* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download recBCD
DNA sequencing wikipedia , lookup
Genome evolution wikipedia , lookup
DNA barcoding wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
DNA profiling wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Human genome wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Metagenomics wikipedia , lookup
Genetic engineering wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Designer baby wikipedia , lookup
Primary transcript wikipedia , lookup
DNA polymerase wikipedia , lookup
SNP genotyping wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Point mutation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genomic library wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Microevolution wikipedia , lookup
Microsatellite wikipedia , lookup
Epigenomics wikipedia , lookup
DNA vaccination wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Molecular cloning wikipedia , lookup
Non-coding DNA wikipedia , lookup
DNA nanotechnology wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
History of genetic engineering wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome editing wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Helitron (biology) wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Homologous recombination wikipedia , lookup
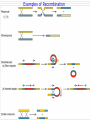
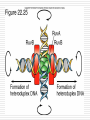
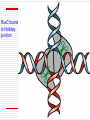
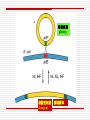
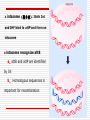
Chapter 11 DNA recombination and Transposon Examples of Recombination 交互的 DNA Recombination Roles Types Homologous recombination in E.coli Transposable elements Biological Roles for Recombination 1. Deleterious mutations would accumulate in each chromosome. Recombination generates genetic diversity 多样性 2. Generating new gene/allele combinations (crossing over during meiosis) 3. Integration of a specific DNA element 4. Role in DNA damage and repair 5. Gene regulation Practical Uses of Recombination 1. Used to map genes on chromosomes (recombination frequency proportional to distance between genes) 2. Making transgenic cells and animals Types of Recombination 1. Homologous - occurs between sequences that are nearly identical (e.g., during meiosis) 2. Site-Specific - occurs between sequences with a limited stretch of similarity; involves specific sites 3. Transposition – DNA element moves from one site to another, usually little sequence similarity involved Homologous Recombination: The Holliday model (1964) Two homologous duplexes are aligned Strand exchange leads to an intermediate with crossed strands This branch can move: Branch migration (~30bp/s) The branch is resolved (拆分) by cleavage and sealing Holliday Model R. Holliday (1964) - Holliday Junctions form during recombination - HJs can be resolved 2 ways, only one produces true recombinant molecules Patch recombination 片段重组体 Holliday中间体 Splice recombination 拼接重组体 Holliday结构一经生成即可 不断地处于异构化--异源双链 hetero duplex DNA Isomerization (Holliday的异构化) Resolution ◘ Holliday resolution (1) 产生含异源双链 的片段重组体 (2) (2) (1) 产生拼接重组体 3‘ 5‘ 切割 5‘ 3‘ Meselson-Radding模型 单链入侵模型(链转移模型) 5’ 置换 侵入 Loop切除 同化 异构化 分支迁移 双链断裂修复模型 Double strand break model Double strand break model Homologous recombination in E.coli ◘ This process involved the exchange of homologous regions between two molecules Conjugation 接合 Transformation 转化 Transduction 转导 Cell fusion 细胞融合 Chi site and RecBCD ptotein E.coli also perform recombination, the nick site in two homologous DNA duplexes created by RecBCD are near a specific sequence called Chi site. Chi site ◘ A Chi site is a short stretch of DNA near which homologous recombination is unusually likely to occur. ◘ In E. coli, the sequence is 5'-GCTGGTGG-3'. 1/5~10kb ◘ Chi serves as a signal to the RecBCD helicaseexonuclease. RecBCD : A complex enzyme ◘ RecBCD enzyme is both a helicase and a nuclease ◘ RecBCD recognizes a specific sequence of Chi site. ◘ RecBCD cut one DNA strand close to Chi sequence 凡能从多核苷酸链的末端开始水解核酸的酶称为核酸外切酶 凡能从多核苷酸链中间开始水解核酸的酶称为核酸内切酶 而能识别特定的核苷酸顺序,并从特定位点水解核酸的内切酶称为限制性核酸内切酶 recBCD Pathway of Homologous Recombination •RecBCD binds an end of linear dsDNA •RecD helicase travels on the strand with a 5' end and RecB on the strand with a 3' end •RecB is slower than RecD, so that a ssDNA loop accumulates ahead of RecB •This produces DNA structures with two ss tails and one ss loop •ss tails can anneal to produce a second ss loop complementary to the first one; such twin-loop structures were referred to as “rabbit ears.” •When RecBCD encounters a Chi site on the 3' ended strand, unwinding pauses and digestion of the 3' tail is reduced. • The RecA protein is then actively loaded onto the 3' tail by RecBCD and form RecAssDNA filaments. • RecA catalyzes branch migration and makes it possible to complete recombination The recBCD Pathway of Homologous Recombination Part I: Nicking and Exchanging Assimilation 同化 recBCD Pathway of Homologous Recombination Part I: Nicking and Exchanging 1. 