* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Personal genomics as a major focus of CSAIL research
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Transposable element wikipedia , lookup
Metagenomics wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Gene therapy wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Copy-number variation wikipedia , lookup
Oncogenomics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Ridge (biology) wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
History of genetic engineering wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenomics wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Minimal genome wikipedia , lookup
Gene expression profiling wikipedia , lookup
Non-coding DNA wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Gene desert wikipedia , lookup
Pathogenomics wikipedia , lookup
Human genetic variation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Human genome wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Helitron (biology) wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Designer baby wikipedia , lookup
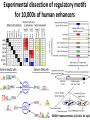
Computational personal genomics: selection, regulation, epigenomics, disease Manolis Kellis Broad Institute of MIT and Harvard MIT Computer Science & Artificial Intelligence Laboratory Understanding human variation and human disease Gene annotation (Coding, 5’/3’UTR, RNAs) Evolutionary signatures Roles in gene/chromatin regulation Activator/repressor signatures CATGACTG CATGCCTG Non-coding annotation Chromatin signatures Disease-associated variant (SNP/CNV/…) Other evidence of function Signatures of selection (sp/pop) • Challenge: from loci to mechanism, pathways, drug targets Goal: A systems-level understanding of genomes and gene regulation: • The regulators: Transcription factors, microRNAs, sequence specificities • The regions: enhancers, promoters, and their tissue-specificity • The targets: TFstargets, regulatorsenhancers, enhancersgenes • The grammars: Interplay of multiple TFs prediction of gene expression The parts list = Building blocks of gene regulatory networks Compare 29 mammals: Reveal constrained positions NRSF motif • Reveal individual transcription factor binding sites • Within motif instances reveal position-specific bias • More species: motif consensus directly revealed Chromatin state dynamics across nine cell types Predicted linking Correlated activity • Single annotation track for each cell type • Summarize cell-type activity at a glance • Can study 9-cell activity pattern across Revisiting diseaseassociated variants xx • Disease-associated SNPs enriched for enhancers in relevant cell types • E.g. lupus SNP in GM enhancer disrupts Ets1 predicted activator HaploReg: Automate search for any disease study (compbio.mit.edu/HaploReg) • Start with any list of SNPs or select a GWA study – Mine publically available ENCODE data for significant hits – Hundreds of assays, dozens of cells, conservation, motifs – Report significant overlaps and link to info/browser Experimental dissection of regulatory motifs for 10,000s of human enhancers 54000+ measurements (x2 cells, 2x repl) Example activator: conserved HNF4 motif match WT expression specific to HepG2 Motif match disruptions reduce expression to background Non-disruptive changes maintain expression Random changes depend on effect to motif match Allele-specific chromatin marks: cis-vs-trans effects • Maternal and paternal GM12878 genomes sequenced • Map reads to phased genome, handle SNPs indels • Correlate activity changes with sequence differences Brain methylation in 750 Alzheimer patients/controls 750 individuals 500,000 methylation probes Phil de Jager, Roadmap disease epigenomics Epigenome Phenotype Genome meQTL 1 Brad Bernstein REMC mapping 2 Epigenome Classification MWAS • 10+ years of cognitive evaluations, post-mortem brains • 93% of functional epigenomic variation is genotype driven! • Global repression in 7,000 enhancers, brain-specific targets Global hyper-methylation in 1000s of AD-associated loci P-value Top 7000 probes Methylation 480,000 probes, ranked by Alzheimer’s association Alzheimer’s-associated probes are hypermethylated • Global effect across 1000s of probes – – – – Rank all probes by Alzheimer’s association 7000 probes increase methylation (repressed) Enriched in brain-specific enhancers Near motifs of brain-specific regulators Complex disease: genome-wide effects Covers computational challenges associated with personal genomics: - genotype phasing and haplotype reconstruction resolve mom/dad chromosomes - exploiting linkage for variant imputation co-inheritance patterns in human population - ancestry painting for admixed genomes result of human migration patterns - predicting likely causal variants using functional genomics from regions to mechanism - comparative genomics annotation of coding/non-coding elements gene regulation - relating regulatory variation to gene expression or chromatin quantitative trait loci - measuring recent evolution and human selection selective pressure shaped our genome - using systems/network information to decipher weak contributions combinatorics - challenge of complex multi-genic traits: height, diabetes, Alzheimer's 1000s of genes Family Inheritance Personal genomics today: 23 and We Recombination breakpoints Me vs. my brother My dad Mom’s dad Disease risk Human ancestry Dad’s mom Genomics: Regions mechanisms drugs Systems: genes combinations pathways Personal genomics tomorrow: Already 100,000s of complete genomes • Health, disease, quantitative traits: – Genomics regions disease mechanism, drug targets – Protein-coding cracking regulatory code, variation – Single genes systems, gene interactions, pathways • Human ancestry: – Resolve all of human ancestral relationships – Complete history of all migrations, selective events – Resolve common inheritance vs. trait association • What’s missing is the computation – – – – – New algorithms, machine learning, dimensionality reduction Individualized treatment from 1000s genes, genome Understand missing heritability Reveal co-evolution between genes/elements Correct for modulating effects in GWAS Collaborators and Acknowledgements • Chromatin state dynamics – Brad Bernstein, ENCODE consortium • Methylation in Alzheimer’s disease – Phil de Jager, Brad Bernstein, Epigenome Roadmap • Mammalian comparative genomics – Kerstin Lindblad-Toh, Eric Lander, 29 mammals consortium • Massively parallel enhancer reporter assays – Tarjei Mikkelsen, Broad Institute • Funding – NHGRI, NIH, NSF Sloan Foundation MIT Computational Biology group Compbio.mit.edu Mike Lin Ben Holmes Angela Yen Matt Eaton Soheil Feizi Luke Bob Ward Altshuler Stefan Washietl Pouya Kheradpour Manolis Kellis Jason Jessica Ernst Wu Irwin Daniel Jungreis Marbach Louisa DiStefano Sushmita Roy Stata3 Stata4 Chris Bristow Mukul Bansal Rachel Sealfon Dave Hendrix Loyal Goff Human constraint outside conserved regions Active regions Average diversity (heterozygosity) Aggregate over the genome Ward and Kellis, Science 2012 • Non-conserved regions: • Conserved regions: – ENCODE-active regions show reduced diversity Lineage-specific constraint in biochemically-active regions – Non-ENCODE regions show increased diversity Loss of constraint in human when biochemically-inactive