in the population. The double arrow represents the boundaries of the cloned DNA used in the Southern blot analysis. Which of the following is consistent with the arrangement of EcoRI restriction sites in human genomic DNA? * a) 1.0 * 2.1 1.0 1.9 d) c) 1.0 4.0 1.0 1.9 1.9 2.1 * * 1.0 * e) * * b) 2.1 1.9 2.1 5. Suppose individuals 1 and 4 married. Assuming no recombination occurs within the region in question, how many potentially different patterns would be observed among their progeny on Southern blot analysis? a) only 1; b) 2; c) 3; d) 4 e) 6. Question 6 pertains to the following. This region of the genome is known to contain a particular gene, which encodes a very large protein of 1600 amino acids. A cDNA library primed with oligo dT was made and a clone derived from that library hybridized to the 5 kb and 3.1 kb restriction fragments only. When sequenced, this clone was 720 nucleotides in length. A synthetic oligonucleotide that corresponded to amino acids 3 through 11 of this protein was produced and labeled; it hybridized to the 5 kb, 4 kb and 1.9 kb fragments. 6. True or false. The 3’ end of the mRNA made from this region would be located in the 1 kb restriction fragment. 7. True or false. It would be impossible to produce a cDNA library of genes expressed in human red blood cells, since red blood cells do not contain a nucleus. Questions 8-9 pertain to the following diagram, which represents a region of a DNA sequencing gel produced using the Sanger chain termination method. The individual lanes represent the reaction products from the four separate labeling reactions (A,G,C,T) which were loaded at the top of the gel. 8. As deduced from this experiment, the sequence A G C T of the strand being copied (template) in this reaction is: a) 5’-AGACTATCTCTA-3’ b) 5’-ATCTCTATCAGA-3 c) 5 -TAGAGATAGTCT-3’ d) 5- TCTGATAGAGAT-3’ e) 5- GAGTCGCTCTCG-3’ 9. Which of the following was NOT present in the test tube during the ‘A’ DNA synthesis reaction? a) dATP; b) dGTP; c) ddATP; d) dUTP; e) all of the above were present in the reaction. Question 10 pertains to the following. Six independently derived mutants are recovered in Neurospora which are all unable to grow unless supplemented with compound Z. The mutants are then grown on minimal media supplemented with one of 6 chemicals all believed to be precursors to compound Z. A summary of the ability of the mutants to grow on media containing these chemicals is indicated below, where a “+” sign indicates growth and a “-” sign indicates no growth. mutant number precursors added to minimal medium A B C D E F Z 1 + + + - - - + 2 + + + + - - + 3 + + + + + - + 4 - - + - - - + 5 - - - - - - + 6 - + + - - - + 10. From the above chart, the enzyme that catalyzes the production of compound D appears to be mutant in strain: a) 1; b) 2; c) 3; d) 4; e) none of the above.