* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download XIXth INTERNATIONAL CONFERENCE OF GENETIC DAYS, 5th …
Epigenetics in learning and memory wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Human genome wikipedia , lookup
DNA polymerase wikipedia , lookup
Population genetics wikipedia , lookup
Primary transcript wikipedia , lookup
Gene therapy wikipedia , lookup
Genome evolution wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
DNA profiling wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Genetic engineering wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Point mutation wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
DNA vaccination wikipedia , lookup
Genetic drift wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Genomic library wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Molecular cloning wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Non-coding DNA wikipedia , lookup
Epigenomics wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
SNP genotyping wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome editing wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Designer baby wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Helitron (biology) wikipedia , lookup
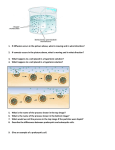
Novel approaches for linkage mapping in dairy cattle. ”Selective DNA pooling” Chandra S. Pareek Dept. of Animal Genetics, Faculty of Animal Bio-Engineering, University of Warmia and Mazury, Olsztyn, Poland. Main sub-headings Definition Principle Experimental design Experimental design to locate the QTL region through selective DNA pooling. Microsatellite genotyping Statistical methods for accurate estimation of gene frequency from pooled samples. Problems in determination of gene frequency. Problems in interpreting pooling results by visual inspection. Application of Selective DNA pooling in farm animals. Advantages of Selective DNA pooling. Success of selective DNA pooling in dairy cattle. Definition • “Selective DNA pooling” is an advanced methodology for linkage mapping of quantitative, binary and complex traits in farm animals. • In human, this methodology is termed as DNA pooling where it serves as mapping of complex disease traits. • It is defined as densitometric genotyping of physically pooled samples from phenotypically extreme individuals. • The DNA pooling is performed by taking equal aliquots of DNA from the pooled individuals. Principle • ¨ The principle is based on densitometric estimates of marker allele frequency from the pooled phenotypically extreme individuals. • ¨ In this regards, the marker of choice is: STR or microsatellite markers. • ¨ The microsatellite allele linked to any QTL gene can be identified by any shift or deviation of allele from the pools. • ¨ The QTL linked allele, and then further tested for its feasibility by statistical analysis. • ¨ The power of statistics is relied on the accurate estimates of gene frequency from the pooled samples. • ¨ Several methods have been described for accurate estimation of gene frequencies (Daniels et al. 1998, Barcellos et al. 1998, Lipkin et al. 1998, and Collins et al. 2000). • Affected pool Figure A and B: showing allelic patterns of a linked marker. Unaffected pool Here figure A is displaying a shift of marker allele in affected individuals pool Affected pool Figure C and D: showing allelic patterns of a unlinked marker. Unaffected pool Here both figures are not displaying any shift or deviation of the alleles. Experimental design A well-defined experimental design is an essential prerequisite to perform the selective DNA pooling. • The experimental design should include the following conditions. • 1. Identification of resource families having extreme phenotypic values for the given analysed trait. • 2. Systemic selection of highly polymorphic STR markers from the analysed genome. • Experimental design to locate the QTL region through selective DNA pooling • Daughter design: In case of cattle, by utilizing multiple half-sib families with multiple STR markers. Granddaughter design: In case of cattle, poultry and swine, by utilizing F2 full-sib daughters including sire and grand sire. Microsatellite genotyping • The most commonly used touch down protocol of Don et al. 1991, can be used for typing of microsatellte markers, followed by visualisation of electrophoresis results in any DNA sequencing machine (Perkin Elmer ABIPrism, Pharmacia ALF, and LICOR genetic analyser). Statistical methods for accurate estimation of gene frequency from pooled samples The following three methods have been described: By measuring the relative intensity of shadow bands (RI): Method proposed by Lipkin et al. 1998. By measuring the Allelic Image Pattern (AIP) from the pools: Method proposed by Daniels et al. 1998. By measuring the Total Allelic content (TAC) from the pools: Method proposed by Collins et al. 2000. first Method: Lipkin et al. 1998 Measuring relative intensity of shadow