* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Lecture 33
Transcriptional regulation wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Lipid signaling wikipedia , lookup
Silencer (genetics) wikipedia , lookup
Eukaryotic transcription wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
RNA polymerase II holoenzyme wikipedia , lookup
Magnesium transporter wikipedia , lookup
Citric acid cycle wikipedia , lookup
Ribosomally synthesized and post-translationally modified peptides wikipedia , lookup
Expression vector wikipedia , lookup
Peptide synthesis wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Ancestral sequence reconstruction wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Interactome wikipedia , lookup
Point mutation wikipedia , lookup
Western blot wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Metalloprotein wikipedia , lookup
Messenger RNA wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Biochemistry wikipedia , lookup
Protein purification wikipedia , lookup
Protein structure prediction wikipedia , lookup
Gene expression wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Proteolysis wikipedia , lookup
De novo protein synthesis theory of memory formation wikipedia , lookup
Genetic code wikipedia , lookup
Biosynthesis wikipedia , lookup
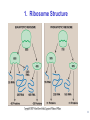
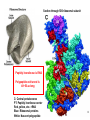
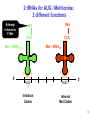
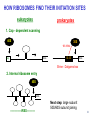
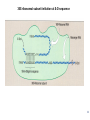
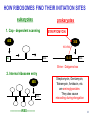
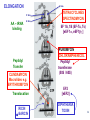
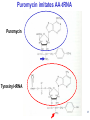
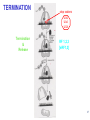
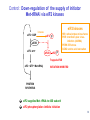
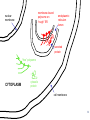
LecLtures 33 and 34 Lectures 33-34 GENETIC CODE and PROTEIN SYNTHESIS Mukund Modak, Ph.D. Adapted from . M. Mathews, Ph.D. 1 Proteins are important… ~44% of the dry wt. of the human body. ~5% of human caloric intake goes for protein synthesis. catalyze most of the reactions in living organisms. serve many roles (enzymatic, structural, transport, regulation, ...) …in sickness and in health protein synthesis is tightly regulated by environmental stimuli as well as intrinsic processes (e.g., hormonal, developmental). dysregulation can cause disease. many antibiotics act at the level of protein synthesis. 2 2 I. INTRODUCTION Central Dogma Ribosomes and polysomes Genetic Code Mutations with effects at the translation level II. TRANSLATIONAL MACHINERY III. MECHANISM OF TRANSLATION AND INHIBITORS OF PROTEIN SYNTHESIS IV. ENERGETICS AND REGULATION OF TRANSLATION 3 3 POLYSOMES E.M. 4 CENTRAL DOGMA DNA RNA PROTEIN The central dogma states that once “information” has passed into protein it cannot get out again. The transfer of information from nucleic acid to nucleic acid, or from nucleic acid to protein, may be possible, but transfer from protein to protein, or from protein to nucleic acid is impossible. Information means here the precise determination of sequence, either of bases in the nucleic acid or of amino acid residues in the protein. Francis Crick, 1958 5’ N- or aminoterminus RNA protein 3’ C- or carboxyterminus 5 Coupled transcription & translation in bacteria [ N terminus to [ 5’ to 3’ ] C terminus ] Not so in Eukaryotes 6 1st position (5’ end) GENETIC CODE U C A G 2nd position 3rd position (3’ end) U C A Phe Phe Leu Leu Ser Ser Ser Ser Tyr Tyr STOP STOP Cys Cys STOP Trp U C A G Leu Leu Leu Leu Pro Pro Pro Pro His HIs Gln Gln Arg Arg Arg Arg U C A G Ile Ile Ile Met Thr Thr Thr Thr Asn Asn Lys Lys Ser Ser Arg Arg U C A G Val Val Val Val Ala Ala Ala Ala Asp Asp Glu Glu Gly Gly Gly Gly U C A G codon # 5 6 7 CCU GAG GAG Pro Glu Glu CCU GUG GAG Pro