2. 3. 4. 5. A nick is created in one strand by recBCD at a Chi sequence (GCTGGTGG), found every 5000 bp. Unwinding of DNA containing Chi sequence by recBCD allows binding of SSB and recA. recA promotes strand invasion into homologous DNA, displacing one strand. The displaced strand base-pairs with the single strand left behind on the other chromosome. The displaced and now paired strand is nicked (by recBCD?) to complete strand exchange. recBCD Pathway of Homologous Recombination Part II: Branch Migration and Resolution Generation of a chi intermediate Electron micrograph of the chi form Without functional RecA protein in E.coli, the exogenous plasmid DNA is left unaltered by the bacteria. Purification of this plasmid from bacterial cultures can then allow high-fidelity PCR amplification of the original plasmid sequence. RecA 38 kDa protein that polymerizes 聚合 onto SS DNA 5’-3’ Catalyzes strand exchange, also an ATPase Also binds DS DNA, but not as strongly as SS RecA Function Dissected Three steps of strand exchange: 1. Pre-synapsis: recA coats single stranded DNA (accelerated by SSB, get more relaxed structure) 2. Synapsis 联会 : alignment of complementary sequences in SS and DS DNA (paranemic or side-by-side structure) 3. Post-synapsis or strand-exchange: SS DNA replaces the same strand in the duplex to form a new DS DNA (requires ATP hydrolysis) RecA promotes the assimilation of invading single strands (单链同化)into duplex DNA so long as one of the reacting strands has a free end. Ruv protein and resolution of Holliday structures Need three genes in E.coli, ruvA,ruvB and ruvC a、 RuvA recognize the junction of Holliday b、 RuvB provide energy for migration (ATPase 10~20bp/s) c、 RuvC nuclease cut Holliday junction specifically. RuvA and RuvB DNA helicase that catalyzes branch migration RuvA tetramer binds to HJ (each DNA helix between subunits) RuvB is a hexamer ring, has helicase & ATPase activity 2 copies of ruvB bind at the HJ (to ruvA and 2 of the DNA helices) Branch migration is in the direction of recA mediated strand-exchange RuvC : resolvase Endonuclease that cuts 2 strands of HJ Binds to HJ as a dimer Consensus sequence: (A/T)TT (G/C) - occurs frequently in E. coli genome - branch migration needed to reach consensus sequence! RuvC bound to Holliday junction Action of E. coli proteins in branch migration and resolution of Holliday structures Site specific recombination 位点特异性重组 Viruses and transposable elements often integrate their genomes into the host chromosome Site specific recombination is used by both eukaryotes and prokaryotes to regulate gene expression and to increase the organisms genetic repertoire Site specific recombination 1、Occurs between sequences with a limited stretch of similarity; involves specific sites and proteins 依赖于小范围同源序列的联会,重组也只 发生在同源的短序列的范围之内,需要位点特异性的蛋白 质分子参与催化 2、Based on position and direction λphage integrate and cleavage 1、Via Site-specific recombination 2、attachment site ◘ E.coli (att) attB BOB’ 23bp ◘ λphage attP POP’ 240bp ◘ “O”complete same(15bp) 3、Integration ◘ attachment site attL(BOP’) attR(POB’) ◘ Integration site---attB、attP Excision site---attL、attR ◘ need integrase Int(λ encoded) and integration host factor 溶菌周期 (lysis) 溶源性细菌 (lysogen) 原噬菌体 4、Mechanism ◘ Core sequence is “O” 15bp,A-T rich ◘ exchange within O region:7bp ◘ Int Binding site:attP 240bp、attB 23bp ◘ intasome(整合体): Both Int and IHF bind to attP and form an intasome ◘ intasome recognize attB a、attB and attP are identified by Int b、 Homologous sequences is important for recombination ◘ Int cleavages DNA lead to crosswise pairing holliday J ◘ Recombinant junction are resolved, sealed to generate intergrated prophage DNA 5、The control of integration - excision Control of the integration-excision reaction depends on: (1) the forward (insertion) reaction, which requires only Int among phage-specified proteins (2) the reverse (excision) reaction, which requires the phagecoded Int and Xis proteins Knockout mouse A genetically engineered mouse in which one or more genes have been turned off through a targeted mutation. The first knockout mouse was created by Mario R. Capecchi, Martin Evans and Oliver Smithies in 1989, for which they were awarded the Nobel Prize for Medicine in 2007 Generate Knockout mice based on HR Conditional knockout mice