* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download 20. Transposable Genetic Elements
X-inactivation wikipedia , lookup
Human genetic variation wikipedia , lookup
Frameshift mutation wikipedia , lookup
DNA vaccination wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Population genetics wikipedia , lookup
Molecular cloning wikipedia , lookup
Minimal genome wikipedia , lookup
Oncogenomics wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Public health genomics wikipedia , lookup
DNA supercoil wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Primary transcript wikipedia , lookup
Human genome wikipedia , lookup
Gene expression programming wikipedia , lookup
Genomic library wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genome evolution wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Point mutation wikipedia , lookup
Genetic engineering wikipedia , lookup
Designer baby wikipedia , lookup
Genome editing wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genome (book) wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
History of genetic engineering wikipedia , lookup
Transposable element wikipedia , lookup
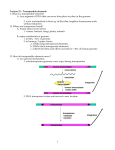
Transposable Genetic Elements 20. Transposable Genetic Elements Key Concepts A series of genetic elements can occasionally move, or transpose, from one position on a chromosome to another position on the same chromosome or on a different chromosome. In bacteria, insertion sequences, transposons, and phage mu are examples of transposable genetic elements. Transposable elements can mediate chromosomal rearrangements. In higher cells, transposable elements have been extensively characterized in yeast, Drosophila, and maize and in mammalian systems. In eukaryotes, some transposable elements utilize an RNA intermediate during transposition, whereas, in prokaryotes, transposition is exclusively at the DNA level. Introduction Starting in the 1930s, genetic studies of maize, undertaken independently by Barbara McClintock and Marcus Rhoades, yielded results that greatly upset the classical genetic picture of genes residing only at fixed loci on the main chromosome. The research literature began to carry reports suggesting the existence of genetic elements of the main chromosomes that can somehow mobilize themselves and move from one location to another. These findings were viewed with skepticism for many years, but it is now clear that such mobile elements are widespread in nature. A variety of colorful names (some of which help to describe their respective properties) have been applied to these genetic elements: controlling elements, cassettes, jumping genes, roving genes, mobile genes, mobile genetic elements, and transposons. We choose the term transposable genetic element, which is formally most correct and embraces the entire family of types. The term transposition has long been used in genetics to describe transfer of chromosomal segments from one position to another in major structural rearrangements. In the present context, what is being transposed seems to be a gene or a small number of linked genes or a gene-sized fragment. Any genetic entity of this size can be called a genetic element. Transposable genetic elements can move to new positions within the same chromosome or even to a different chromosome. The normal genetic role of these elements is not known with certainty. They have been detected genetically through the abnormalities that they produce in the activities and structures of the genes near the sites to which they move. A variety of physical techniques have been used to detect them as well, including DNA sequencing. Transposable genetic elements have been found in most organisms in which they have been sought. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Today, transposable elements provide valuable tools both in prokaryotes and in eukaryotes for genetic mapping, creating mutants, cloning genes, and even producing transgenic organisms. Let us reconstruct some of the steps in the evolution of our present understanding of transposable elements. In doing so, we will uncover the principles relevant to these fascinating genetic units. Controlling elements in maize In 1938, Marcus Rhoades analyzed an ear of Mexican black corn. The ear came from a selfing of a pure-breeding pigmented genotype, but it showed a surprising modified Mendelian dihybrid segregation ratio of 12:3:1 among pigmented, dotted, and colorless kernels. Analysis showed that two events had occurred at unlinked loci. At one locus, a pigment gene A1 had mutated to a1, an allele for the colorless phenotype; at another locus, a dominant allele Dt (Dotted) had appeared. The effect of Dt was to produce pigmented dots in the otherwise colorless phenotype of a1/a1 (Figure 20-1). Thus, the original line was very probably A1/A1 ; dt/dt, and the mutations generated an A1/a1 ; Dt/dt plant, which on selfing gave the observed ration of progeny. But what was causing the dotted phenotype? A reverse mutation of a1 → A1 in somatic cells would be an obvious possibility, but the large numbers of dots in the Dotted kernels would require extremely high reversion rates. Using special stocks, Rhoades was able to find anthers in the flowers of a1/a1 ; Dt/– plants that showed patches of pigment (Figure 20-2). He reasoned that these anthers might contain pollen grains bearing the reverted pigment genotype, and he used the pollen from these anthers to fertilize a1/a1 tested females. Sure enough, some of the progeny were completely pigmented, showing that each dot in the parental plants was in fact the phenotypic manifestation of a genetic reversion event. Thus, a1 is one of the first known examples of an unstable mutant allele—an allele for which reverse mutation occurs at a very high rate. However, the allelic instability is dependent on the presence of the unlinked Dt gene. When the reverse mutations occur, they are stable; the Dt gene can be crossed out of the line with no loss of the A1 character. This outcome would not be surprising if the a1 phenotype arose from the insertion of a defective transposable element that by itself is unable to move. In the presence of a transfactor produced by the Dt locus, however, the element can move, yielding reversion to A1. The A1 allele remains stable in the absence of Dt. McClintock's experiments: the Ds element In the 1950s, Barbara McClintock demonstrated an analogous situation in another study of corn. She found a genetic factor Ds (Dissociation) that causes a high tendency toward chromosome breakage at the location at which it appears. These breaks can be located either cytologically (Figure 20-3a) or genetically by the uncovering of recessive genes (Figure 20-3b). This action of Ds is another kind of instability. Once again, this instability proves to be dependent on the presence of an unlinked gene, Ac (Activator), in the same way that the instability of a1 is dependent on Dt. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements McClintock found it impossible to map Ac. In some plants, it mapped to one position; in other plants of the same line, it mapped to different positions. As if this were not enough of a curiosity, the Ds locus itself (Figure 20-3) was constantly changing position on the chromosome arm, as indicated by the differing phenotypes of the variegated sections of the seeds (as different recessive gene combinations were uncovered in a system such as the one illustrated in Figure 20-3b). The wanderings of the Ds element take on new meaning for us in the context of this chapter when we consider the results of the following cross: Here C allows color expression and Ds+ and Ac+ indicate the lack of the element. Most of the kernels from this cross were of the expected types (Figure 20-4), but one exceptional kernel was very interesting. In Figure 20-4, the first seed shows the normal solid pigment pattern owing to the presence of the dominant C allele. The second seed shows the same basic background pigmentation but with the expected white mottling caused by the loss of the C allele through chromosome breakage in some of the cell lines within the seed, with the resultant expression of the recessive c. Because of the clonal nature of cell growth in the seed, the size of a white patch is an indication of when in the seed's development the breakage occurred. A small white area suggests that the break came late in development, because it gave rise to only a small number of affected cells. A large patch suggests an early break, because many descendant cells are affected. The bottom seed in Figure 20-4 indicates expression of the c allele at the very beginning of development, inasmuch as the background is white, not pigmented. However, the presence of pigmented blotches on a white background suggests a reversible process at work that allows the C allele to be reexpressed. Chromosomal breakage could not be the explanation in this case, because, on breakage, the C alleles are left on acentric fragments that are lost in mitosis, and therefore the white phenotype is not reversible in such cases. The white coloration in the bottom seed appears to be the result of a second type of action in which the Ds gene transposes into the C allele, disrupting its function but not inducing the chromosome to break. If the Ds allele is then excised and transposes elsewhere, the C allele regains its function, yielding the pigmented cells that form the patches. The Ac allele is still required, but in this case it has the effect of mediating a reversible instability at the C locus. If Ac is crossed out of the line, the cu allele becomes a stable mutant. The analogy of this system with the a1 Dt system is obvious. Perhaps the earlier situation also is due to the insertion of a Ds-like element into the A1 gene. It is natural to ask whether a1 will respond to Ac or cu will respond to Dt. The answer is no; some kind of specificity prevents this cross-activation of mutational instability. The wx (waxy) locus 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements The Ds element can wander not only into the middle of the C gene, but also into other genes, rendering them unstable mutants dependent on Ac. One such locus, wx (waxy), has been the subject of an intense study on the effects of the Ds element. Oliver Nelson paired many unstable waxy alleles in the absence of the Ac mutation. In such wxm−1/wxm−2 heterozygotes, he looked for rare wild-type Wx recombinants by staining the pollen with KI-I2 reagent, which stains Wx pollen black and wx pollen red. By counting the frequency of Wx pollen grains in each kind of heterozygote, Nelson was able to do fine-structure recombination mapping of the waxy gene. He showed that the different “mutable waxy” mutant alleles are in fact due to the insertion of the Ds element in different positions in the gene. Continuing the experiment, he allowed the Wx-bearing pollen to fertilize wx plants and produce rare Wx kernels, which could be detected by staining sliced-off slivers. The Wx kernels were then raised into plants, and the exchange of flanking markers expressed in the adults showed that the Wx pollen grains had arisen from chromosome exchange. General characteristics of controlling elements Several systems like a1-Dt and Ds-Ac have now been found in corn. Each shows similar action, having a target gene that is inactivated, presumably by the insertion of some receptor element into it, and a distant regulator gene that maintains the mutational instability of the locus, presumably through its ability to “unhook” the receptor element from the target locus and return the locus to normal function. The receptor and the regulator are termed controlling elements. In the examples considered so far, the unstable allele is said to be nonautonomous: it can revert only in the presence of the regulator. Sometimes, however, a system such as the Ac-Ds system can produce an unstable allele that is autonomous. Such mutants are recognizable because they show Mendelian ratios (such as 3:1 for pigmented to dotted) that apparently are independent of any other element. In fact, such alleles appear to be caused by the insertion of Ac itself into the target gene. An allele of this type can subsequently be transformed into a nonautonomous allele. In such cases, the nonautonomy seems to