* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download ESTs to genome
Extrachromosomal DNA wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Oncogenomics wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Designer baby wikipedia , lookup
Polyadenylation wikipedia , lookup
Messenger RNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Transposable element wikipedia , lookup
Epigenetics of human development wikipedia , lookup
RNA interference wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Point mutation wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Minimal genome wikipedia , lookup
Pathogenomics wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genomic library wikipedia , lookup
Microsatellite wikipedia , lookup
RNA silencing wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Human Genome Project wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Metagenomics wikipedia , lookup
Helitron (biology) wikipedia , lookup
Genome evolution wikipedia , lookup
Non-coding RNA wikipedia , lookup
Non-coding DNA wikipedia , lookup
Human genome wikipedia , lookup
History of RNA biology wikipedia , lookup
Epitranscriptome wikipedia , lookup
Genome editing wikipedia , lookup
RNA-binding protein wikipedia , lookup
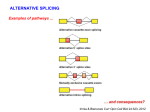
Alternative Splicing 1 Eukaryotic genes Splicing Mature mRNA 2 The mechanism of RNA splicing 3 The mechanism of splicing 5’ splice site 1 1 Branch point CAG GTRAGT A -OH A 3’ splice site 2 A 1 2 YYYYYYYYYNCAG G 2 4 Alternative splicing 1 2 50-70% of mammalian genes Mature splice variant I 1 23 3 4 Can be specific to tissue, developmentalstage or condition (stress, cell-cycle). Alternative Splicing 4 13 4 Mature splice variant II 5 Some types of alternative splicing Exon skipping Alternative Acceptor Alternative Donor Mutually exclusive Intron retention 6 Sex determination in fly 7 Sex determination in fly 8 Sex determination in fly 9 Many variants in one gene 10 DSCAM 11 Antibody secretion 12 Antibody secretion immunoglobulin μ heavy chain 13 Tissue specific alternative splicing 14 Detection of alternative splicing By sequencing of RNA Old methods (1995-2007) – ESTs New methods: – Splicing-sensitive microarrays – RNA-seq 15 Expressed Sequence Tags (ESTs) AAA AAA AAA AAA AAAAAAAAA mRNA RT AAAAAAAAA TTTTTTTTTT cDNA Cloning Vector 16 EST preparation 5’ EST Picking a clone 3’ EST Random-primed EST Average size of EST ~450bp 17 Alignment of ESTs to the genome DNA EST EST EST EST EST EST 8 million public human ESTs, collected over >10 years (NCBI) 18 Splicing microarrays 19 Massive sequencing of RNA (RNA-seq) 20 RNA-seq on multiple tissues 21 Wang et al Nature 2008 Splicing regulation 22 Tissue specific alternative splicing How is this process regulated? 23 Regulation of alternative splicing Splicing Enhancers/Silencers Specifically bind SR proteins 24 Model for ESE action SR brain Y(n) AG Weak splice site Exon Exonic Splicing Enhancer (ESE) 25 SR proteins structure 26 27 Discovery of ESEs Exon Silent mutations can cause exon skipping 28 Regulators of splicing Signal transduction ISE SR proteins (Splicing factors) ESE/ESS ISS • Complex regulation usually exists • Hard to find intronic elements • For most alt exons – regulation unknown 29 How can we break the regulatory code? 1. Comparative genomics 2. High throughput methods 30 Comparative genomics: Use the mouse genome to find sequences that regulate alternative splicing 31 Human-mouse comparisons 32 The mouse genome 100 million years of evolution Average conservation in exons: 85% Only 40% of intronic sequences is alignable Average conservation in alignable intronic sequences: 69% Average conservation in promoters: 77% Function => evolutionary conservation 33 Conservation of near introns 34 (from VISTA genome browser, http://pipeline.lbl.gov) Collection of exons Human DNA AF010316 AF217965 AF217972 BE614743 BE616884 AI972259 35 Finding the mouse homolog Mouse DNA Human DNA AF010316 AF217965 AF217972 BE614743 BE616884 AI972259 243 Alt. 1753 Const. 36 Conservation in the intronic sequence near exons Mouse DNA Human DNA AF010316 AF217965 AF217972 BE614743 BE616884 AI972259 243 Alt. 1753 Const. 