Val Glu G Normal Hb – β Sickle cell Hb – βS 7 GENETIC CODE: Co-linear triplet code Nearly universal – variations in mitochondria, mycoplasma, ciliates Degenerate (or redundant) Non-overlapping Unpunctuated – although some codons are signals Mutations - in coding region can cause various ill-effects, such as, change in desired amino acids, early or late stop, insertion, etc. 8 I. INTRODUCTION II. TRANSLATIONAL MACHINERY Ribosomes: prokaryotic / eukaryotic Messenger RNA Transfer RNA Aminoacyl-tRNA synthetases; Met-tRNA forms (m, f, i) Initiation, elongation and termination enzymes III. MECHANISM OF TRANSLATION AND INHIBITORS OF PROTEIN SYNTHESIS IV. ENERGETICS AND REGULATION OF TRANSLATION 9 9 TRANSLATIONAL COMPONENTS 1. Ribosomes (large and small subunits) 2. Messenger RNA (mRNA) 3. Transfer RNAs (tRNAs) 4. Amino Acids (aa’s) 5. Enzymes (“factors”) 6. Energy (ATP, GTP) 10 1. Ribosome Structure 11 Section through 50S ribosomal subunit Peptidyl transferase is RNA Polypeptide exit tunnel is 40~50 aa long C: Central protuberance PT: Peptidyl tranferase center Red, yellow, etc.: rRNA Blue: Ribosomal proteins White: Nascent polypeptide 12 12 2. mRNA Eukaryotic: Monocistronic (spliced) 5’ end ( 1 coding region ) 5’ UTR cap 3’ end 3’ UTR poly A AAA ~150 only 1 7-MeGpppGXY Cistron = coding region = open reading frame (ORF) Prokaryotic: Polycistronic ( >1 coding region ) 5’ ppp #1 #2 3’ #3 13 3. tRNA Translational Adaptor 14 4. Amino Acids tRNAs carry “activated” amino acids: aaRS (1) AA + tRNA + ATP AA ~tRNA + AMP + PPi G ~0 Kcal/mole PPase (2) PPi + H2O 2 Pi G = -6.6 Kcal/mole Overall free energy change for aminoacylation of tRNA G ~ -6.6 Kcal/mole aaRS = aminoacyl-tRNA synthetase PPase = pyrophosphatase 15 Formation of aminoacyl-tRNA The amino acid is first activated by reacting with ATP The activated amino acid is transferred from aminoacyl-AMP to tRNA These enzymes are vital for the fidelity of protein synthesis: 2 steps allow “proofreading” 16 Genetic Code 20 AA’s Translation Machinery 20 AA – tRNA synthases ( i.e., 1 per AA ) 61 Codons for AA’s ~50 tRNA species (at least 1 per AA, but less than 1 per codon) “WOBBLE” Pairing Wobble Position e.g. CUU 1 anti–codon anti-codon stem-loop of tRNA GAA GAG tRNA 3’ 2 codons 5’ 3 2 1 5’ 1 2 3 mRNA CODON ANTI-CODON 17 2 tRNAs for AUG / Methionine: 2 different functions N-formyl in bacteria: F-Met Met Met CCA CCA Met – tRNA F or I 3’ 5’ Met – tRNA M UAC AUG 1 5’ Initiation Codon 3’ UAC AUG 5’ 3’ Internal Met Codon 18 5. Translation Factors Enzymes Translation Step Prokaryotes Charging of tRNA Eukaryotes Aminoacyl – tRNA synthetases 1. Initiation IF1- IF3 eIF1- eIF5 (multiple) 2. Elongation EF1, EF2 eEF1, eEF2 3. Termination RF1- RF3 eRF1, eRF3 Modifications, cleavage, etc. 19 I. INTRODUCTION II. TRANSLATIONAL MACHINERY III. MECHANISM OF TRANSLATION AND INHIBITORS OF PROTEIN SYNTHESIS Initiation Elongation Termination Antibiotics Toxins IV. ENERGETICS AND REGULATION OF TRANSLATION 20 20 HOW RIBOSOMES FIND THEIR INITIATION SITES eukaryotes prokaryotes 1. Cap - dependent scanning 40S 30S 16S rRNA cap AUG... S-D AUG.. Shine - Dalgarno box 2. Internal ribosome entry 40S AUG.. ---------------IRES----------- Next step: large subunit 50S/60S subunit joining 21 30S ribosomal subunit initiation at S-D sequence 22 HOW RIBOSOMES FIND THEIR INITIATION SITES eukaryotes prokaryotes 1. Cap - dependent scanning STREPTOMYCIN 40S 30S 16S rRNA cap AUG.. S-D AUG.. Shine - Dalgarno box 2. Internal ribosome entry 40S AUG... ---------------IRES----------- Streptomycin, Gentamycin, Tobramycin, Amikacin, etc. are aminoglycosides. They also cause miscoding during elongation 23 ELONGATION A Site TETRACYCLINES SPECTINOMYCIN P Site AA – tRNA binding E