* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Ribosome binding site Polysomes (多聚核糖体)
Peptide synthesis wikipedia , lookup
NADH:ubiquinone oxidoreductase (H+-translocating) wikipedia , lookup
Biochemistry wikipedia , lookup
Signal transduction wikipedia , lookup
Magnesium transporter wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Polyadenylation wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Point mutation wikipedia , lookup
Silencer (genetics) wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Paracrine signalling wikipedia , lookup
Expression vector wikipedia , lookup
Metalloprotein wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Interactome wikipedia , lookup
Bimolecular fluorescence complementation wikipedia , lookup
Acetylation wikipedia , lookup
Transcriptional regulation wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Protein structure prediction wikipedia , lookup
Western blot wikipedia , lookup
Protein purification wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Gene expression wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Genetic code wikipedia , lookup
Proteolysis wikipedia , lookup
Messenger RNA wikipedia , lookup
Biosynthesis wikipedia , lookup
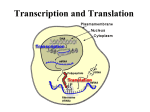
Chapter 8 Protein synthesis Protein synthesis • Aspects of protein synthesis • Mechanism of protein synthesis (Prokaryotic) • Initiation in eukaryotes • Translational control and posttranslational events 8.1: Aspects of protein synthesis • • • • • Codon-anticodon interaction Wobble (变偶性) Ribosome binding site Polysomes (多聚核糖体) Initiators tRNA Codon-anticodon interaction In the cleft of the ribosome, an antiparallel formation of three base pairs occurs between the codon on the mRNA and the anticodon on the tRNA. Some highly purified tRNA molecules were found to interact with more than one codon, and this ability is correlated with the presence of modified nucleosides in the 5’anticodon position, particularly inosine (次黄嘌呤)(formed by posttranscriptional processing of adenosine by anticodon deaminase) Wobble To explain the redundancy of the genetic code. 18 aa are encoded by more than one triplet codons which usually differ at 5’-anticodin base 5'-anticodon base is able to undergo more movement than the other two bases and can thus form nonstandard base pairs as long as the distances between the ribose units are close to normal. All possible base pairings at the wobble position No purine-purine or pyrimidine-pyrimidine base pairs are allowed as ribose distances would be incorrect (Neat!). U is not found as 5’-anticodon base Wobble pairing: non Waston-crick base paring Ribosome binding site (Shine-Dalgarno sequence) • Solely for prokaryotic translation • A purine-rich sequence usually containing all or part of the sequence 5'-AGGAGGU-3' • Upstream of the initiation codon in prokaryotic mRNA • To position the ribosome for initiation of protein synthesis Shine-Delgarno element Polysomes • Each mRNA transcript is read simultaneously by more than one ribosome. • A second, third, fourth, etc. ribosome starts to read the mRNA transcript before the first ribosome has completed the synthesis of one polypeptide chain. • Multiple ribosomes on a single mRNA transcript are called polyribosomes or polysomes. • Multiple ribosomes can not be positioned closer than 80 nt. Polysomes (多聚核糖体) • Electron micrographs of ribosomes actively engaged in protein synthesis revealed by "beads on a string" appearance. 