* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Post-transcriptional modifications Cap a
Zinc finger nuclease wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Non-coding DNA wikipedia , lookup
Human genome wikipedia , lookup
Point mutation wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Gene nomenclature wikipedia , lookup
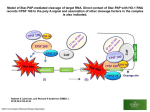
Transposable element wikipedia , lookup
Gene therapy wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Gene desert wikipedia , lookup
Gene expression programming wikipedia , lookup
Genome evolution wikipedia , lookup
Genome (book) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
X-inactivation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genome editing wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Helitron (biology) wikipedia , lookup
Microevolution wikipedia , lookup
Gene expression profiling wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Designer baby wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
History of RNA biology wikipedia , lookup
Messenger RNA wikipedia , lookup
Epigenetics of human development wikipedia , lookup
RNA interference wikipedia , lookup
RNA-binding protein wikipedia , lookup
Non-coding RNA wikipedia , lookup
Epitranscriptome wikipedia , lookup
Primary transcript wikipedia , lookup
Post-transcriptional modifications 5’ capping Cap a methylated guanosine bound through a 5’-5’ triphosphate linkage to the 5’phosphate of RNA 1 Steps: 1. RNA triphosphatase, removes terminal phosphate 2. Guanylyl transferase adds the capping GMP from GTP 3. Two methyl transferases methylate the N7 of the capping guanosine and the 2’Omethyl group of the penultimate nt. Occurs early in transcription process, before chain reaches 30nt. Functions: Protection: a cap is joined through special linkage found nowhere else in RNA. Might be expected to protect from RNases that start at 5’ end of substrates Translatability Eukaryotic mRNA gains access to the ribosome for translation via cap binding protein. If there is no cap, the CBP can not bind – translation is very poor. Transport of mRNA out of the nucleus Proper splicing of mRNA – removal of the first intron is dependent on the cap in vitro. This effects may be mediated by cap-binding complex that is involved in spliceosome formation 2 Experiment: LUC mRNA with and w/o cap, with and w/o polyA introduced in tobacco cells Outcome: polyA and cap act synergistically to stabilize and enhance translation of mRNA Polyadenylation •rRNA and tRNA do not have poly A tail. •Process of adding poly A nts is called polyadenylation •Most eukaryotic mRNAs and their precursors have a chain of about 250nt long at their 3’ end. •It is added post-transcriptionally by poly (A) polymerase. 3 Functions: Protection Translatability Stability Functions: Effects of polyadenylation on translatability and stability of mRNA 4 Mechanism of polyadenylation: •Transcription of genes goes beyond the polyadenylation site. •Then transcript is cleaved and polyadenylated at the 3’end created by cleavage. •Polyadenylation signals: in mammalian genes – AATAAA about 20-30 bp before the polyadenylation site. •In RNA – sequence AAUAAA – in most mammalian RNAs 20nt before their polyA Experiments: importance of AAUAAA was tested First the nts were deleted between this sequence and polyA – deletion shifts the polyA site downstream ~ the number of nts deleted •No deviation in AATAAA could occur w/o destroying polyadenylation •AAUAAA is not sufficient for polyadenylation, otherwise polyadenylation would occur downstream of AAUAAA sequences in many introns, but it does not •Polyadenylation is destroyed by deleting sequences immediately downstream of polyadenylation site. Region downstreams might contain another element of polyadenylation signal. This signal is not highly conserved, but is GT or T reach. 