* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Evolucijska genomika 2
Copy-number variation wikipedia , lookup
Genomic imprinting wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Oncogenomics wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Point mutation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
X-inactivation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Human genetic variation wikipedia , lookup
Metagenomics wikipedia , lookup
Public health genomics wikipedia , lookup
Gene desert wikipedia , lookup
Pathogenomics wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Gene expression programming wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Gene therapy wikipedia , lookup
Epigenomics wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Genetic engineering wikipedia , lookup
Minimal genome wikipedia , lookup
Epitranscriptome wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Transposable element wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
History of RNA biology wikipedia , lookup
Human Genome Project wikipedia , lookup
Genome (book) wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Non-coding RNA wikipedia , lookup
Genomic library wikipedia , lookup
Primary transcript wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Human genome wikipedia , lookup
Genome editing wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Microevolution wikipedia , lookup
RNA silencing wikipedia , lookup
Non-coding DNA wikipedia , lookup
RNA interference wikipedia , lookup
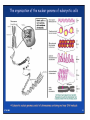
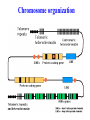
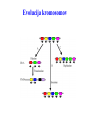
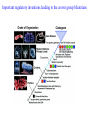
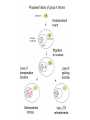
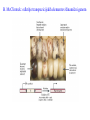
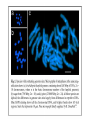
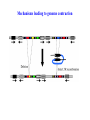
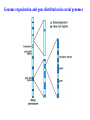
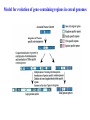
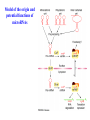
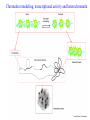
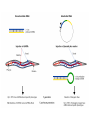
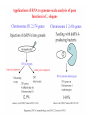
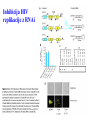
Skrivnostni svet nekodirajočih delov evkariontskih genomov Kordiš Dušan Odsek za biokemijo in molekularno biologijo, IJS, Ljubljana Nekodirajoča DNA zavzema pretežni del genomov pri eukariontih Human Genome At Glance Genomska arhitektura Chromosome organization Human Chromosome Y: Organization of Genes and DNA Elements * Heterochromatin Euchromatin *Centromere Evolucija kromosomov Eukaryote gene structure • Coding sequence gene not continuous 2 Fig 18-8 The forces affecting genome size evolution Velikosti genomov in število genov pri prokariontih in eukariontih Funkcionalna evolucija nekodirajoče DNA Human vs. Chimpanzee • A difference every 100 bases. • A new transposon every 50000 bases • Two chromosome in one species fused compared to the other. Deletion of a conserved noncoding sequence selectively reduces expression of cytokine genes over a long distance Percent identity plots comparing human and other sequences at three loci (CNS:red, exons:blue, introns:yellow) Cis-regulatorna evolucija gradual, stepwise accumulation of sites that impart quantitatively greater Hox influence over gene regulation Important regulatory inventions leading to the crown group bilaterians Cis-regulatory logic during development Introni: izvor, evolucija in vloga v genomu • What are introns? Stretches of DNA that are transcribed into RNA, then spliced out during RNA processing. Contain functional elements such as splicing signals, regulatory promoters, and other genes. Evolve very rapidly in size and content. Constitute 26%, 11%, and 24% of the nematode, fly, and human genomes. What forces drive the evolution of intron size? Inton Size - 10 to 100,000 nt Introns are transcribed Precursor RNA (dotted) hybridized with DNA (red) Mature mRNA (dotted) hybridized with DNA (red) Introns vary in size and number The complexity problem Gene numbers do not increase as much as expected with complexity: - worm and fly gene numbers (12-14,000) are only about twice those of yeast (6,000) and P. aeruginosa (5,500) - mammalian (human, mouse) gene numbers (~30,000) are only about twice those of invertebrates. Phenotypic variation in mammals is primarily associated with noncoding regions: - only ~10,000 out of ~3,000,000 polymorphisms between individual humans (0.3%) occur in protein coding sequences - only 1% of genes are different between humans and mice. This suggests that: - animals have a relatively stable core proteome, whose components are multitasked in differentiation and development - variations in phenotype occurs mainly by variation in the control architecture (unlike prokaryotes) 98% of transcriptional output in humans is noncoding RNA Transpozicijski elementi B. McClintock: odkritje transpozicijskih elementov/dinamični genom Four classes of parasitic DNA elements are found interspersed throughout the human genome Retrotransposons 1. 2. 3. Transposons 4. Kakšen % genoma zavzemajo TE Spremembe sesalskih genomov po retrotranspoziciji L1 elementov Do LINEs Mediate Genomic Plasticity ? Whole Genome Breakpoints (3300 Mb) (245 Mb) LINEs 20.4% 45.3% SINEs 13.1% 12.5% LTRs 8.3% 8.3% Kakšen vpliv ima načina razmnoževanja na preživetje TE nespolno spolno Povezava ekologije in velikosti genoma: dinamika genoma na populacijskem nivoju Kako lahko Alu elementi poškodujejo človeški genom ? Homologna rekombinacija med Alu elementi Fig 18-11 Mechanisms of genome expansion in the grass genomes Mechanisms leading to genome contraction Genome organization and gene distribution in cereal genomes Model for evolution of gene-containing regions in cereal genomes Genome defense and regulation by small RNAs RNA interferenca MicroRNA (miRNA) Epigenetsko utišanje transpozicijskih elementov Model of the origin and potential functions of microRNAs Model delovanja RNAi How RNAi initiates chromatin silencing Chromatin remodeling, transcriptional activity and heterochromatin Epigenetsko reprogramiranje med gametogenezo chromatin remodeling and demethylation during (a) normal fertilization and (b) during cloning by nuclear transfer. Chromatin as a template of genetic inheritance epigenetics as process inducing differentiated cellular states Central role for RNAi in genome maintenance: RNAi responds not only to exogenous nucleic acids but also to endogenous DNA parasites Uporaba RNA interference v funkcionalni genomiki in medicini Potencialna uporaba RNAi pri sesalcih RNA interferenca in zdravljenje raka (a) A viral vector delivers a gene encoding a small interfering RNA (siRNA) to silence the mutant allele of a cancer-causing gene. The vector encodes a short RNA hairpin, which is processed in the cytoplasm by the ribonuclease Dicer into the siRNA. (b) The siRNA acts as a sequence-specific guide for the RNA-induced silencing complex (RISC) to target cleavage of the mRNA from a specific gene, in this case, the mutant allele of an oncogene. The genetic flow and mRNA processing, indicating possible strategies for gene regulation HIV infection and replication have been targeted by RNA interference (RNAi) (red). preventing HIV entry and subsequent replication: RNAi has also been used to suppress CD4 expression in host cells (blue). Inhibicija HIV replikacije z RNAi Future possibilities for therapy treatments using RNAi vectors