bands (RI) By giving the densitometric values of main and shadow bands, the relative intensity of a given shadow band for a given allele can be calculated as: RIn.i = Dn.i / Dn Where, n = is the number of repeats in the native genomic tract of the allele An I = is the order of the shadow band RIn.i = is the relative intensity of the ith shadow band derived from the genomic tract of An Dn = is the densitometric intensity of the main band derived from the genomic tract of An Dn.i =is the densitometric intensity of the ith shadow band derived from the genomic tract of An 2nd method: Daniels et al.. 1998 Measuring Allelic Image Patterns (AIP) from the pools • The principle of this method is based on the analysis of microsatellite allele image patterns (AIP) generated from the DNA pools. • The AIP statistic is calculated from the differences in the area between two allele image pattern expressed as a fraction of total shared and non-shared area. AIP = Dif / (Dif + Com) The AIPs from the pools and X2 values from individual genotyping were compared. The results demonstrated a high correlation between AIPs and X2 values. Figure showing overlaid AIPs of two different pools amplified with the microsatellite marker. Area ”Dif” and ”Com” are the non-shared and common areas between the two AIPs. 3rd method: Collins et al.. 2000 Measuring Total Allelic content (TAC) from the pools • This is a modified method of Daniels et al.. 1998. • The principle of this method is based on simple measurement of total allele differences by comparing the two pools. • The pool comparison is done by comparing the relative peak height differences between electrophoregrams for each allele of a microsatellite. Figure A: Showing peak image profile from individual genotyping illustrating sutter profile and amplitude variation. Figure B: showing peak image profiles from pooled genotyping. Problems in determination of gene frequency Feasibility and reliability of selective DNA pooling is depend upon the accurate estimates of the gene frequency from pooled samples, which is mostly confounded with Sutter banding and Differential amplification. • 1. Sutter banding • 2. Differential amplification. Problems in interpretating pooling results by visual inspection Visual inspection of numerous STR genotyping of pooled samples can be performed by visual eye balling of the peak image files. There are 2 problems encountered during visual inspection of the peak image files. True negative peaks False positive peaks control Shifted allele Figure 1: Showing shifting of microsatellite allele in affected group. This figure represents the True result with correct peak profile image. unaffected Example of correct result Figure 2: Showing shifting of microsatellite allele in affected group. False .Shifted allele This figure represents a good example of False positive peak profile image. Example of false positive result Figure 3: Showing no shifting of microsatellite alleles but there is one linked marker allele in this locus. This figure represents a good example of True Negative peak profile image. Example of true negative result Application of selective DNA pooling in farm animals In rapid genome scanning for the identification of unknown gene or linked gene fragment. In rapid estimation of STR gene frequency. More recently in estimation of SNP frequencies as well. In identification of complex gene fragment within the genome through linkage analysis of STR marker linked to that gene fragment. In QTL mapping of the identified gene or gene fragment. To detect genes with small effect, for e.g., complex disease traits in human. Figure representing detection of linked allele by comparing affected and unaffected DNA pools. In this figure: Marker D5S393 is showing the linked allele to the disease trait whereas, marker D5S410 showing no allele linked to the disease trait. Advantages of selective DNA pooling ¨To detect any linkage between marker and QTL: Multiple families with large numbers of daughters are required to get reasonable statistical power. This requirement leads to genotyping of hundreds of thousands individuals with high cost of experiment. By means of selective DNA pooling, the cost of numerous genotyping can be substantially reduced. Thus selective DNA pooling is an ideal and potential approach for analysing multiple large families with multiple microsatellite markers. ¨ Selective DNA pooling reduces not only the genotype cost by many folds, but also minimizes the valuable experimental time. For example:individual v/s Pooled genotyping In case of individual G: 2000 markers x 2000 individuals = 4 x 106 individuals In case of Pooled G: The genotyping becomes 2000 x 2 = 4000. Success of selective DNA pooling in dairy cattle ¨ Mapping of QTL genes for milk protein percentage in Israeli HF cattle (Lipkin et al. 1998). ¨ Detection of loci that affect quantitative traits like milk production in New Zealand HF and Jersey cattle population (Spelman et al. 1998).