* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download lecture 20 notes
Comparative genomic hybridization wikipedia , lookup
Gene expression programming wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Gene expression profiling wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Genealogical DNA test wikipedia , lookup
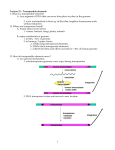
Nutriepigenomics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
DNA vaccination wikipedia , lookup
Genomic imprinting wikipedia , lookup
Molecular cloning wikipedia , lookup
Epigenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
X-inactivation wikipedia , lookup
DNA supercoil wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Primary transcript wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Copy-number variation wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Genetic engineering wikipedia , lookup
Pathogenomics wikipedia , lookup
Oncogenomics wikipedia , lookup
Point mutation wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome (book) wikipedia , lookup
Microsatellite wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Designer baby wikipedia , lookup
Human genome wikipedia , lookup
Human Genome Project wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genomic library wikipedia , lookup
Minimal genome wikipedia , lookup
History of genetic engineering wikipedia , lookup
Transposable element wikipedia , lookup
Genome editing wikipedia , lookup
Business • Helpful video on translocations: https://www.youtube.com/watch?v=WqHEndYJEkQ • Homework due WEDNESDAY because of holiday • Practice problems and sample exam on web site 1 Roadmap • One more deeply weird amphibian • Selfish DNA: – Repeated sequences – Tranposable elements (transposons) 2 Amphibian weirdness • Pelophylax esculenta appears to be a species of water frog • Originally a hybrid of P. ridibundus and P. lessonae • It prefers to mate with P. lessonae • Why doesn’t it get diluted out? Pelophylax esculenta (Rana esculenta) Pelophylax ridibundus (Rana ridibunda) 3 Pelophylax lessonae (Rana lessonae) 4 Sexual and asexual lineages in the same species • Like the triploid toads, P. esculenta has: – A sexual, recombining part – An asexual, nonrecombining part • Will the asexual parts deteriorate over time? • No examples known of diverse, old groups with genomes like this 5 Selfish DNA • How can a gene be “selfish”? • Kinds of selfish DNA – Meiotic drive (previously discussed) – Repeats – Transposons 6 Selfish DNA • Regular genes: – Replicate only when genome replicates – Increase their fitness by increasing organism’s fitness – “Motivated” to work well with the rest of the genome • Selfish genes (or genome regions): – Replicate independently, or otherwise control their own reproduction (t chromosome) – Can increase their fitness without increasing organism’s fitness, by making more copies of themselves – Gene’s fitness is less dependent on working well with the rest of the genome 7 Examples of potentially selfish DNA • Repetitive elements (microsatellites, etc) • Transposons • Meiotic drive loci (t chromosome) 8 “Satellite DNA” • Many repetitions of the same short sequence • In kangaroo rats, half the genome is AAG, TTAGGG, CAACAGCGCGGG repeats • First detected as peaks (“satellites”) in DNA density • Different GC content than the rest of the genome • Small localized groups of repeats are “microsatellites” 9 Satellite DNA: localized repeats • Regions with satellite DNA are frequently in a condensed state called heterochromatin • Most active genes are inactivated if they are moved into heterochromatin • A few genes can remain active there • Drosophila melanogaster heterochromatin patterns: Mb 23.0 21.4 Mb 5.4 11.0 2R Chromosome 2L 24.4 28.0 21.8 20.0 1.2 X 3L 8.2 8.2 Y 3R 40.9 Heterochromatin Euchromatin Centromere X and Y 10 3.1 4 How does repeated DNA increase? • Hard to duplicate repeats accurately: – Replication slippage (same area gets replicated twice) – Unequal crossing over • Genetic drift can then cause copy number to increase or decrease • Microsatellite rate of copy number change up to 10−4 per meiosis 11 How does repeated DNA increase? • Number of repeats cannot decrease below 1 by these mechanisms • Mathematically, a random walk with a barrier in only one direction will tend to move away from the barrier • This might be kept in check by selection against the repeat: – DNA replication is expensive – Cells with lots of DNA divide slower and are larger – One repeat more or less makes little difference! 12 Function vs selfishness • Does highly repeated DNA have a function? It might be involved in: – Chromosome physical structure–spindle fiber attachment, etc – Recognition of homologous centromeres – Chromosome pairing • Possible evidence for selfishness: – Fitness in Drosophila is not reduced when heterochromatin is reduced – Heterochromatin content varies widely between species • Drosophila nasutoides has about 60% satellite DNA, but all on one chromosome. • Drosophila littoralis, D. ezoana have no satellite DNA 13 Mobile elements • Transposons can move or copy themselves into new genomic locations • This can inactivate or truncate genes • It can separate a gene from its usual promoter • It can put a gene under control of the transposon promoter 14 Mobile elements • Conservative transposition: – transposable element leaves its original position and moves to a new position – because of chromosome reassortment, can increase or decrease genomic copy number – can also increase copy number if transposition happens during DNA replication (transposon replicates, moves, and replicates again) – causes mutations which may cleanly revert when transposon leaves – often a small duplication is left at the site • Typical of most DNA transposons 15 Mobile elements • Replicative