* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Problem Set 1A
Epigenetics in learning and memory wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Human genome wikipedia , lookup
Genomic imprinting wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
SNP genotyping wikipedia , lookup
Y chromosome wikipedia , lookup
Primary transcript wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Genetic engineering wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Holliday junction wikipedia , lookup
DNA vaccination wikipedia , lookup
Point mutation wikipedia , lookup
Genome (book) wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Epigenomics wikipedia , lookup
Gene expression programming wikipedia , lookup
Genome evolution wikipedia , lookup
Molecular cloning wikipedia , lookup
Genomic library wikipedia , lookup
Non-coding DNA wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Genome editing wikipedia , lookup
Microsatellite wikipedia , lookup
Homologous recombination wikipedia , lookup
X-inactivation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Helitron (biology) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Designer baby wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
History of genetic engineering wikipedia , lookup
Neocentromere wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Microevolution wikipedia , lookup
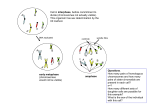
Problem Set 2B 9-26-06 Name and Lab Section:_____________________ 1. Define each of the following rearrangements (mutations) (use one phrase or sentence for each). Then describe what kind of chromosomal structure you might see in cells in meiotic prophase I if those cells are heterozygous for each of these rearrangements (one phrase or one sentence). If possible, give enough detail to distinguish each rearrangement from the others in your description of what you might see. A. deletion: The loss of a seqment of DNA. In prophase I a person might see a loop-out from one of the paired chromosomes (two of the four chromatids). B. duplication: A portion of a chromosome is duplicated, so its present twice. A person might see a loop-out that would look the same as in A above. (Note: it might not be possible to distinguish whether you are looking at a deletion or a duplication, just by looking at the paired chromosomes, unless there is a distinctive banding pattern.) C. inversion: The DNA sequences (or genes) in a portion of a chromosome have rearranged such that they are in the opposite order (run in the opposite direction) from what they originally were. A person might see a loop-out of the chromosomes, but in this case (as opposed to the situation with deletions or duplications) both chromosomes would be involved in the loop. Thus all four chromatids would be involved in the loop. D. translocation: Movement of genetic material between nonhomologous chromosomes. A person might see a cross structure formed that involves four chromosomes. 2. Draw a pair of homologous chromosomes as they would be matched during prophase I if they had a paracentric inversion. (be sure to give yourself plenty of space so drawing is clear) Let the original gene order on these chromosomes be: L, M, N, centromere, O, P, Q, R, S, T. Let one of the chromosomes contain an inversion that includes P, Q, and R. Remember that the chromosomes are duplicated at this time, and thus contain sister chromatids. (For the exam you should be able to do this for deletions and duplications too.) 3. Now assume that a crossover occurs between genes Q and R. Draw 4 possible chromosomes (with letters included) that might arise from this by the end of meiosis II (there might be one of each of these chromosomes in 4 different gametes). 1 Note that the break that occurred at anaphase I in the dicentric chromosome, could have occurred anywhere between the two centromeres. 4. Compare legitimate recombination to illegitimate recombination. Which is more common? Legitimate recombination is recombination between two DNA sequences that share regions of high similarity, as opposed to illegitimate recombination, which is recombination between two DNA sequences which share very little sequence similarity. Legitimate recombination is the most common. 5. Most cases of autosomal aneuploidy in humans involve the small chromosomes. Why is this? Since larger chromosomes contain more genes, aneuploidy in larger chromosomes will result in gene dosage problems with a larger number of genes, which is more likely to result in an embryo not even completing development. 6. Red-green color blindness is an X-linked recessive disorder. Bob has a 47,XXY karyotype (Klinefelter syndrome) and is color blind. His 46,XY brother, Tom, is also color blind. Both of their parents have normal color vision. Where did the nondisjunction occur that caused Bob to be color blind and have Klinefelter syndrome? Probably in meiosis II in their mother. The reasoning: Since Bob and Tom’s father is not color blind, the X-linked color blind allele must be carried on one of their mother’s X chromosomes. Tom got that color blind allele from one of his mother’s X chromosomes. Bob got two X’s from his mom, both of which were carrying the color blind allele, so those two X’s had to have come from nondisjunction in his mother, probably during meiosis II. Note that nondisjunction in meiosis I would give rise to a cell with a regular X and a “color-blind” X, which would have lead to Bob being a carrier with normal sight. A much less-likely possibility would be mitotic nondisjunction in the mother. 