* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download DNA methylation
Zinc finger nuclease wikipedia , lookup
Genome evolution wikipedia , lookup
Human genome wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Genealogical DNA test wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Neocentromere wikipedia , lookup
Genomic library wikipedia , lookup
Oncogenomics wikipedia , lookup
Point mutation wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Transgenerational epigenetic inheritance wikipedia , lookup
Molecular cloning wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Epigenetic clock wikipedia , lookup
DNA vaccination wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Epigenetics of depression wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
DNA supercoil wikipedia , lookup
Genomic imprinting wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Non-coding DNA wikipedia , lookup
Deoxyribozyme wikipedia , lookup
X-inactivation wikipedia , lookup
DNA methylation wikipedia , lookup
Primary transcript wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Behavioral epigenetics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
History of genetic engineering wikipedia , lookup
Helitron (biology) wikipedia , lookup
Histone acetyltransferase wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Epigenetics wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
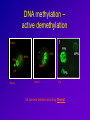
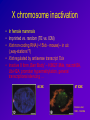
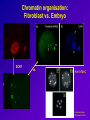
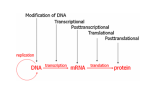
Epigenetic regulation during early embryogenesis – reprogramming and remodelling Helena Fulkova Istitute of Animal Science, www.vuzv.cz [email protected] Basic facts • No loss/gain of genomic DNA during development and differentiation • Somatic cells possess full developmental potential – demonstrated by SCNT •Organisation of DNA in nucleus is nonrandom and heritable •Gene expression is not reset every cell division → A mechanism that is flexible but heritably regulates gene expression and nuclear organisation … DNA + proteins = chromatin This cannot simply be achived by different TFs! What is epigenetic inheritance? A heritable change in gene expression that is not caused by changes in the DNA seguence → DNA methylation → Histone post-translational modifications → Histone variants → non-histone protein composition of chromatin → chromatin remodelling complexes → (RNA – antisense transcripts…) • • • • • • Role in gene expression Differentiation Development Genome integrity, nuclear organisation Recombination DNA repair … Histones • H2A, H2B, H3 and H4 (canonical core histones - octamer) +146bp DNA = nucleosome Histones II • • • • Basic proteins Conserved among Eukaryotes C- and N- terminal tails + globular domain Subjected to post-translational covalent modifications + H1 (linker histone) + other non-histone proteins→ chromatin Histone modifications • • • • Acetylation - HATs/HDACs methylation - HMTs/demethylases Phosphotylation – kinases/phosphatases Ubiquitination… Activating vs. repressive modifications • • Methylation • H3/K9, H3/K27, H4/K20 … repression of transcription and silencing H3/K4 … activation • • Acetylation • • Phoshorylation • • • Ubiquitination • … H3/K9, H4/K16, H4/K12, H4/K8 and H4/K5 … activation H4 acetylation … DS break repair H3/S10 … active transcription, chromosome condensation H2AX/S139 … DS breaks repair • Ubiquitin H2A … X chromosome silencing The histone code hypothesis Histone variants • H3 • H2A Canonical core histone H3.3 CENP A H2AX H2AZ macroH2A H2ABBD • H2B • H4 Transcriptional activation Kinetochore assembly Canonical core histone DNA repair and recombination Gene expression X chromosome inactivation Transcriptional activation? Canonical core histone Canonical core histone Maintaining histone modifications over DNA replication phase Chromatin • Euchromatin – active histone modifications, low DNA methylation … • Heterochromatin – Constitutive – repressive histone modifications, high DNA methylation, specific histone variants – Facultative – repressive histone modifications, high DNA methylation DNA methylation • Cytosine 5-methylcytosine • In CpG dinucleotides (exception: most CpG islands) • Highly mutagenic: mCT • Repression of transcription, mobile element silencing, host defence, genomic imprinting, genome stability … • Genome instability and global hypomethylation is linked to diseases such as cancer DNA methylation – DNA methyltransferases • Dnmt1 • Dnmt1o, p, s isoform (maintenance function) • Dnmt3 family (de novo methylation) • Dnmt2 • Dnmt 3a • Dnmt 3b • Dnmt 3L N-terminal regulatory domain C-terminal catalytic domain Substrate: S-AdenosylMethionin Base flipping (like repair enzymes) De novo methylation vs. maintenance methylation Active vs. passive demethylation How do these systems intergate? • Adaptor proteins – chromodomain/bromodomain (TFs)… • Heterochromatin/Euchromatin (chromatin remodelling) • X chromosome inactivation • Change of gene expression (imprinting, development associated genes – Oct4, Nanog…) Adaptor proteins e.g. Methyl CpG binding proteins: MBD1 …transcriptional repression (not • MBD 1-4 in MeCP1 complex) MBD2 …associated with HDAC1 repression (found in MeCP1, Sin3, Mi2/NURD complexes) MBD3 …species specific binding of methylated/nonmethylated DNA (found in Mi2) MBD4 …repair?, can induce