transposition: – – – – transposon stays where it is; a new copy inserts elsewhere increases copy number causes mutations which do not easily revert this can happen via DNA copying or via DNA to RNA reverse transcription – also tends to cause a small duplication at the site • RNA transposons (retrotransposons) and some DNA transposons 16 Heavy spotting shows activity of Mu transposon 17 Transposons as genome components • Many transposon families have both active and inactive members – Initial transposon codes for genes needed in transposition – A copy can lose these genes but use proteins from other copies to transpose – Eventually copies deteriorate and stop transposing • Within a genome, transposons evolve like organisms on their own • However, if they evolve in ways contrary to host survival they die 18 Transposons clog the human genome • LINE elements – – – – – – “Long interspersed elements” Retrotransposons which code for a reverse transcriptase Three families in humans (LINE1, LINE2, LINE3) LINE1 is still active and can move LINE2 and LINE3 have stopped transposing LINEs have about 850,000 copies in human genome or 21% of genome • SINE elements – – – – – “Short interspersed elements” Have lost the reverse transcriptase gene Can transpose only in presence of related LINE Alu family of SINEs is the biggest gene family in humans SINEs have about 1,500,000 copies or 13% of genome 19 Are retrotransposons viruses? Two hypotheses: • RNA virus from a retrotransposon: – Cell evolved reverse transcriptase for its own use – Transposon picked up a protein that could form a virus capsule and became a virus • Retrotransposon from an RNA virus: – Virus infected cell and then lost ability to leave – Reverse transcriptase now allows it to move within the cell • Both of these may have happened at different times to different viruses/transposons! 20 Drosophilia P element transposon • Found in wild D. melanogaster populations • Lab strains (started about 100 years ago) lack P • In wild flies: – No transposition in somatic cells – Rare transposition in germ line cells • In wild x lab crosses: – High transposition in all cells – “Hybrid dysgenesis”–flies are sick due to too much transposition 21 Drosophilia P element transposon Hypothesis: P element a new invasion of D. melanogaster • Very similar P found in a distant relative (60 million years’ separation) • Lab strains not only lack P, but lack suppressors to keep it from moving • P may have come in via rare cross-species mating • It spreads because it can move around in the genome Drosophilia P element transposon Transposition good or bad? • If P truly spread across the world in 100-200 years, it’s clearly good–for the transposon • P moving in somatic cells kills the flies • P moving at a high rate in germ cells is pretty bad too • P-bearing strains experience strong selection for silencing of P • P element can fight back by evolving immunity 23 Evolutionary effects of transposons–discussion • What would be different between a species with transposons and a similar one without? • Consider effects of: – Mutations – Similarity between different parts of the genome – Genome size 24 Thought problem • Cross P into a lab strain and maintain a population in bottles • Experimentally, population may live or die • Which is likely to do better, a large or small population? 25 Thought problem • Cross P into a lab strain and maintain a population in bottles • Experimentally, population may live or die • Which is likely to do better, a large or small population? • When this experiment has been done, large populations are more likely to survive 26 Genome shock hypothesis for transposons • Barbara McClintock hypothesized that transposons allow an organism to control its own mutation rate: – Healthy vigorous plants suppress transposition – Sickly plants allow transposition, leading to mutations which could possibly be useful – If you are sick already, you have little to lose and a lot to gain • Alternative hypothesis: transposons are purely selfish – Healthy vigorous plants suppress transposition – Sickly plants aren’t able to do so because they’re too sick, so they are vulnerable to transposition • Not easy to test these alternatives 27 One supportive observation • Copia transposons in flies carry a heat-shock promoter • Transposition is triggered when flies are stressed • Hard to distinguish between: – Heat-shock regulation of transposons helps the fly – Heat-shock regulation of transposons helps the transposon 28 Finding a use for transposons 29 Oxytricha genome rearrangement • Two nuclei per cell: – Micronucleus used for reproduction, but genes not active – Macronucleus expresses genes • Macronucleus genome is highly rearranged: – – – – – Cut into around 16,000 tiny chromosomes Mainly 1 gene per chromosome Genes are re-assembled from fragments Some are duplicated (dosage control?) 95% of germline genome is destroyed 30 Transposons harnessed to chew up genome • Germline genome of Oxytricha full of transposons • Macronucleus has none • If transposases are inactivated, the macronucleus fails to develop properly • Transposons and transposase probably central in rearrangement process 31 Other “tamed” genome rearrangements • Vertebrate rearrangement of antibody genes – Genome contains assortment of V, D and J segments – Targeted rearrangement makes a final antibody gene – This system greatly increases antibody diversity • Yeast sex switching – Genome contains inactive copies of a and α genes – Targeted gene conversion allows sex switching • Evolution can “use” elements which were initially selfish 32 One-minute responses • Tear off a half-sheet of paper • Write one line about the lecture: – Was anything unclear? – Did anything work particularly well? – What could be better? • Leave at the back on your way out 33