5 •Thus: efficient mammalian polyadenylation signal consists of an AAUAAA motif about 20nts upstream of polyadenylation site in the re-mRNA, followed 23 or 24 nt later by a GU-rich motif, followed immediately by the U-motif. •Plant signals usually contain the AAUAAA motif, but more variation in this region is allowed that in mammalian AAUAAA. •Yeast polyadenylation signals are more different, and rarely contain the AAUAAA. Polyadenylation requires cleavage of the pre-mRNA and polyadenylation at the cleavage site. Cleavage – proteins – CPSF – cleavage and polyadenylation specificity factor •CstF - cleavage stimulation factor •CFI – cleavage factor I •CFII – cleavage factor II •polyA polymerase •And RNA polymerase II, in particular the CTD of RPB1. 6 Initiation of polyadenylation •The signal to initiate the polyadenylation of the cleaved substrate – AAUAAA followed by at least 8 nt. •Once polyA reaches about 10nt, further polyadenylation becomes independent of the signal and depends on polyA itself •Two proteins participate in initiation of the process: CPSF that binds to the AAUAAA motif and polyA polymerase Elongation •Required specificity factor called PolyA binding protein (PABII). Binds to a preinitiated oligoA and aids polymerase to elongate the polyA to 250nt •PABII acts independently of the AAUAAA motif •It depends only of polyA and its activity is enhanced by CPSF. Gene silencing Multiple copies of transgenes introduced into plant cells can interact with each other and with homologous host genes in trans, resulting in the inactivation of expression of both genes. This phenomenon called homology-dependent gene silencing (HDGS) occurs generally in plants and is similar to certain types of gene silencing described for fungi, protozoa, nematodes, insects and mammals. 7 Gene silencing The results of nuclear run-on transcriptioin experiments suggest there are: Transcriptional (TGS) and Posttranscriptional (PTGS) forms of HDGS. Both types of HDGS are often associated with de novo DNA methylation, which is usually confined to promoters in cases of TGS and transcribed regions in instances of PTGS. PTGS In the case of PTGS, the interaction in trans of genes with similar transcribed sequences results in sequence-specific degradation of RNAs derived from the genes involved. Highly expressed single-copy loci, transcribed inverted repeats, and poorly-transcribed complex loci can act as sources of diffusible signals that trigger PTGS. In plants, posttranscriptional gene silencing (PTGS) is implicated as an antiviral mechanism. Intercellular spreading of the PTGS effect allows the plant to respond in a systemic manner after a localized viral challenge. 8 Gene silencing In some cases, mobile, sequence-specific silencing signals can move from cell-to-cell or even over long distances in the plant. Several current models hold that silencing signals are “aberrant” RNAs (aRNA), that differ in some way from normal mRNAs. The most likely candidates are small antisense RNAs (asRNA) and double-stranded RNAs (dsRNA). Most current models assume that silencing signals interact with target RNAs in a sequence specific fashion. This results in degradation usually in cytoplasm, by exonucleolytic as well as endonulceolytic pathways, which are not necessarily PTGS specific. Possible mechanisms Gene silencing is probably often the result of more than one mechanism. Transcriptional gene silencing (TGS) is often associated with methylation of the gene, which may inhibit transcription. In posttranscriptional gene silencing (PTGS), high levels of normal mRNA can cause activation of RNA-dependent RNA polymerases (RdRP), which can synthesize antisense transcripts. Antisense transcripts can also be synthesized when a gene is present in high copy number, especially where tandem-inverted repeated copies are present. 9 Possible mechanisms Double-stranded RNAs resulting from either RdRP activity or base-pairing between antisense transcripts and mRNAs become targets for ribonucleases, which degrade dsRNAs into small fragments of about 21-25bp. This process appears to be part of the natural defense against viral dsRNAs. Small dsRNAs may serve to target nuclear copies of the gene for methylation, resulting in a feedback mechanism for gene silencing. dsRNAs can also be transmitted intercellularly via plasmodesmata, causing systemic gene silencing. Again, this systemic component of gene silencing is likely part of the natural defense against viruses. In RNAi, introduced double-stranded RNA (dsRNA) (i) is recognized by proteins (ii) which, starting from the dsRNA termini, cleave it (iii) into ~22 nt fragments to produce nucleoprotein complexes (iv). 