result from the spontaneous generation of a Ds element from the inserted Ac element. In other words, Ds is in all likelihood an incomplete version of Ac itself. Figure 20-5 summarizes the overall behavior of the controlling elements in maize as inferred from genetic data. Note that the mutation events that gave rise to Rhoades's original ratio are nicely explained by this model: a nonautonomous Ds-like element is generated from an Ac-like progenitor; transposition of the Ds-like element into the A1 gene produces an inactive but mutationally unstable allele a1. Molecular studies in the past few years on Ac, Ds, and other controlling elements in maize have confirmed McClintock's genetic model. MESSAGE 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Controlling elements in maize can inactivate a gene in which they reside, cause chromosome breaks, and transpose to new locations within the genome. Complete elements can perform these functions unaided; other forms with partial deletions can transpose only with the help of a complete element elsewhere in the genome. Figures 20-6 and 20-7 show examples of the effects of transposons in maize and similar effects in the snapdragon. Figure 20-1. A formal genetic explanation of the appearance of the dotted phenotype in corn. The A1/a1 ; Dt/dt genotype is created by simultaneous mutations of A1 → a1 and dt → Dt. On selfing, this genotype yields the observed 12:3:1 ratio of kernel phenotypes. Figure 20-2. Rhoades used special stocks of a1/a1 ; Dt/–corn plants carrying certain genes that allow pigmented sectors to be detected in tissue other than the kernels. (After M. M. Rhoades, Genetics 23, 1938, 382.) 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-3. Detection of chromosomal breakage (instability) due to action of the Ds element in corn. (a) Cytological detection of the breakage. (Ds+ indicates a lack of Ds.) (b) Genetic detection of the breakage. Figure 20-4. Results that indicate the transposition of Ds into the C gene in corn. (C allows color expression; c does not. Ac = activator.) The action of Ds is dependent on the presence of the unlinked gene Ac. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-5. Summary of the main effects of controlling elements in corn. Ac and Ds are used as examples, acting on two hypothetical genes 1 and 2. Bacterial insertion sequences Insertion sequences, or insertion-sequence (IS) elements, are now known to be segments of bacterial DNA that can move from one position on a chromosome to a different position on the same chromosome or on a different chromosome. When IS elements appear in the middle of genes, they interrupt the coding sequence and inactivate the expression of that gene. Owing to their size and in some cases the presence of transcription and translation termination signals, IS elements can also block the expression of other genes in the same operon if those genes are downstream from the promoter of the operon. IS elements were first found in E. coli in the gal operon—a set of three genes taking part in the metabolism of the sugar galactose. Physical demonstration of DNA insertion Recall that phage λ inserts next to the gal operon and that it is a simple matter to obtain λdgal phage particles that have picked up the gal region (page 229). When the IS mutations in gal are incorporated into λdgal phages and the buoyant density in a cesium chloride (CsCl) gradient of the phages is compared with that of the normal λdgal phages, it is evident that the DNA carrying the IS mutation is longer than the wild-type DNA. This experiment clearly demonstrates that the mutations are caused by the insertion of a significant amount of DNA into the gal operon. Figure 20-8 depicts this experiment in more detail. Direct visualization of inserted DNA When denatured λdgal DNA containing the insertion mutation is hybridized to denatured wild-type λdgal DNA, the extra piece of DNA can be located under the 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements electron microscope. In such experiments, some of the DNA molecules that form in the mixture are not the parent duplexes but are heteroduplexes between one mutant and one wild-type strand. When point mutations are analyzed, the heteroduplexes are indistinguishable from the parental DNA molecules. However, in DNA containing the IS mutations, each heteroduplex shows a single-stranded buckle, or loop (Figure 20-9). This single-stranded buckle confirms the presence of an inserted sequence in the mutated DNA that has no complementary sequence in the wild-type DNA. The length of this single-stranded loop can be calibrated by including standardized marker DNA in the preparation. It proves to be approximately 800 nucleotides in length. Identification of discrete IS elements Are the segments of DNA that insert into genes merely random DNA fragments or are they distinct genetic entities? Hybridization experiments show that many different insertion mutations are caused by a small set of insertion sequences. In these experiments, the λdgal phages, which contain the gal− gene, are isolated from the IS mutant bacteria, and their DNA is used to synthesize radioactive RNA in vitro. Certain fragments of this RNA are found to hybridize with the mutant DNA but not with wild-type DNA, indicative of the fact that the mutant contains an extra piece of DNA. These particular RNA fragments also hybridize to DNA from other IS mutants, showing that the same bit of DNA is inserted in different places in the different IS mutants. On the basis of their patterns of cross-hybridization, the insertion mutants are placed into categories. The first sequence, the 800-bp segment identified in gal, is termed IS1. A second sequence, termed IS2, is 1350 bp long. Table 20-1 lists some of the insertion sequences and their sizes. The inverted repeats listed in Table 20-1 will be dealt with shortly under “Transposons.” We now know that the genome of the standard wild-type E. coli is rich in IS elements: it contains eight copies of IS1, five copies of IS2, and copies of other less well studied IS types. It should be emphasized that the sudden appearance of an insertion sequence at any given