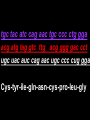* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download gene regulation
Transposable element wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Cancer epigenetics wikipedia , lookup
RNA interference wikipedia , lookup
History of RNA biology wikipedia , lookup
Protein moonlighting wikipedia , lookup
Gene desert wikipedia , lookup
Human genome wikipedia , lookup
Gene nomenclature wikipedia , lookup
Genome evolution wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Epigenomics wikipedia , lookup
Genetic engineering wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Genome (book) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Messenger RNA wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Gene expression profiling wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Non-coding RNA wikipedia , lookup
History of genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Non-coding DNA wikipedia , lookup
Transcription factor wikipedia , lookup
Microevolution wikipedia , lookup
Point mutation wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Designer baby wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Helitron (biology) wikipedia , lookup
Epitranscriptome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Primary transcript wikipedia , lookup
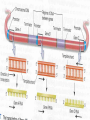
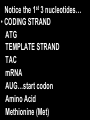
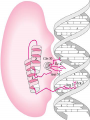
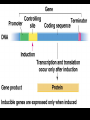
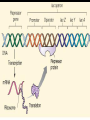
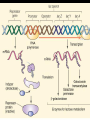
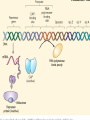
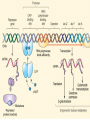
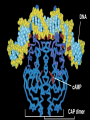
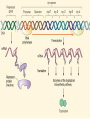
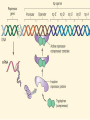
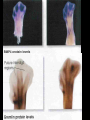
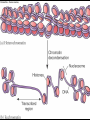
GENE REGULATION OXYTOCIN • The following gene is from mRNA that has been copied with reverse transcriptase to produce cDNA • Sequence provided is from the coding strand 1 accagtcacg gaccctggac ccagcgcacc cgcaccatgg ccggccccag cctcgcttgc 61 tgtctgctcg gcctcctggc gctgacctcc gcctgctaca tccagaactg ccccctggga 121 ggcaagaggg ccgcgccgga cctcgacgtg cgcaagtgcc tcccctgcgg ccccgggggc 181 aaaggccgct gcttcgggcc caatatctgc tgcgcggaag agctgggctg cttcgtgggc 241 accgccgaag cgctgcgctg ccaggaggag aactacctgc cgtcgccctg ccagtccggc • 301 cagaaggcgt gcgggagcgg gggccgctgc gcggtcttgg gcctctgctg cagcccggac • 361 ggctgccacg ccgaccctgc ctgcgacgcg gaagccacct tctcccagcg ctgaaacttg • 421 atggctccga acaccctcga agcgcgccac tcgcttcccc catagccacc ccagaaatgg • 481 tgaaaataaa ataaagcagg tttttctcct ct SIGNAL SEQUENCE • 37-93 • Before signal sequence is the leader sequence • Mature Oxytocin 94-120 27 nucleotides = 9 amino acids • Preproprotein 37- 411 • Signal seq, oxytocin + neurophysin I Proprotein 94-411 Oxytocin + neurophysin I • Mature Peptides modified from Proprotein • Trailer sequence responsible for PolyA-polymerase action 1 accagtcacg gaccctggac ccagcgcacc cgcaccatgg ccggccccag cctcgcttgc 61 tgtctgctcg gcctcctggc gctgacctcc gcctgctaca tccagaactg ccccctggga 121 ggcaagaggg ccgcgccgga cctcgacgtg cgcaagtgcc tcccctgcgg ccccgggggc 181 aaaggccgct gcttcgggcc caatatctgc tgcgcggaag agctgggctg cttcgtgggc 241 accgccgaag cgctgcgctg ccaggaggag aactacctgc cgtcgccctg ccagtccggc • 301 cagaaggcgt gcgggagcgg gggccgctgc gcggtcttgg gcctctgctg cagcccggac • 361 ggctgccacg ccgaccctgc ctgcgacgcg gaagccacct tctcccagcg ctgaaacttg • 421 atggctccga acaccctcga agcgcgccac tcgcttcccc catagccacc ccagaaatgg • 481 tgaaaataaa ataaagcagg tttttctcct ct Notice the 1st 3 nucleotides… • CODING STRAND ATG TEMPLATE STRAND TAC mRNA AUG…start codon Amino Acid Methionine (Met) tgc tac atc cag aac tgc ccc ctg gga acg atg tag gtc ttg acg ggg gac cct ugc uac auc cag aac ugc ccc cug gga