核糖体是蛋白质合成的部位 • 放射性同位素标记氨基酸注射小鼠,取肝脏制备 亚细胞器,发现微粒体中放射性强度最高,证明 核糖体是蛋白质合成的场所。 • 核糖体存在的形式: 基本类型 附着核糖体 游离核糖体 70S的核糖体 80S的核糖体 主要成分 r蛋白质:40%,核糖体表面 rRNA:60%,,核糖体内部 11-14 1 rRNA • 小亚基: 16s RNA在识别mRNA上的多肽合成起始位 点时起重要作用;SD sequence. • 大亚基: 23s RNA存在一段与tRNAMet序列互补的 片段。在23s RNA靠近5‘’端处,有一段序列(143 -157nt)与5s RNA(72-83)结合。 • 5s RNA有两个保守区,一个有保守序列CGAAC,与 tRNA上TC环相互作用。另一个保守序列与23s RNA互补。这是小亚基和大亚基的相互作用。 11-15 核糖体小亚单位rRNA (a) E.coli 16S rRNA;(红色为高度保守区) (b) 酵母菌18S rRNA,它们都具有类似的40个臂环结构(图中1~40),其长度和位 置往往非常保守;P、E分别代表仅在原核或真核细胞存在的rRNA的二级结构。 (Darnell et al.,1990) 2 核糖体蛋白 • 原核生物 • 30s 小亚基有21种蛋白,16s rRNA; • 50s 大亚基有36种蛋白,5s和23s rRNA; • 真核生物 • 40s 小亚基有33种蛋白,18s rRNA; • 60s 大亚基有49种蛋白,5s,5.8s, 28s rRNA; 11-17 原核生物与真核生物核糖体成分的比较 11-18 3 核糖核蛋白复合体 E.coli (a)核糖体小亚单位中的部分r蛋白与rRNA的结合位点) (b)及其在小亚单位上的部位 (引自Albert et al.,1989,图a; Lewin,1997,图b 11-19 4 原核生物的核糖体 11-20 5 真核生物的核糖 体 11-21 Initiator tRNA • Methionine (甲硫氨酸)is the first amino acids incorporated into a protein chain in both prokaryotes (modified to N-formylmethionine) and eukaryotes. • Initiator tRNAs are special tRNAs recognizing the AUG (GUG) start codons in prokaryotes and eukaryotes. • Initiator tRNAs differ from the one that inserts internal Met residues. Initiator tRNA, fMet-tRNAfMet in E. coli Lacking alkylated A endorses more flexibility in recognition in base pairing (both AUG and GUG). Initiator tRNA formation in E. coli 1. Both initiator tRNA and noninitiator tRNAmet are charged with Met by the same methionyltRNA synthetase to give the methionyl-tRNA 2. Only the initiator methionyl-tRNA is modified by transformylase to give N-formylmethionyltRNAfmet. First step: ATP links to AA + H - P P P H3N C COO R + HO H3N C C-O- P O + OH OH Amino acid R AARS OH OH ATP Second step: Aminoacyl-AMP reacts with tRNA + HO H3N C C-O- P Aminoacyl-AMP O O + R OH OH OH OH Aminoacyl-AMP O tRNA P P AARS O + O C C OH O + H 3N C H R Aminoacyl-tRNA P O OH OH AMP 原核细胞中起始氨基酸活化后,还要甲酰化, 形成甲酰蛋氨酸tRNA,由N10甲酰四氢叶酸提 供甲酰基。而真核细胞没有此过程。 运载同一种氨基酸的一组不同tRNA称为同 功tRNA。一组同功tRNA由同一种氨酰基tRNA合 成酶催化。氨基酰tRNA合成酶对tRNA和氨基酸 两者具有专一性,它对氨基酸的识别特异性很 高,而对tRNA识别的特异性较低。 8.2: Mechanism of protein synthesis (Prokaryote) Protein synthesis falls into three stages . 1.initiation-the assembly of a ribosome on an mRNA molecule. 2.elongation-repeated cycles of amino acid addition. 3.termination-the release of the new protein chain. Initiation In prokaryotes, initiation requires • the large and small ribosome subunits, • the mRNA • the initiator tRNA • three initiation factors . Size comparisons show that the ribosome is large enough to bind tRNAs and mRNA. IF1 and IF3 bind to a free 30S subunits. IF2 complexed with GTP then bind to the small subunits, forming a complex at RBS. The initiator tRNA can then bind to the complex at the P site paired with AUG codon. 30S initiation complex The 50S subunits can now bind. GTP is then hydrolyzed and IFs are released to give the 70S initiation complex The assembled ribosome has two tRNA-binding sites, which are called A- and P-site, for aminoacyl (氨酰基位点) and peptidyl sites(肽酰基 位点) respectively. Only fMet-tRNAfMet can be used for initiation by 30S subunits; all other aminoacyl-tRNAs are used for elongation by 70S ribosomes. Elongation With the formation of the 70S initiation complex, the elongation cycle can begin. Elongation involves the three factors, EFTu, EF-Ts, EF-G, as well as GTP, charged tRNA and the 70S initiation complex. The three steps of elongation 1.Charged tRNA is delivered as a complex with EF-Tu and GTP .