* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Gene Tagging with Transposons
RNA interference wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Ridge (biology) wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
X-inactivation wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Public health genomics wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
RNA silencing wikipedia , lookup
Gene nomenclature wikipedia , lookup
Genomic imprinting wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Metagenomics wikipedia , lookup
Copy-number variation wikipedia , lookup
Oncogenomics wikipedia , lookup
Gene therapy wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Primary transcript wikipedia , lookup
Genetic engineering wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Gene desert wikipedia , lookup
Point mutation wikipedia , lookup
Pathogenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
Human genome wikipedia , lookup
Gene expression programming wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Microsatellite wikipedia , lookup
Genome (book) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Minimal genome wikipedia , lookup
History of genetic engineering wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Microevolution wikipedia , lookup
Genomic library wikipedia , lookup
Designer baby wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome editing wikipedia , lookup
Genome evolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
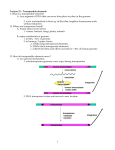
Transposable Genetic Elements MBIOS 520/420 September 22, 2005 MBios 420/520 Announcements • PowerPoint lectures are now available for download at the web site http://mbios420.tripod.com/mbios420.htm • No longer holding office hours; please e-mail me at [email protected] for an appointment • Thoughts on a review session? • Excellent source for questions and review: Go to http://www.ncbi.nlm.gov Search in BOOKS. Enter keywords. Hit GO. Transposon Introduction • Transposable elements are stretches of DNA that can move to new locations in a genome • These elements can contain genes or be non-coding • Large portions of higher eukaryotes’ genomes are composed of either inert or active transposons (often as repetitive DNA) • Transposons are thus important evolutionarily • Transposons can also be used to isolate genes or introduce foreign genes into cells Bacterial Transposon Discovery • First transposons characterized, these are the simplest • Detected because experimental lac- strains kept reverting back to wild type (ie, colonies kept turning blue) • lac- mutants were due to transposons which then moved back out of the gene Agar w/X-Gal transposon Agar w/X-Gal Transposition event lac gene lac gene lac gene Bacterial Transposons • Can occur in bacterial genome or in plasmid, and can move between these two • Consist of two major types: Insertion Sequences (IS) small, <2500 bp Composite Transposons large, flanked by two IS elements Insertion Sequences • Consists of a pair of inverted terminal repeats at each end (cannot be mutated without loss of transposition activity) • Between this is a stretch of DNA, often containing the gene for transposase – the enzyme that catalyzes transposition • Flanking the terminal repeats are a pair of direct repeats that result from the transposition process Insertion Sequence Transposition Transposase moves the element by creating a staggered cut at either end in a random spot of the genome The IS element then moves then inserts into this region DNA polymerase fills in the resulting singlestranded areas The result is termed a target site duplication Composite Transposons • Denoted Tn • Created when two IS elements insert near each other • The elements can be either in inverse or direct orientation to each other • These two then move together and transpose the sequence between them (often carrying genes) • Movement of these large elements is how bacteria become antibiotic resistance (often using viral intermediates) Composition Transposon Transposition • Involves both IS elements • Two types: Replicative Transposon transposon is copied & moved (ie, a copy remains in place) Non-Replicative Tranposon (aka “cut & paste”) the whole element is moved (no copy remains) • Similar to conserved & semi-conservative DNA replication Tn3 Transposition • A combination of replication & recombination Tn10 Transposition How can we determine if a transposon uses replicative or “cut and paste” transposition? Gene Tagging with Transposons • We can use transposons to tag and isolate genes • Ex: let’s say we want to isolate a blue flower color gene STEP 1 transposon A. Transform plant with a vector containing a transposon. M2 (Thousands of plants needed, depending on genome size) transposon Blue color gene B. Grow progeny from that plant and pick out the mutant phenotype (in this case, an uncolored flower). C. This plant should have the transposon inserted somewhere in the gene. Gene Tagging with Transposons STEP 2 A. Isolate genomic DNA from the mutant plant. BAC clones (many thousands of these) B. Make genomic DNA library from this sample (ex: using BAC vectors). C. Pool clones of the library into 96-well plate. THE MATH Ex: Rice Genome size = 400 million bases BAC insert size = 200,000 bases # clones needed for 1X = 2,000 # clones needed for 6X = 12,000 # of clones per well = 125 Gene Tagging with Transposons PCR w/ transposon primers 1 2 3 4 5 STEP 3 1 2 3 4 5 A. Find the well containing the BAC clone with the transposon (use PCR). B. Grow the cells from this well on a plate. P32-labeled transposon probe C. Hybridize with transposon probe to locate exact colony that has the clone. D. Sequence this clone. You’ve found the gene! Gene Tagging with Transposons & Inverse PCR • There is a faster way of identifying the gene without having to build an entire library, by using a technique called inverse PCR STEP 2b EcoRI A. Follow STEP 1 like before. Isolate mutant & extract its DNA. B. Cut DNA with restriction enzymes, then re-ligate to form circular segments. DNA ligase (high volume reaction) SEQUENCE C. Use the transposon-based primers & do PCR. This amplifies the flanking gene regions. Sequence it. Gene Tagging with Transposons - Troubleshooting • What if we get many transposons inserting into our mutant genome? How do we tell which one is in our color gene? OPTION 1 Do inverse PCR as in STEP 2B. SEQUENCE BLAST sequence & search for similarity with other known pigment genes. (GENBANK) Gene Tagging with Transposons - Troubleshooting OPTION 2 Cross mutant plant back to wild-type plant. Produce an F2 (or more) generation. Do Southern blot with tranposon probe. Find marker that segregates with mutation. Many plants needed. P X F1 F2 λ HindIII Marker 23.0 kb 9.4 kb 6.5 kb 4.3 kb 2.3 kb 2.0 kb Make a library out of a plant with only this marker (Plant D in our example). 