7. Why does recombination in the inversion regions appear to be suppressed in individuals heterozygous for paracentric and pericentric inversions? A crossover within a paracentric inversion produces a dicentric and an acentric recombinant chromatid. The acentric fragment is lost, and the dicentric fragment breaks, resulting in chromatids with large deletions that lead to nonviable gametes or embryonic lethality. A crossover within a pericentric inversion produces recombinant chromatids that have duplications or deletions. Gametes with these recombinant chromatids also do not lead to viable progeny. So even though there is the same amount of recombination, those recombinant products never make it into viable offspring. 8. Give two ways that translocations can produce phenotypic effects. If a translocation break point is within a gene, the translocation will disrupt that gene, inactivating it. The translocation of a gene, causing it to become adjacent to a new 2 region of a chromosome, changes the linkage pattern. That is, it changes the types of sequences that are near the gene, which can lead to position effects, which are changes in expression of the gene. 9. Draw a temperature profile graph for 2 cycles of PCR, giving an approximate temperature at each stage, and tell in one short phrase what happens at each of those stages. You could write each of the phrases alongside each step in the profile. 10. Use an entire sheet of paper for this drawing to give you plenty of room. It doesn’t need to be neat, only clear. Draw three base pairs of a DNA molecule that has the sequence 5’-GCA-3.’ Draw one strand down the page on the left, and the other strand base-paired with it just to the right. Use the same amount of detail as we used in class. Show the negative charge on each phosphate group, label the 5’ and 3’ carbon positions with numbers, and show the hydrogen bonding between paired bases with dashed lines (even though they don’t need to be placed accurately with respect to specific atoms). Label the 5’ and 3’ ends of each strand. The only nitrogen atoms you need to show are the ones in the bases that form the covalent bonds with the deoxyriboses. Place a small star next to each carbon atom that would have a hydroxyl (-OH) if the left strand were an RNA molecule. 11. In the 1920’s Griffith did an experiment with bacteria and mice that showed there exists a “transforming” material that can change the genetic characteristics of an organism. What was the characteristic that was passed from one bacterium to another? The ability to kill mice, a phenotype that could also be seen as the ability to produce smooth colonies. 3 What did he do to ensure that the bacteria which originally had the characteristic weren’t merely passed through the critical experiment? He killed them by heating them. 12. What is euchromatin? (one phrase or sentence) What is heterochromatin? (one phrase or sentence) In which type of material does most transcription occur? Euchromatin is chromatin that undergoes the normal process of condensation and decondensation in the cell cycle. Heterochromatin is DNA that remains in a condensed state throughout the cell cycle. Most transcription occurs in euchromatin. 13. What is a nucleosome and what are two of its functions? Name the 8 proteins classically considered to be an essential part of the nucleosome core. A nucleosome is a core particle of chromatin structure consisting of 8 histones around which about 145-150bp of DNA are wrapped. Two of its functions are to aid in the physical organization of DNA into a shorter structure, and to help control expression of genes. The eight proteins are: two copies each of histones H2A, H2B, H3 and H4 14. In this problem you will do four very simple drawings (a-d). a. Draw a DNA molecule as it would look before it is denatured, preparatory for PCR amplification, using only lines with arrows at the ends to show the 5’ and 3’ ends. Here is that molecule to get you started. Ignore the letters A and B for the moment. A B b. Now draw that DNA as it would look during the heat denaturation step in the first cycle of a PCR. c. Now draw that DNA as it might look during the annealing (or hybridization) step just after two PCR primers have annealed to it. Let the primers anneal at positions A and B and draw them as short arrows oriented with their 3’ ends pointing in the direction of the arrow. d. Now draw those DNA molecules as they would look after the first extension step in which Taq polymerase had extended from the primers. Let any newly synthesized DNA be shown as a wavy line with an arrow at one end to signify the 3’ end. Cool! You’ve got more DNA than you started out with! 4 5