(进位反应) 2.Peptidyl tranferase (肽酰基转移酶)(50S ribosomal subunit) makes a peptide bond by joining the two adjacent amino acid without the input of more energy. (转肽反应) 3.Translocase (EF-G), with the energy from GTP, moves the ribosome one codon along the mRNA, ejecting the uncharged tRNA and transferred the ribosome peptide from the mRNA. (移位反应) EF-Tu-Ts exchange cycle Peptide bond formation takes place by reaction between the polypeptide of peptidyl-tRNA in the P site and the amino acid of aminoacyl-tRNA in the A site. Translocation • In bacteria, the discharged tRNA leaves the ribosome via another site, the E site. • In eukaryotes, the discharged tRNA is expelled directly into the cytosol. • EF-G (translocase) and GTP binds to the ribosome, and the discharged tRNA is ejected from the P-site in an energy consuming step. • the peptigly-tRNA is moved from A-site to Psite and mRNA moves by one codon relative to the ribosome P-site E-site A-site Translocation in E. coli Termination Protein factors called release factors interact with stop codon and cause release of completed polypeptide chain. RF1 and RF2 recognizes the stop codon with the help of RF3 The release factors make peptidyl transferase transfer the polypeptide to water, and thus the protein is released Release factors and EF-G: remove the uncharged tRNA and release the mRNA,. 8.3: Initiation in eukaryotes Most of the differences in the mechanism of protein between prokaryotes and eukaryotes occur in the initiation stage, where a greater numbers of eIFs and a scanning process are involed in eukaryotes. The eukaryotic initiator tRNA does not become N-formylated. 真核细胞蛋白质合成的起始 真核细胞蛋白质合成起始复合物的形 成中需要更多的起始因子参与,因此起 始过程也更复杂。 (1)需要特异的起始tRNA即,tRNAfmet,并且不需要N端甲酰化。已发 现的真核起始因子有近10种(eukaryote Initiation factor,eIF) (2)起始复合物形成在mRNA5’端AUG 上游的帽子结构,(除某些病毒mRNA外 ) (3)ATP水解为ADP供给mRNA结合所需 要的能量。真核细胞起始复合物的形成 过程是:翻译起始也是由eIF-3结合在 40S小亚基上而促进80S核糖体解离出60S 大亚基开始,同时eIF-2在辅eIF-2作用 下,与Met-tRNAfmet及GTP结合,再通过 eIF-3及eIF-4C的作用,先结合到40S小 亚基,然后再与mRNA结合。 mRNA结合到40S小亚基时,除了eIF-3参 加外,还需要eIF-1、eIF-4A及eIF-4B并 由ATP小解为ADP及Pi来供能,通过帽结 合因子与mRNA的帽结合而转移到小亚基 上。但是在mRNA5’端并未发现能与小亚基 18SRNA配对的S-D序列。目前认为通过帽 结合后,mRNA在小亚基上向下游移动而 进行扫描,可使mRNA上的起始密码AUG在 Met-tRNAfmet的反密码位置固定下来, 进行翻译起始。 prokaryotic Initiation factor IF1 IF3 IF2 Elongation factor EF-Tu EF-Ts EF-g eukaryotic function eIF3 eIF4c eIF6 eIF4B eIF4F eIF2B eIF2 eIF5 Bind to ribosome submits Bind to mRNA Initiator tRNA delivery Displacement of other factors eEF1α eEF1βγ eEF2 Aminoacyl tRNA delivery Recycling of EF-Tu or eEF1α Translocation Termination factors RF1, RF2, RF3 Polypeptides Chain release eRF Scanning The eukaryotic 40s ribosome submit complex bind to the 5’cap region of the mRNA and moves along it scanning for an AUG start codon. Eukaryotic ribosomes migrate from the 5’ end of mRNA to the ribosome binding site, which includes an AUG initiation codon. Initiation In contrast to the events in prokaryotes, initiation involves the initiation tRNA binding to the 40S subuits before it can bind to the mRNA. Phosphorylation of eIf2, which delivers the initiation tRNA, is an important control point. The initiation factor can be grouped to there function as follow Binding to ribosomal subunits eIF6 eIF3 eIF4c Binding to the mRNA eIF4B eIF4F eIF4A eIF4E Involved in initiation