* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lect11_DNAMethylation
DNA damage theory of aging wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Genome (book) wikipedia , lookup
Ridge (biology) wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Point mutation wikipedia , lookup
Epitranscriptome wikipedia , lookup
Molecular cloning wikipedia , lookup
Genomic library wikipedia , lookup
DNA vaccination wikipedia , lookup
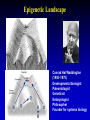
Minimal genome wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
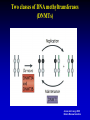
Metagenomics wikipedia , lookup
Primary transcript wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Human genome wikipedia , lookup
DNA supercoil wikipedia , lookup
Genome evolution wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
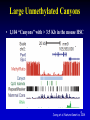
Extrachromosomal DNA wikipedia , lookup
Epigenetic clock wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Genomic imprinting wikipedia , lookup
Gene expression profiling wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Designer baby wikipedia , lookup
Oncogenomics wikipedia , lookup
Epigenetics of depression wikipedia , lookup
Genome editing wikipedia , lookup
Microevolution wikipedia , lookup
Epigenetics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Behavioral epigenetics wikipedia , lookup
Non-coding DNA wikipedia , lookup
History of genetic engineering wikipedia , lookup
Helitron (biology) wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
DNA methylation wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Epigenomics wikipedia , lookup
ChIP-seq Downstream Analysis Xiaole Shirley Liu STAT115, STAT215, BIO298, BIST520 Human TF Binding Distribution • • • • • • Most TF binding sites are outside promoters How to assign targets? Binding in promoter (how far)? Nearest distance? Number of binding? Other knowledge? • Still open Q 2 Higher Order Chromatin Interactions Interactions ~ follows exponential decay with distance Lieberman-Aiden et al, Science 2009 How to Assign Targets for Enhancer Binding Transcription Factors? • Regulatory potential: sum of binding sites weighted by distance to TSS with exponential decay • Rank1 of genes based on binding TSS 4 How to Assign Targets for Enhancer Binding Transcription Factors? • Regulatory potential: sum of binding sites weighted by distance to TSS with exponential decay • Rank1 of genes based on binding • Rank2 of genes based on expression – Differential expression upon TF perturbation – Gene expression correlation in cohorts • Rank ordered list of targets: Rank1 * Rank2 • Only minority of binding sites is functional in a condition 5 BETA • Binding expression target analysis • Is TF an activator, repressor or both? – Do genes with higher regulatory potential show more up/down expression than random genes 6 BETA • Binding expression target analysis • Is TF an activator, repressor or both? – Do genes with higher regulatory potential show more up/down expression than random genes • • • • Functional analysis of targets? How can a factor be both activator and repressor? Collaborating transcription factors Motif analysis on the binding sites near up vs down genes separately ER TF?? 7 Summary • ChIP-seq identifies genome-wide in vivo protein-DNA interaction sites • MACS for ChIP-seq peak calling – Shift reads, Poisson with dynamic λ • ChIP-seq QC: FDR, fold, conservation, etc • Functional analysis of ChIP-seq data: – Target identification – Activator / repressor function – Motif analysis Break 8 Epigenetics Xiaole Shirley Liu STAT115, STAT215, BIO298, BIST520 Epigenetics • Heritable changes in gene function that occur without a change in the DNA sequence – How come not all the motif sites are bound by the factor? – How come TF binding only regulate some of the nearby genes? Epigenetics • Functionally relevant changes to the genome that do not involve a change in the nucleotide sequence • The study of heritable (transgenerational) changes in gene activity that are not caused by changes in the DNA sequence • The study of stable, long-term alterations in the transcriptional potential of a cell that are not necessarily heritable The Making of a Queen Larvae From Ting Wang, Wash U Queen Worker Agouti Mice and DNA Methylation In Human • Nature vs nurture • Zygotic twins: same DNA different epigenome • North American Ice Storm of 1998 Epigenetic Landscape Conrad Hal Waddington (1905–1975) Developmental biologist Paleontologist Geneticist Embryologist Philosopher Founder for systems biology Components • DNA-methylation • Nucleosome position and histone modifications • Chromatin accessibility • Higher order chromatin interactions Break DNA Methylation 17 DNA Methylation Distribution in Mammalian Genomes • In human somatic cells, 60%-80% of all CpGs (~1% of total DNA bases) are methylated – Most methylation is found in “repetitive” elements • “CpG islands”, GC-rich regions that possess a high density of CpGs, remain methylation-free – The promoter regions of ~70% of genes have CpG islands Two classes of DNA methyltransferases (DNMTs) Jones and Liang, 2009 Nature Review Genetics Inheritance of DNA Methylation DNA Methylation Detection • Bisulfite sequencing – Unmethyl C T, sequence 30X – High resolution, quantitative, but expensive! – RRBS enriches CpG rich regions for sequencing ACGGGCTTACTTGCTTTCCTACGGGCTTACTTGCTTTCCTACGGGCTTACTTGCTTTCCTACGGGCTTACTTGC CGGGTTTATTTGCTTTTTTATGGGC TGGGTTTATTTGCTTTTTTATGGGC TGGGTTTATTTGCTTTCCTATGGGC CGGGCTTATTTGCTTTCCTATGGGC CGGGCTTATTTGCTTTCCTATGGGC 3/5 60% methylated 0/5 0% methylated BS-seq Methylation Call • Bismark: Krueger & Andrews, Bioinfo 2011 – Create additional sequence in the BWA index to account for the C -> T conversion • Methylation or genome sequence difference? + ACGAACGCAACG ACGAATGCAATG + ACGAATGCAATG - TGCTTGTGTTGT ACGAATGCAATG TGCTTACGTTAC Analysis Issues • Methylation level within a short distance is consistent: smoothing across bigger regions • Most regions are either mostly methylated or mostly unmethylated (dichotomy) • Estimate tumor purity 24 DNA Methylation Controls Gene Expression • Methylation at CpG islands often repress nearby gene expression • Many highly expressed genes have CpG methylation on their exons Some genes could be imprinted, so maternal and paternal copies have different DNA methylation Large Unmethylated Canyons • 1,104 “Canyons” with > 3.5 Kb in the mouse HSC 26 Jeong et al. Nature Genetics. 2014 DNA Methylation in Cancer • Prevalent misregulation of DNA methylation in cancer: global hypomethylation and CpG island hypermethylation • Methylation variable regions in cancer DNA Demethylation • Recently, another type of DNA methylation called hydroxyl methylation (hmC) is found • hmC is an intermediate step between mC and C. • TET family of proteins are important for DNA demethylation • Mutation in TET is linked to many cancers Acknowledgement • Evan Johnson, BU • Ting Wang, Wash U • Wei Li, Baylor 29