locus under study means that these elements are truly mobile, with a capability for transposition throughout the genome. Presumably, they produce a mutation or some other detectable alteration of normal cell function only when they happen to end up in an “abnormal” position, such as the middle of a structural gene. Insertion sequences also are commonly observed in the F factor. Figure 20-10 shows an example of an F-lac episome. MESSAGE The bacterial genome contains segments of DNA, termed IS elements, that can move from one position on the chromosome to a different position on the same chromosome or on a different chromosome. Orientation of IS elements 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Because of the base sequence, the two strands of λdgal+ DNA happen to have different buoyant densities. After DNA denaturation, they can be recovered separately in an ultracentrifuge. In some cases, the same strands (that is, parallel strands) from two different IS1 mutants form an unexpected hybrid with each other. Under the electron microscope, these hybrids have a peculiar appearance: each is a double-stranded region with four single-stranded tails (Figure 20-11). This observation is explained by assuming that the IS1 elements are inserted in opposite directions in the two mutants (Figure 20-12). Figure 20-8. Mutation by insertion is demonstrated with phage λ particles carrying the bacterial gene for galactose utilization (gal+) or the mutant gene gal−. The viruses are centrifuged in a cesium chloride solution. The gal− particles are found to be denser. Because the virus particles all have the same volume and their outer shells have the same mass, the increased density of gal− particles shows that they must contain a larger DNA molecule; the gal− mutation was caused by the insertion of DNA. (From S. M. Cohen and J. A. Shapiro, “Transposable Genetic Elements.” Copyright © 1980 by Scientific American, Inc. All rights reserved.) 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-10. Genetic and physical map of the F factor. The positions of the resident insertion sequences, IS2, IS3, and Tn1000 (boxes), are shown relative to the map coordinates 0–100. The arrow indicates the origin and direction of F DNA transfer during conjugation. Figure 20-11. Diagrammatic representation of the appearance of a heteroduplex DNA molecule formed by annealing corresponding strands (that is, strands that have the same base sequence, excepting at the site of the IS) of λdgalm DNA from two specific mutants caused by insertion sequences. This unexpected hybridization between corresponding strands (normally having the same rather than complementary base sequences) can be explained by the model shown in Figure 20-12. Figure 20-12. A model to explain the heteroduplex DNA structure shown in Figure 20-11. IS1 is represented by the maroon lines; α and β represent the ends of the IS1 molecule; H and L represent the strands of λdgal DNA with high and low buoyant densities, respectively. The hybrid can be explained by assuming that the IS1 sequence is inserted in opposite directions in the two mutants. Prokaryotic transposons A frightening ability of pathogenic bacteria was discovered in Japanese hospitals in the 1950s. Bacterial dysentery is caused by bacteria of the genus Shigella. This bacterium initially proved to be sensitive to a wide array of antibiotics that were used to control the disease. In the Japanese hospitals, however, Shigella isolated from patients with dysentery proved to be simultaneously resistant to many of these drugs, including penicillin, tetracycline, sulfanilamide, streptomycin, and chloramphenicol. This multiple drugresistance phenotype was inherited as a single genetic package, and 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements it could be transmitted in an infectious manner—not only to other sensitive Shigella strains, but also to other related species of bacteria. This talent is an extraordinarily useful one for the pathogenic bacterium, and its implications for medical science were terrifying. From the point of view of the geneticist, however, the situation is very interesting. The vector carrying these resistances from one cell to another proved to be a self-replicating element similar to the F factor. These R factors (for resistance) are transferred rapidly on cell conjugation, much like the F particle in E. coli (Chapter 7). In fact, these R factors proved to be just the first of many similar F-like factors to be discovered. These elements, which exist in the plasmid state in the cytoplasm, have been found to carry many different kinds of genes in bacteria. Table 20-2 lists just some of the characteristics that can be borne by plasmids. What is the mode of action of these plasmids? How do they acquire their new genetic abilities? How do they carry them from cell to cell? The discovery of transposons made it possible to answer some of these questions. Physical structure of transposons If the DNA of a plasmid conferring drug resistance (carrying the genes for kanamycin resistance, for example) is denatured to single-stranded forms and then allowed to renature slowly, some of the strands form an unusual shape under the electron microscope: a large, circular DNA ring is attached to a “lollipop”-shaped structure (Figure 20-13). The “stick” of the lollipop is double-stranded DNA, which has formed through the annealing of two inverted repeat (IR) sequences in the plasmid (Figure 20-14). Subsequent studies have shown that the IR sequences are a pair of IS elements in many cases. For instance, IS10 is present at the ends of the region carrying the genes for tetracycline resistance (Figure 20-15). In some cases, however, the IR sequences are much smaller. The genes for drug resistance or other genetic abilities carried by the plasmid are located between the IR sequences in the lollipop head. The IR sequences together with their contained genes have been collectively called a transposon (Tn). (Transposons are therefore longer than IS elements, because they contain extra protein-encoding genes.) The remainder of the plasmid, bearing the genes encoding resistance-transfer functions (RTF), is called the RTF region (Figure 20-16). Table 20-3 lists some of the known transposons. Movement of transposons A transposon can jump from a plasmid to a bacterial chromosome or from one plasmid to another plasmid. In this manner, multiple drug-resistant plasmids are generated. Figure 20-17 is a composite diagram of an R plasmid, indicating the various places at which transposons can be located. MESSAGE 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Transposons were originally detected as mobile genetic elements that confer drug resistance. Many of these elements consist of recognizable IS elements flanking a gene that encodes drug resistance. IS elements and transposons are now grouped together under the single term transposable elements. Phage mu Phage mu is a normal-appearing phage. We consider it here because, although it is a true virus, it has many features in common with IS elements. The DNA double helix of this phage is 36,000 nucleotides long—much larger than an IS element. However, it does appear to be able to insert itself anywhere in a bacterial or plasmid genome in either orientation. Once inserted, it causes mutation at the locus of insertion—again like an IS element. (The phage was named for this ability: mu stands for “mutator.”) Normally, these mutations cannot be reverted, but reversion can be produced by certain kinds of genetic manipulation. When this reversion is produced, the phages that can be recovered show no deletion, proving that excision is exact and that the insertion of the phage therefore does not involve any loss of phage material either. Each mature phage particle has on each end a piece of flanking DNA from its previous host (Figure 20-18). However, this DNA is not inserted anew into the next host. Its function is unclear. Phage mu also has an IR sequence, but neither of the repeated elements is at a terminus. Mu can also act like a genetic snap fastener, mobilizing any kind of DNA and transposing it anywhere in a genome. For example, it can mobilize another phage (such as λ) or the F factor. In such situations, the inserted DNA is flanked by two mu genomes (Figure 20-19). It can also transfer bacterial markers onto a plasmid; here again, the transferred region is flanked by a pair of mu genomes (Figure 20-20). Finally, the phage mu can mediate various kinds of structural chromosome rearrangements (Figure 20-21). 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-14. An explanation at the nucleotide level for the lollipop structure seen in Figure 20-13. The structure is called a transposon. (a) The transposon in its double-stranded form before denaturing. Note the presence of oppositely oriented copies of an insertion sequence (IS). (b) The lollipop structure formed by intrastrand annealing after denaturing. The two IS regions anneal to form the inverted repeat (IR) of the transposon; the transposon genes are carried in the lollipop loop. Figure 20-15. Two different transposons having different inverted repeat (IR) regions and carrying different drug-resistance genes. (a) Tn9 has a short IR region, because the two IS1 elements are in the same orientation and each element has a short inverted repeat. (b) Tn10 has a large IR region because the two IS10 components have opposite orientations, and the entire IS10 sequence constitutes the inverted repeat. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-16. The insertion of a transposon (Tn) into a plasmid. RTF represents the resistance-transfer functional genes of the plasmid. Tn includes both the IS elements and the drug-resistance genes. Figure 20-17. The role of transposable elements in the evolution of antibiotic-resistance plasmids is illustrated by a schematic map of a plasmid carrying many resistance genes. The plasmid appears to have been formed by the joining of a resistance-determinant segment and a resistance-transfer segment; there are insertion elements (IS1) at the junctions, where the two segments sometimes dissociate reversibly. Genes encoding resistance to the antibiotics chloramphenicol (cmR), kanamycin (kanR), streptomycin (smR), sulfonamide (suR), and ampicillin (ampR) and to mercury (HgR) are clustered on the resistance-determinant segment, which consists of multiple transposable elements; inverted repeat termini are designated by arrows pointing outward from the element. A transposon encoding resistance to tetracycline (tetR) is on the resistance-transfer segment. Transposon Tn3 is within Tn4. Each transposon can be transferred independently. (From S. N. Cohen and J. A Shapiro, “Transposable Genetic Elements.” Copyright © 1980 by Scientific American, Inc. All 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements rights reserved.) Figure 20-18. The DNA of a free mu phage has tails derived from its previous host. (From S. N. Cohen and J. A. Shapiro, “Transposable Genetic Elements.” Copyright © 1980 by Scientific American, Inc. All rights reserved.) Figure 20-19. Phage mu can mediate the insertion of phage λ into a bacterial chromosome, resulting in a structure like the one shown here. Figure 20-20. Phage mu can mediate the transposition of a bacterial gene into a plasmid. The selection procedure for detecting the transposition is indicated here. An auxotrophic mutant gene is indicated by aux−. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-21. Phage mu can cause the deletion or inversion of adjacent bacterial segments. (a) The gal region is deleted by transposition of phage mu. (b) The F-factor region of an Hfr strain is inverted by transposition of phage mu. Mechanism of transposition in prokaryotes Several different mechanisms of transposition are employed by prokaryotic transposable elements. And, as we shall see later, eukaryotic elements exhibit still additional mechanisms of transposition. In E. coli, we can identify replicative and conservative (nonreplicative) modes of transposition. In the replicative pathway, a new copy of the transposable element is generated in the transposition event. The results of the transposition are that one copy appears at the new site and one copy remains at the old site. In the conservative pathway, there is no replication. Instead, the element is excised from the chromosome or plasmid and is integrated into the new site. Replicative transposition When transposition is from one locus to a second locus for certain transposons, a copy of the transposable element is left behind at the first locus. An analysis of transposon mutants revealed an interesting fact about the mechanism of transposition. Using the transposon Tn3 (Figure 20-22), researchers grouped the mutations that prevent transposition into two categories. A trans-recessive class maps in the gene that encodes the transposase enzyme, a catalyst of transposition. A second class of cis-dominant mutations results in the buildup of an intermediate in the transposition process. Figure 20-23 diagrams the transposition pathway in the Tn3 transposition from one plasmid to another. The intermediate is a double plasmid, with both donor and recipient plasmid being fused together. The combined circle resulting from the fusion of two circular elements is termed a cointegrate. Apparently, the mutations in this second class delete a region on the transposon at which a recombination event takes place that resolves cointegrates into two smaller circles. This region, called the internal resolution site (IRS), appears in Figure 20-22. The finding of a cointegrate structure as an intermediate in transposition helped establish a replicative mode of transposition for certain elements. In Figure 20-23, note how the transposable element is duplicated in the fusion event and how the recombination event that resolves the cointegrate into two smaller circles leaves one copy of the transposable element in each plasmid. Conservative transposition Some transposons, such as Tn10, excise from the chromosome and integrate into the target DNA. In these cases, DNA replication of the element does not occur, and the element is lost from the site of the original chromosome. Researchers demonstrated 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements this lack of replication by constructing heteroduplexes of λTn10 derivatives containing the lac region of E. coli. The researchers used DNA from Tn10-lacZ+ and Tn10-lacZ− derivatives. The heteroduplexes, therefore, contain one strand with the wild-type lac region and a second strand with the mutated (Z−) lac region. Figure 20-24 diagrams this part of the experiment. The heteroduplex DNA is used to infect cells that have no lac genes, and transpositions of the TetR Tn10 are selected. Different types of colonies arise from the transposition of a heteroduplex Z−/Z+ carrying transposon (Figure 20-25). If replication takes place (the replicative mode of transposition), all colonies are either completely Lac+ or completely Lac−, because the replication will convert the heteroduplex DNA into two homoduplex daughter molecules. The mechanism by which this conversion takes place will be examined in detail in the next section. However, if the transposition is conservative and does not include replication, each colony arises from a lacZ+/lacZ− heteroduplex. Such colonies are partly Lac+ and partly Lac−. By using media that stain Lac+ and Lac− cells different colors, researchers can observe the Lac+ and Lac− sectors in colonies. Therefore, the determination of whether Tn10 undergoes replicative or conservative transposition can be made by observing whether differently colored sectors exist within the same colony resulting from the transposition. Sectored colonies are observed in a majority of cases (Figure 20-26). Thus, Tn10—and perhaps other transposable elements in E. coli—transpose by excising themselves from the donor DNA and integrating directly into the recipient DNA. Molecular consequences of transposition The molecular consequences of transposition reveal an additional piece of evidence concerning the mechanism of transposition: on integration into a new target site, transposable elements generate a repeated sequence of the target DNA in both replicative and conservative transposition. Figure 20-27 depicts the integration of IS1 into a gene. In the example shown, the integration event results in the repetition of a 9-bp target sequence. Analysis of many integration events reveals that the repeated sequence does not result from reciprocal site-specific recombination (as is the case in phage λ integration; see page 229); rather, it is generated in the process of integration itself. The number of base pairs is a characteristic of each element. In bacteria, 9-bp and 5-bp repeats are most common. The preceding observations have been incorporated into somewhat complicated models of transposition. Most models postulate that staggered cleavages are made at the target site and at the ends of the transposable element by a transposase enzyme that is encoded by the element. One end of the transposable element is then attached by a single strand to each protruding end of the staggered cut. Subsequent steps depend on the mode of transposition (replicative or conservative). Rearrangements mediated by transposable elements 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Transposable elements generate a high incidence of deletions in their vicinity. These deletions emanate from one end of the element into the surrounding DNA (Figure 20-28). Such events, as well as element-induced inversions, can be viewed as aberrant transposition events. Transposons also give rise to readily detectable deletions in which part of the element is deleted together with varying lengths of the surrounding DNA. This process of imprecise excision is now recognized as deletions or inversions emanating from the internal ends of the IR segments of the transposon. The process of precise excision—the loss of the transposable element and the restoration of the gene that was disrupted by the insertion—also occurs, although at very low rates compared with the frequencies of the events just described. MESSAGE Some DNA sequences in bacteria and phages act as mobile genetic elements. They are capable of joining different pieces of DNA and are thus capable of splicing DNA fragments into or out of the middle of a DNA molecule. Some naturally occurring mobile or transposable