Site EF 1A, 1B (EF-Tu, Ts) [eEF 1α, eEF1βγ ] PUROMYCIN CHLORAMPHENICOL Peptidyl transferase (50S / 60S) Peptidyl Transfer CLINDAMYCIN Macrolides e.g. ERYTHROMYCIN Translocation RICIN -SARCIN GTP EF2 [eEF2] DIPHTHERIA TOXIN 24 Puromycin imitates AA-tRNA Puromycin Tyrosinyl-tRNA 25 Inhibition of ribosome translocation 1) Diphtheria toxin inactivates eEF2 2) Erythromycin inhibits EF2 26 26 TERMINATION stop codons UAG UAA UGA Termination & Release RF 1,2,3 [eRF1,3] 27 ENERGETICS OF PROTEIN SYNTHESIS 1. Charging ATP, 2~ 2. Initiation Unwinding and scanning Met-tRNAi binding ATP (several), 1~ GTP, 1~ Elongation AA-tRNA binding Translocation GTP, 1~ (see later) GTP, 1~ Termination GTP (number unknown), 1~ 3. 4. TOTAL: 4~ per AA polymerized + initiation + termination > 1200~ for an average protein Compared to 36-38 ATP’s generated by Glucose CO2 28 Down-regulation of the supply of initiator Met-tRNAi via eIF2 eIF2 • GDP eIF2B eIF2 • GTP eIF2 • GTP • Met-tRNAi PROTEIN SYNTHESIS eIF2 supplies Met- tRNAi to 40S subunit 29 Control : Down-regulation of the supply of initiator Met-tRNAi via eIF2 kinases eIF2 • GDP eIF2 kinases kinases eIF2 HRI: reticulocytes minus heme PKR: interferon plus virusinfection (dsRNA) PERK: ER stress GCN2: amino acid starvation P eIF2B eIF2 • GTP eIF2B eIF2 P Trapped eIF2B eIF2 • GTP • Met-tRNAi INITIATION INHIBITED PROTEIN SYNTHESIS eIF2 supplies Met- tRNAi to 40S subunit eIF2 phosphorylation inhibits initiation 30 GTP/GDP exchange during elongation by (e)EF1 (aka EF-Tu) Terminology EF-Tu • GDP EF-Ts EF-Tu • GTP PROK. Old New EUK. aa-tRNA complex Tu 1A 1α GEF Ts 1B 1βγ EF-Tu • GTP • aa-tRNA PROTEIN SYNTHESIS This factor supplies aa- tRNA to ribosome during elongation. 31 membrane-bound polysome on “rough” ER nuclear membrane endoplasmic reticulum lumen secreted protein “free” polysome CYTOPLASM cytosolic protein cell membrane 32 Inhibitors of Protein Synthesis: Antibiotics and Toxins Class Aminoglycosides Target 30S Tetracylines 30S 50S 50S Action (1) Inhibits initiation (2) Causes misreading Inhibits binding of AA-tRNA to A-site Inhibits peptidyl transferase Inhibit translocation 50S Ile-tRNA synthase Inhibit translocation Inhibits isoleucine tRNA charging PUROMYCIN 50S, 60S Premature release of nascent polypeptide Cycloheximide DIPHTHERIA TOXIN RICIN (castor beans) -Sarcin (fungus) 80S eEF2 60S 60S Inhibits translocation Inhibits translocation¤ Inhibits binding of AA-tRNA to A-site♦ Inhibits binding of AA-tRNA to A-site & translocation# Inhibitor STREPTOMYCIN, Gentamicin, Kanamycin, Neomycin, etc. TETRACYCLINE, doxycycline CHLORAMPHENICOL ERYTHROMYCIN, Clarithromycin, Azithromycin Clindamycin, Lincomycin Mupirocin (pseudomonic acid) Macrolides Lincosamides CAPITALIZED: most important Catalytic activities of toxins ¤ ADP ribosylation ♦ 28S rRNA depurination (A) # 28S rRNA cleavage 33 PROKARYOTES EUKARYOTES Nucleus No Transcription & translation Coupled Yes Separated mRNA Polycistronic Ribosomes 70S (50S, 30S) Monocistronic, Capped & Polyadenylated 80S (60S, 40S) Initiator f Met – tRNAi Met – tRNAi Site selection Shine-Dalgarno mediated internal initiation 1) Scanning 2) IRES mediated internal entry Initiation factors 3 >12 Order of events 1) mRNA binding 2) f Met – tRNAi binding 1) Met – tRNAi binding 2) mRNA binding Antibiotics Sensitive Resistant Toxins Resistant Sensitive 34 Protein Modifications 1. Phosphorylation - (Tyr, Ser,Threo) Metabolic Regulation, Signal transduction, etc 2. Hydroxylation - (Proline) in collagen, Endoplasmic Reticulum 3. Glycosylation – (O-linked as with Ser/Threo- OH or N-Linked as in lysine) 4. Other - biotinilation, farnesyl, etc Protein Degradation - Mostly thru specific proteases and ubiquitin-proteosome system 35