10 The strands of the short dsRNA are separated and one strand is used as a guide to recognize single-stranded RNA (ssRNA) with complementary sequences (v). Each complex cleaves the ssRNA at a position approximately in the middle of the guide sequence (v, vi). PTGS in plants In PTGS in plants (b), an RNAi-like cleavage mechanism operates. The dsRNA directing the mechanism is introduced by a replicating virus or an inverted repeat transgene that produces self-complementary (hairpin) RNA. 11 PTGS in plants The short RNAs from the cleavage (or alternatively uncleaved dsRNA) enter the nucleus to guide a methyltransferase complex to sequences for methylation and also spread into other cells to direct the cleavage of homologous ssRNAs. PTGS in plants The possible points of action of viral silencing suppressors, p25 and HC-Pro, are shown. p25 prevents the spread of the mobile PTGS signal but does not inhibit existing PTGS. HCPro is a viral silencing suppressor. 12 Posttranscriptional RNA modification -editing RNA editing – is a posttranscriptional changes of RNA sequences involving: a) Insertions b) Deletions c) Substitutions (conversions) …of nucleotides within an RNA chain. In contrast: Capping, splicing, 3’ end formation, trimming of ends, modification of bases (e.g. methylation), etc., etc. … are RNA processing. RNA editing Type of RNA editing Base Mechanism Posttransciptional insertion and deletion of nucleotides. U Guide RNAs C ? Cotranscriptional insertion of nucleotides A Modificationsubstitution of nucleotides C (to U) Polyadenylati on Polymerase stuttering Deamination G U (to C) A (to I) Genes and Organism Mitochondrial genes of Trypanosomes Mitochondrial genes of Physarum polycephalum Mitochondrial genes of vertebrates Phosphoprotein gene of Paramyxoviruses Apolipoprotein gene of mammals. Mitochondrial and chloroplast genes of plants Wilm’s tumor susceptibility gene, WT1 of mammals Mammalian glutamate receptor genes, matrix gene of Measles virus. 13 Conversions Insertion/deletions Examples 1. Transcipts of trypanosomes 2. Mitochondria of P. polycephalum 3. Paramyxoviruses (P gene) 4. Mitochondria of vertebrates 5. Apolipoprotein B gene of mammals 6. Mitochondria of higher plants 7. Chloroplasts of higher plants 8. Glutamate-gated ion channels of mammals ..AAUUUAUGUUGUCUUU ..AUCUCUAAGGGUUUAACCGG Frameshift ..AUUAAAAAGGGGCACAC… Frameshift ..CAGUAAAAAAAAAA… Stop C ..GAUAUAAUUUGAUCAGUAUA… C Stop C ..UUUUUCAUUGUGGUUUAC… PHE TYR C ..AAUAAUAUGGCGAAACAU… Start A ..UUUAUGCGGCAAGGA… ARG Discovery of RNA Editing It was noted early on that several of the structural genes had frameshifts and lacked translation initiation codons. In 1986, Benne et al. discovered that U residues were inserted into the mRNA after transcription and that this overcame the frameshifts. U deletions also were found to occur at a lower frequency. The extent of this "RNA editing" in the lizard Leishmania, L. tarentolae, is indicated. 14 Discovery of RNA Editing Several mRNAs are edited at the 5' end and several are" panedited" throughout the gene, creating an open reading frame from nonsense sequence. Discovery of RNA Editing Blum et al. discovered that a novel class of small RNA molecules, transcribed both from the maxicircle and the thousands of minicircles, contained the editing information. The gRNAs were discovered by a computer search for short maxicircle sequences complementary to edited mRNA sequences. 15 Discovery of RNA Editing These guide RNAs or gRNAs had a 5' sequence complementary to the mRNA just 3' of the first editing site. They also had a non-encoded 3' oligo[U] tail. The central portion of the gRNA contained sequence that was complementary to edited mRNA sequence, provided G was allowed to pair with U and well as with C. Therefore the gRNA would form a perfect duplex with edited mRNA except for the oligo[U] tail. Reading: p. 471-488, 521-526 16