elements carry antibiotic-resistance genes. Figure 20-22. The structure of Tn3. Tn3 contains 4957 base pairs and encodes three polypeptides: the transposase is required for transposition, the repressor is a protein that regulates the transposase gene (see Chapter 11), and β-lactamase confers ampicillin resistance. Tn3 is flanked by inverted repeats (IR) of 38 base pairs and contains a site designated the internal resolution site necessary for the resolution of 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Tn3 cointegrates. Figure 20-23. Transposition of Tn3 takes place through a cointegrate intermediate. Cointegrates in Tn3 transposition are observed for some internal deletions in the transposon. The correct explanation for this observation is that the cointegrates are intermediates in Tn3 transposition and that their resolution is blocked because the internal deletion has removed an internal resolution site, where recombination takes place. (From F. Heffron, in J. A. Shapiro, ed., Mobile Genetic Elements, pp. 223–260. Copyright © 1983 by Academic Press.) Figure 20-24. Generation of heteroduplex and homoduplex Tn10 elements. The denaturation and reannealing of a mixture of two parental λ phages carrying Tn10 elements that differ only at three single bases in the transposon yields a mixture of heteroduplex and homoduplex products. The base differences in the Z gene allow the 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements ultimate determination of the heteroduplex or homoduplex nature of a cell that has received the Z gene through transposition. (After J. Bender and N. Kleckner, Cell 45, 1986, 801–815.) Figure 20-25. Consequences of conservative and replicative transposition. (a) The heteroduplex or homoduplex nature of DNA (see Figure 20-24) is transposed into a target gene. If the starting DNA is heteroduplex, then the resulting DNA will still be heteroduplex only in a conservative, or nonreplicative, pathway. (b) Because the heteroduplex results in a transposed cell that maintains the heteroduplex nature of the DNA during conservative transposition, colonies will arise that are partly Z+ and partly Z−. However, in a replicative pathway, transposition results in individual cells that are either all Z+ or all Z−, and all the colonies will be either Z+ or 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Z−. Figure 20-27. Duplication of a short sequence of nucleotides in the recipient DNA is associated with the insertion of a transposable element; the two copies bracket the inserted element. Here the duplication that attends the insertion of IS1 is illustrated in a way that indicates how the duplication may come about. IS1 insertion causes a nine-nucleotide duplication. If the two strands of the recipient DNA are cleaved (arrow) at staggered sites that are nine nucleotides apart, as shown in part a, followed by insertion of IS1 between the resulting single-stranded ends, then the subsequent filling in of single strands on each side of the newly inserted element, indicated by red letters in part b, with the right complementary nucleotides could account for the duplicated sequences (boxes). (From S. N. Cohen and J. A. Shapiro, “Transposable Genetic Elements.” Copyright © 1980 by Scientific American, Inc. All rights 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements reserved.) Figure 20-28. Deletion formation mediated by a transposable element. In this example, the transposable element IS1 is shown at a point in the E. coli chromosome near the gal genes. Deletions can be generated from each end of the IS1 element, extending into the neighboring DNA sequences. In cases where the deletions extend into the gal regions, they can be detected as a result of the Gal− phenotype. Review of transposable elements in prokaryotes Let's examine what we have learned so far about prokaryotic transposable elements: 1. There are several different types of transposable elements, including insertion sequences (IS1, IS2, . . . ) and transposons (Tn1, Tn2, . . . ). 2. Two copies of a transposable element can act in concert to transpose the DNA segments in between them. Some of the transposons that confer antibiotic resistance are formed in this manner, with two insertion sequences flanking the genes for antibiotic resistance. 3. Most of the transposable elements have recognizable inverted repeat (IR) structures, some of which can be observed under the electron microscope after denaturation and renaturation. 4. Transposable elements are found in bacterial chromosomes, as well as in plasmids. 5. After insertion into a new site on the DNA, transposable elements generate a short repeated sequence, commonly consisting of 9 or 5 bp. 6. The detailed mechanism of transposition is not known, but two different pathways for transposition have been identified. In some cases, transposition takes place by replication of a new copy of the element into the target site, with one copy being left behind at the original site. In 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements other cases, transposition consists of the excision of the element from the original site and its reintegration into a new site. These two modes of transposition are called replicative and conservative, respectively. Transposable elements have been found in eukaryotes, and some have close similarities to those observed in bacteria, transposing through DNA intermediates. Interestingly, other mobile elements transpose through RNA intermediates and, in certain cases, resemble mammalian retroviruses. Molecular nature of transposable elements in eukaryotes Transposable genetic elements are even more prevalent in eukaryotic chromosomes than in bacterial chromosomes. For instance, from 25 percent to 40 percent of mammalian chromosomes consist of transposable elements that have accumulated in the course of evolution. In addition, half of the spontaneous mutations seen in Drosophila are attributed to the movement and insertion of transposable elements. The phenotype conferred by the transposed element was used to detect the first elements in maize, as well as in bacteria—for instance, the patched kernels in corn and the