0.5 kb A B C D E F Gene Tagging with Transposons - Troubleshooting OPTION 3 A B C D Pick out all BAC clones with transposons via PCR (STEP 3). Do RFLP ofF2 mutant plant using transposon as a probe. λ HindIII F2 Mutant Marker Plant 23.0 kb Cut BACs with same restriction enzyme as in your RFLP. Hybe with tranposon probe. Transposon probe should bind at same MW in mutant and BAC digest. 9.4 kb 6.5 kb 4.3 kb 2.3 kb 2.0 kb 0.5 kb A B C D Eukaryotic Transposons • Similar in structure to bacterial transposons • Most are thought to be derived from viral genomes that have integrated into a host cell genome • Some eukaryotic transposons move via an RNA intermediate • Some transpositions are utilized for programmed genome rearrangements • Movement of transposons in genomes can inactive or activate genes, and can cause cancer • The movement and buildup of transposable sequences has had an effect on the evolution of eukaryotice genomes Maize Transposons • Two best characterized transposons are Ac (Activator) and Ds (Dissociation) elements (both are “cut & paste” types) • Can occur in many copies of the cell, but must work together • Ac is ~4.6 kb, with same basic structure as insertion elements • Ds is similar (identical inverted terminal repeats) but has deletions of various sizes in it Transposase Gene Transposase Missing • Because of the deletions, Ds does not have transposase & cannot transpose itself (Ac needed) Discovery of Transposons • Barbara McClintock, maize geneticist • Observed mosaic corn kernels, despite presence of CI, a dominant colorless allele • Concluded that chromosome was breaking, causes CI loss • Only occurred when another segregating factor, Ac, was present CI Present CI Absent Using Ac & Ds Elements for Mutagenesis • When we do transposon mutagenesis, how can we control when transposition stops & starts? • Ac can be introduced via a vector Ac+/+ Ds-/P Ac-/- Ds+/+ X Ac+/- Ds+/- F1 F2 • But how do we know that Ac won’t transpose into our gene instead of Ds? Ac-/- Ac+/? Ds+ X Mutate the inverted terminal repeats of Ac, then it can’t transpose! BC1F1 BC1F2 Stable Mutant Ac-/- Ds+ Drosophila Transposons • Known as P elements, similar in structure to Ac & Ds • P elements can be incomplete (no transposase) or complete (functional transposase) – analogous to Ac/Ds • P elements are only active in germ line cells, because a stop codon exists in transposase • In germ cells, alternative splicing removes exon 2 to remove the codon • Demonstrated by engineering a P element without intron 3 Retro-Transpososons • These transpose via an RNA intermediate • Transposon is transcribed into RNA, then reverse transcriptase creates “cDNA” that inserts into genome • Two categories exist, based on their origin & structure: Retroviruslike Elements Retroposons Possess Long Terminal Repeats Have viral genes gag & pol Derived from retroviruses Non-LTR retrotransposons Have poly A:T tract Derived from reverse transcribed mRNA Retroviruslike Elements • Ty1 of yeast is best studied • Long terminal repeats called δ regions (not inverted) flank a coding region • Coding region has TyA & TyB; these genes are gag & pol derived • ~35 copies per yeast cell; sometimes solo δ regions are found (formed by recombination between δ) • Form target site duplications Ty1 Transposition • RNA is synthesized by normal RNA pol II transcription • δ elements act as strong promoters (can activate genes) • TyB gene has reverse transcripase activity and produces dsDNA from the RNA • DNA integrates into the genome (δ element is replicated) • copia and gypsy are Drosophila retrotransposons very similar to Ty1 (gypsy even has viral env gene) Proof of Ty1 Transposition • How can we prove Ty1 transposes as an RNA molecule? • Constructed Ty1 element with a galactose-inducible promoter and an intron • Used galactose to stimulate transcription, then found that all the new copies transposed had the intron spliced out Retroposons • F, G and I elements in Drosophila; LINEs in humans • Also called non-LTR retrotransposons because they lack inverted or direct repeats at their ends (do have target site repeats) • Retroposons all have a poly-A region at the end, evidence that these are reverse transcribed mRNAs that re-inserted in the genome • These function by reverse transcription, followed by insertion • LINE-1 or L1 = only known active human transposon (make up 5% of human genome) & can cause mutations (ex: hemophilia) Transposition & Chromosome Rearrangement • Can create duplications, deletions, inversions, translocations • Means of creating pseudogenes – no selection pressure (can gain novel function) Deletions & Duplication Created by Transposition: Transposition & Chromosome Rearrangement Deletions & Inversions Created by Transposition: Transposition & Cancer • Antibody genes have powerful enhancers and use recombination to produce diverse sets of antibodies • c-myc can recombine with this region & cause cancer Enhancer Trapping • Technique use transposons to identify tissue-specific enhancers • Add lacZ gene between the inverted terminal repeats of a P element • Transform Drosophila with P element • Dissect Drosophila & grind tissue in X-Gal • Blue color change shows that P element inserted near a tissue specific enhancer Or do inverse PCR Transposons & Genome Evolution • All active transposons have the potential to cause mutations • These can be deleterious or potentially beneficial • Duplication of genes via chromosome rearrangements can produce pseudogenes which can eventually gain new function • Which came first Retroviruses or Retrotransposons?