tRNA delivery eIF2 eIF2B Displace other factors eIF5 Initiator eIF3+4C+ tRNA+eIF2+GTP 40S Ternary complex + 43S ribosome complex 43S preinitiation complex ATP ADP+Pi +mRNA+eIF4F +eIF4B 48S preinitiation complex Scanning More factors involved Scanning to find AUG Elongation The protein synthesis elongation cycle in prokaryotes and eukaryotes is quite similar. The factors EF-Tu EF-Ts EF-G have direct eukaryotic equivalents called eEF1α eEF1βγ eEF2 Termination Eukaryotes use only one release factors eRF, which requires GTP,recognize all three termination codons. Termination codon is one of three (UAG, UAA, UGA) that causes protein synthesis to terminate. 8.4: Translational control and post-translational events • • • • • Translational control Polyproteins Protein targeting Protein modification Protein degradation Translational control • In prokaryotes, the level of translation of different cistrons can be affected by: (a) the binding of short antisense molecules, (b) the relative stability to nucleases of parts of the polycistronic mRNA , (c) the binding of proteins that prevent ribosome access. In eukaryotes, 1. protein binding can also mask the mRNA and prevent translation, 2. repeats of the sequence 5'-AUUUA -3' can make the mRNA unstable and less frequently translated. Polyprotein • A single translation product that is cleaved to generate two or more separate proteins is called a polyprotein. Many viruses produce polyprotein. Protein targeting • The ultimate cellular location of proteins is often determined by specific, relatively short amino acid sequence within the proteins themselves. These sequences can be responsible for proteins being secreted, imported into the nucleus or targeted to other organelles. Prokaryotic protein targeting: secretion Eukaryotic protein targeting • Targeting in eukaryotes is necessarily more complex due to the multitude of internal compartments: • There are two basic forms of targeting pathways 1. 2. The secretory pathway in eukaryotes (co-translational targeting) • The signal sequence of secreted proteins causes the translating ribosome to bind factors that make the ribosome dock with a membrane and transfer the protein through the membrane as it is synthesized. Usually the signal sequence is then cleaved off by signal peptidase. Protein modification • Cleavage: – To remove signal peptide – To release mature fragments from polyproteins – To remove internal peptide as well as trimming both Nand C-termini • Covalent modification: – Acetylation; – Hydroxylation; – Phosphorylation; – Methylation; – Glycosylation; – Addition of nucleotides. Phosphorylation Protein degradation • Different proteins have very different halflives. Regulatory proteins tend to turn over rapidly and cells must be able to dispose of faulty and damaged proteins. Protein degradation: process Faulty and damaged proteins are attached to ubiquitins (ubiquitinylation). The ubiquitinylated protein is digested by a 26S protease complex (proteasome) in a reaction that requires ATP and releases intact ubiquitin for re-use. • In eukaryotes, it has been discovered that the N-terminal residue plays a critical role in inherent stability. – 8 N-terminal aa correlate with stability: Ala Cys Gly Met Pro Ser Thr Val – 8 N-terminal aa correlate with short t1/2: Arg His Ile Leu Lys Phe Trp Tyr – 4 N-terminal aa destabilizing following chemical modification: Asn Asp Gln Glu