37 Results Constitutive exons Constitutive exons 17% 83% Alternative exons Flanking conserved introns Alternative exons 77% 23% ~100 bp from each side of the 38exon Conservation of introns 39 Alternative splicing regulatory sequences? Could serve as binding sites for splicing regulatory proteins 40 Motif searching Top scoring hexamer in conserved downstream regions: TGCATG (9-fold over expected) Not over-represented downstream to constitutive exons. Binding site for FOX1 (splicing regulatory protein) 41 Functional elements in the human genome 5% of the human genomic sequence is considered functional 42 Functional elements in the human genome Composition of functional 5% genomic sequence Unknown 50% Coding exons 30% UTR and promoters 20% 43 Impact of splicing regulatory elements ~12,000 alt. spliced exons in the genome 77% have conserved flanking intronic sequences ~100bp conserved on each side 12,000 exons * 100 bp * 2 introns * 0.77= 2M bases ==>At least 2 Million bases in the human genome might be involved in alternative splicing regulation. >1% of all functional DNA in the genome regulates alt splicing! 44 How can we break the regulatory code? 1. Comparative genomics 2. High throughput methods 45 CLIP-seq Ule et al, Science 2003: 340 sequences Licatalosi et al, Nature 2008: 412,686 sequences 46 Nova, a brain-specific splicing regulator Ule et al, Science 2003: 340 sequences 47 48 Ule et al, Science 2003: 340 sequences Extracting the regulatory motifs 49 The power of deep sequencing (2008) 50 Mutations causing aberrant splicing Exon ~15% of all point mutations linked to genetic disorders involve splicing alterations 51 Mutations causing aberrant splicing: SMN 52 Summary – alt splicing Increases the coding capacity of genes We have 25,000 genes but much more protein isoforms 53 RNA EDITINA 54 RNA EDITING 55 What is RNA editing? Alters the RNA sequence encoded by DNA in a single-nucleotide, site-specific, manner If splicing is “cut and paste” editing is the “spelling checker”. 56 Mode of operation: A-to-I editing Editing performed by ADAR enzymes (dsRNA specific adenosine deaminases) Double strand RNA is required A-> G 57 Mechanism of RNA-editing (A-to-I) 58 Functions of RNA editing Defense against dsRNA viruses Also involved in endogenous regulation 59 60 Functional consequences of RNA editing Splicing Protein change RNA stability In human, RNA editing is particularly pronounced in brain tissues, due to excess of ADAR expression in brain Neural disorders (glioblastoma, epilepsy, ALS) are linked to changes in RNA-editing patterns Editing levels vary in other tissues (minimal editing in skeletal muscle, pancreas). 61 Finding RNA-editing sites Theoretically easy : find mismatch between genome to RNA Huge number of sequencing errors Mutations Duplications SNPs Signal drowns in noise 62 Computational approach for identification of editing sites Alignment of ESTs to genome Find potential intramolecular dsRNA Data cleaning 63 Levanon et al, Nature Biotech 2004 Intramolecular dsRNA RNA Exon Intron 64 Levanon et al, Nature Biotech 2004 ESTs to genome 65 Levanon et al, Nature Biotech 2004 •dsRNA regions 66 Levanon et al, Nature Biotech 2004 •dsRNA regions •Masking EST’s ends 67 Levanon et al, Nature Biotech 2004 •dsRNA regions •Masking EST’s ends •Masking poor sequence regions 68 •dsRNA regions •Masking EST’s ends •Masking poor sequence regions •Removing known genomic SNPs 69 Levanon et al, Nature Biotech 2004 •dsRNA regions •Masking EST’s ends •Masking poor sequence regions •Removing SNPs •Collecting candidates 70 Levanon et al, Nature Biotech 2004 Results DNA RNA (ESTs) 71 72 Levanon et al, Nature Biotech 2004 73 RNA-editing – a source for human transcripts diversity >12,000 editing sites in >1,600 human genes Vast majority of editing – in UTRs Vast majority of editing – in Alu (repetitive) A few editing sites in protein-coding regions 74 Levanon et al, Nature Biotech 2004 And the obligatory next generation sequencing study… (Li, Levanon et al, Science 2009) Editing sites in non-repetitive regions 75 Connection between editing and splicing ADAR gene (editing enzyme) Negative feedback loop 76 Evolution of a new exon 77 Summary – alt splicing and RNA editing Increases the coding capacity of genes We have 25,000 genes but much more protein isoforms 78