nicks, has glycosylase activity, binds TpG (deamination of 5-MeC) • MeCP2 Rett syndromeX linked, predomimantly ♀ HP1 Targeting: methylated H3/K9 (triMe) • HP1 alpha • Centromeric heterochromatin • HP1 beta • Pericentric heterochromatin, but also euchromatin ? • HP1 gamma • Euchromatin, active genes? Chromatin remodelling complexes • ATP dependent Three classes: SWI/SNF-like ISWI-like CHD-like All three possess in vitro nucleosome remodelling activity in the absence of associated subunits (2-12 in the complex) Composition of remodelling complexes Swi/Snf family CHD family Possible mechanism • Trans-transfer of nucleosomes (SWI/SNF) • Cis-transfer = nucleosome sliding (ISWI) • generating superhelical torsion (SWI/SNF, ISWI, Mi2) • Generating loops (RSF – a member of ISWI family; SWI/SNF binds DNA in two positions) Snf5p subunit of SWI/SNF can interact with H2B; SWR1 complex (Swr1p) directs specifity towards H2A.Z Other complexes contain histone chaperones (ACF and CHRAC – NAP1) Epigenetics and development Critical developmental time points: • Fertilization: gametes vs. zygote • Differentiation: ICM vs. TE (~ 3,5 dpc), stem cells vs. specific cell types • PGC establishment and migration (~ 10 dpc) Epigenetic „life cycle“ PGCs TE Embryo ICM + placenta Fertilization Gametes – terminally differentiated and highly specialized Genomes - (oocyte, sperm) transcriptionally inactive after fertilization •One cell stage embryo is totipotent •Fertilization represents RADICAL REPROGRAMMING OF BOTH GENOMES Post-fertilization epigenetic remodelling ♂ ♀ •Protamines are exchanged for histones •DNA is demethylated •High histone acetylation, methylated histones are rare Histones are highly methylated DNA is highly methylated Acetylation of histones rises more slowly DNA methylation – active demethylation 5-MeC FPN MPN FPN MPN PB Mouse Human Pig All species tested excluding Sheep! Histone modifications • Asymmetric distribution • Symmetric distribution • Associated with specific DNA sequences (chromatin structures) What happens next – passive demethylation Differentiation • First differentiation – Blastocyst (ICM vs. TE) • Pluripotent stem cells → specific cell types • X chromosome inactivation • Associated with the change of gene expression (silencing of Oct4, Nanog, Sox2 by DNA methylation…+ activation of tissue specific genes) X chromosome inactivation • In female mammals • Imprinted vs. random (TE vs. ICM) • Xist non-coding RNA (~15kb - mouse)– in cis („way-stations“?) • Xist regulated by antisense transcript Tsix • Inactive X form „Barr Body“ – H3K27-3Me, macroH2A, Ub-H2A, promoter hypermethylation, general transcriptional silencing … 46 XX 47 XXX Clemson et al, 1996, J Cell Biol Bivalent domains • Regions that contain both activating and repressive histone marks (prior to differentiation) • H3/K4-3Me + H3/K27-3Me • Typical for tissue-specific genes with CpG island promoters (PRC targets) • Upon differentiation the modifications stabilize to either active or repressive state Primordial Germ Cells (PGCs) • Epiblast (embryonic ectoderm) – 7.25 dpc • Expression of Stella and Fragilis • Oct4 and other pluripotency-associated genes (AP…) • Migration to genital rigdes • Erasure of imprinting and establishment according to the sex of the embryo (8.5 -10.5-11.5 dpc), X chromosome reactivation Imprinting • Gene expression according to parental origin • Localized in clusters • +/- 80 genes (predicted 150) • Expression based on methylation of parental alleles (DMRs/DMDs) • Growth of embryonic/extraembryonic tissues •Human syndromes (Prader-Willi/Angelman, Silver-Russel) – mostly mental retardation •Higher frequency in ART conceived children! Epigenetics and SCNT • Chromatin organisation maintained (according to donor cell) • Initial global demethylation, abnormal remethylation (aberrant Dnmt1 localization) • Imprinted genes dysregulated • X chromosome initially reactivated, later on mosaicism Chromatin organisation: Fibroblast vs. Embryo SCNT vs. Anti 5-MeC Santos and Dean, Reproduction 2004 Consequence • • • • Low SCNT efficiency LOS (Igf2?) Placental hyperplasia Death soon after birth …TSA, 5-AzaC – better efficiency …ESCs better donors for SCNT Methods DNA methylation: – – – – Immunofluorescence Methylation sensitive restriction Bisulfite sequencing COBRA Histone modifications: – Immunofluorescence – Chromatin IP DNA methylation • Convertion of DNA by sodium bisulfite: 5-MeC unreactive, C → U • PCR: 5-MeC(C)/G and U(T)/A, Sequence and compare differences • COBRA – sequence differences based restriction Main problems • PCR is often biased towards unmethylated teplate!!! • Primer design is not easy • High quality DNA required …OK • DNA degraded during treatment, most template unconverted …but if working – lots of information! Chromatin IP - ChIP • Crosslink of DNA and associated proteins • Antibody against desired protein – pulldown of all fragments crosslinked • PCR with primers for certain region • Or microarray experiment Main problems • Usually HIGH amount of starting material required!!! • Good antibody necessary • Modification – „Carrier ChIP“ (starting material still high, radioactively labelled primers …) Recommended literature • Epigenetics, Ed. Allis, Jenuwein, Reinberg, Cold Spring Harbor Press, 2006 • Gene Expression at the Beginning of Animal Development, Ed. DePamphilis, Elsevier, 2002 Thank you for your attention!