Cys-tyr-ile-gln-asn-cys-pro-leu-gly WOBBLE EFFECT • rd 3 base of tRNA may form H-bonds with more than 1 kind of nucleotide • Ie AAU and AAC Asn TRANSCRIPTION FACTORS RECOGNIZE SEQUENCES • Transcription factors recognize DNA sequences inorder to target specific genes ROLE OF REGULATORY PROTEINS • Transcription factors are genetic switches • Master Genes ie HOX genes code for transcription factors • Related “Regulatory Proteins” in prokaryotes PROKARYOTIC REGULATION • No hormonal regulation • No introns no splicing • No capping/tailing • Coupled transcription/translation • Constitutive – always on; can be regulated (enzymes in glycolysis) • Inducible – off but can be switched on • Repressible – on but can be switched off OPERONS • Cluster of genes in which expression is regulated by operatorrepressor protein interactions, operator region, and the promoter. • Promoter • Repressor • Operator (controlling site) • Coding sequences • Terminator Lac Operon • Lactose is an inducer • “Inducible” operon • Negative regulation –-galactosidase (lacZ) • Breaks lactose into glucose + galactose. • Converts lactose to the allolactose, regulates lac operon. –Lactose permease (lacY) • Transports lactose across cytoplasmic membrane. –Transacetylase (lacA) • Positive control when lactose is E. coli’s sole carbon source (but not if glucose also is present). • Catabolite activator protein (CAP) binds cAMP, activates, and binds to a CAP recognition site upstream of the promoter (cAMP is greatly reduced in presence of glucose). • CAP changes the conformation of DNA and facilitates binding of RNA polymerase and transcription. • When glucose and lactose are present, E. coli preferentially uses glucose due to low levels of active CAP (low cAMP). • Regulation of the trp operon: • “repressible” gene • 1. Repressor/operator interaction • When tryptophan is present, tryptophan binds to trpR gene product. • trpR protein binds to the trp operator and prevents transcription. • Is TRP operon neg or pos regulation? • What type of regulation controls repressors/inducers? Jacob and Monod • What is true for E.coli is also true for the elephant! GENETIC SWITCHES • A switch includes.. a) Binding protein b) Binding protein recognizes a strecth of DNA Significance? • Mutate the gene encoding the transcription factor or the DNA sequence to which it binds and gene expression can be altered! GENE INACTIVATION • 1. CHROMATIN structure – Heterochromatin (tight– gene off) vs Euchromatin (loose – gene on) • 2. Methylation – adding methyl group to inactivate genes on DNA • 3. small RNA affects chromatin structure / interferes with transcription (RNAi system) Methylation • Reinforces inactivation • Involved in Barr-Body (inactive X chromosome) TRANSCRIPTIONAL CONTROL • Signals (hormones) in eukaryotes • Environmental- heat shock proteins • Regulator proteins (Transcription factors) – bind to TATA (like prokaryotes) • UPEs (Upstream Promoter Elements) – increase efficiency of RNA pol. Example… • Glucocorticoids released by stress bind to steroid receptor (in liver) forms complex binds to DNA activates genes involved in gluconeogenesis Weakly transcribed Strongly transcribed • TRANSLATIONAL CONTROL • CAPPING – 7’ methyl guanosine added to 5’ role in initiation/ protection • Poly A tail protection, promotes transport out of Nucleus • Splicing – Introns removed Exons for code • Differential mRNA processing –Cells in each tissue produce own version of mRNA –For example, different forms of troponin, a protein that regulates muscle contraction, produced in different muscle tissue • POST TRANSLATIONAL • modification of polypeptide (ie reduction in size of proinsulin to insulin) • chemical modification – addition of phosphate (Kinases) removal of phosphate (Phosphatases) • Examine the insulin pathway as a form of gene regulation