* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Transposons - iPlant Pods
DNA damage theory of aging wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Ridge (biology) wikipedia , lookup
Gene desert wikipedia , lookup
DNA vaccination wikipedia , lookup
Primary transcript wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
DNA supercoil wikipedia , lookup
Public health genomics wikipedia , lookup
Molecular cloning wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Genomic imprinting wikipedia , lookup
Genetic engineering wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Copy-number variation wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Gene expression programming wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
Metagenomics wikipedia , lookup
Oncogenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Point mutation wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Microsatellite wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Pathogenomics wikipedia , lookup
Human Genome Project wikipedia , lookup
Designer baby wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Human genome wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Non-coding DNA wikipedia , lookup
Minimal genome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome editing wikipedia , lookup
Genomic library wikipedia , lookup
Genome evolution wikipedia , lookup
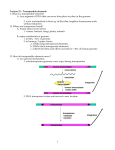
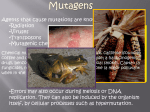
The Dynamic Genome Transposons What are Transposons? Transposable element (transposon, TE): DNA sequence competent to insert into new places Transposition of DNA explains mottled kernels in maize Some definitions and figures from Lisch 2009: Annu. Rev. Plant Biol. 2009.60:43-66. What are Transposons? Transposable element (transposon, TE): DNA sequence competent to insert into new places (1) At the beginning of kernel development, the Ds transposon inserts into the colored (C) gene, resulting in colorless tissue. (2) Ds transposition early in kernel development restores the C gene, giving rise to a large colored sector. (3) Transposition later in kernel development results in smaller sectors. Learn more at: weedtowonder.org/jumpingGenes.html What are Transposons? Transposable element (transposon, TE): DNA sequence competent to insert into new places (Ac/Ds) “Cut & Paste” (Alu) “Copy & Paste” What are Transposons? •Most genomes contain multiple transposon families. •Each family contains autonomous and non-autonomous elements. •Autonomous elements encode their own moving competency. •Non-autonomous elements are moved by other elements. Autonomous element Gene(s) Nonautonomous elements Class I transposons are being copied multiplicative. Class II transposons can undergo copying, too, if transposing during DNA replication What are Transposons? Transposons make up most of (most) eukaryotic genomes • ~50% of the genomes of human, chimp, mouse, gorilla • ~75% of the maize genome • ~85% of the barley genome • ~98% of the iris genome Iris brevicaulis Iris fulva Hs 11: http://dnalc.org/resources/3d/chr11.html What are Transposons? Effect of transposons & genome duplications on genomes Sorghum 700 Mb Rice 450 Mb Barley 5,000 Mb Wheat 20,000 Mb Maize 2,500 Mb Oats ~20,000 Mb Transposons in Action How do organisms live with TEs? • Most TEs are broken (cannot tranpose; “fossils”). • Active TEs evolved to insert into “safe-havens.” • Host regulates TE movement. • TEs can provide advantages. Ping/mPing mPing: MITE (Multi-insertional TE) Deletion-derivative of Ping Requires Ping transposase to jump MITEs are being amplified to high copy numbers mPing copy number in O. japonica OVER 1000 mPing copies mPing Japonica strains Over 1000 copies of mPing in 4 related strains…. Naito et al PNAS (2006)) Takatoshi Tanisaka lab (Kyoto University) mPing insertions in genome • predominantly in genic regions in euchromatin • even inserts in heterochromatin are in genes • where does mPing insert in and around genes? mPing insertions in genes 12 shared (n=926) 10 unshared (n=736) (%) 8 expect. 6 4 2 0 5'UTR UTR exon Exon intron 3'UTR UTR mPing insertions rare in coding-exons TEs can alter gene expression Os02g0135500 (-41) 2.5 NB EG4 (mPing+) A123 (mPing+) A157 2 1.5 1 0.5 0 control cold salt dry mPing found to confer cold and salt inducibility TEs can alter gene expression Can this have phenotypic consequences? Nipponbare EG4 EG4 is salt tolerant Rapid mPing amplification (burst) • Massive amplification largely benign • Subtle impact on the expression of many genes • Produces stress-inducible networks (cold, salt, others?) • Generates dominant alleles Naito et al, Nature, 2009 TEs as tools of evolutionary change • TEs usually inactive. • “Stress” conditions may activate TEs. • Active TEs increase mutation frequency. • Most mutations caused by TEs neutral or harmful. • A rare TE-induced mutation (or rearrangement) may be adaptive. Transposable elements can shake up otherwise conservative genomes and generate new genetic diversity. TEs for student research projects • (relatively) simple • incredibly abundant • evolve rapidly • promote rapid genome evolution • largely ignored (discovery) Suppl.: DNA transposons can be copied, too Gap repair from sister chromatid Jump into site that is then replicated Yellow Line Walk-through (Advanced Yellow Line Example) • Find homologs using DNA • Find homologs using protein • Locate transposons • Examine surroundings of transposon insertions • Identify active transposons and “molecular fossils” • Show recent transposon activity