Gal− mutants in bacteria. Now, most elements are detected by recognizing their characteristic structures after sequencing large chromosomal regions. Some eukaryotic elements, such as the maize Ac and Ds elements, and the Drosophila P elements discussed in a later section, transpose as DNA, as do all prokaryotic elements. However, most eukaryotic elements use a different mechanism, transposing in the same manner as that of RNA viruses. Because many transposable elements appear to be related to single-stranded RNA animal viruses, we shall examine some aspects of these viruses. Retroviruses Retroviruses are single-stranded RNA animal viruses that employ a double-stranded DNA intermediate for replication. The RNA is copied into DNA by the enzyme reverse transcriptase. The life cycle of a typical retrovirus is shown in Figure 20-29. Some retroviruses, such as mouse mammary tumor virus (MMTV) and Rous sarcoma virus (RSV), are responsible for the induction of cancerous tumors. When integrated into host chromosomes as double-stranded DNA, these retroviruses are termed proviruses. Proviruses, like the mu phage in bacteria, can be considered transposable elements, because they can in effect transpose from one location to another. Figure 20-30 depicts transposition by a retroviral mechanism. Retroviruses are structurally similar to transposable elements in other organisms, as we shall see later. Additionally, integration results in the duplication of a short target sequence in the host chromosome. Retrotransposons Transposable elements that utilize reverse transcriptase to transpose through an RNA intermediate are termed retrotransposons. They are widespread among eukaryotes 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements and are generally divided into two classes. Viral retrotransposons have properties similar to those of retroviruses. For instance, they display long terminal repeats, or LTRs, as shown in Figure 20-31. Two examples of viral retrotransposons are the yeast Ty elements, and the Drosophila copia elements. Figure 20-32 shows the structure of one of the Ty elements in yeast: the Ty1 sequence, which is present in approximately 35 copies in the yeast genome. The 330-bp-long termini (terminal sequences), called δ (delta) sequences, are present in about 100 copies in the genome. Yeast δ sequences, as well as Ty elements as a whole, show significant sequence divergence. Ty elements generate a repeated sequence of target DNA (in this case, 5 bp) during transposition, like prokaryotic transposons. Ty elements cause mutations by insertion into different genes in the yeast chromosome. In 1985, Jef Boeke and Gerald Fink use Ty1 to show that the transposition of Ty elements is through an RNA intermediate. Figure 20-33 diagrams the experimental design used by Boeke and Fink and their colleagues to alter a yeast Ty element, cloned on a plasmid. First, a promoter was inserted near the end of an element that could be activated by the addition of galactose to the medium. The use of a galactose-sensitive promoter allows the manipulation of the expression of Ty RNA. Galactose enhances the transcription of Ty RNA. Second, an intron from another yeast gene was introduced into the coding region of the Ty transposon. The addition of galactose greatly increases the frequency of transposition of the altered Ty element. This increased frequency suggests the involvement of RNA, because galactose-stimulated transcription begins at the galactose-sensitive promoter (Figure 20-33). The key experimental result, however, is the fate of the transposed Ty DNA. When the researchers examined the Ty DNA resulting from transpositions, they found that the intron had been removed. Because introns are excised only in the course of RNA processing (see Chapter 10), the transposed Ty DNA must have been copied from an RNA intermediate transcribed from the original Ty element and then processed by RNA–RNA splicing. The DNA copy of the spliced mRNA is then integrated into the yeast chromosome. The copia-like elements of Drosophila constitute at least seven families, ranging in size from 5 kb to 8.5 kb. Members of each family appear at 10–100 positions in the Drosophila genome. Each member carries a long, direct terminal repeat and a short, imperfect inverted repeat (Figure 20-34) and is structurally similar to a yeast Ty element. Copia-like elements also cause a duplication of a characteristic number of base pairs of Drosophila DNA on insertion. Certain classic Drosophila mutations result from the insertion of copia-like and other elements. For example, the white-apricot (wa) mutation for eye color is caused by the insertion of an element from the copia family into the white locus. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-35 shows the similarities in the arrangements of genes between integrated retrovirus DNA and the Ty and copia elements. Note how each of the proteins encoded by the retrovirus have counterparts in Ty and copia transposons. Nonviral retrotransposons: LINES and SINES Nonviral retrotransposons are the most frequently encountered transposons in mammals. The LINEs (long interspersed elements) and SINEs (short interspersed elements; including the Alu sequences) discussed in Chapter 3 are the two most abundant. All of these elements are repeated many times in mammalian genomes. The human genome has from 20,000 to 40,0000 LINE elements and about 500,000 SINE elements, most of which display some sequence divergence. Some idea of the profusion of repetitive elements in the human genome can be obtained from Figure 20-36, which shows the distribution of repetitive elements in the gene for homogentisate 1,2-dioxygenase [the alkaptonuria gene (AKU or HGO)]. Function of transposable elements Transposable elements play a role in the biology of organisms. They cause mutations by insertion into genes and affect the regulation of genes by inserting near promoters. They also provide substrates for genetic rearrangements and thus act as agents of genome evolution. For instance, in E. coli growing in natural environments, a sizable fraction of spontaneous mutations are caused by the IS series of transposable elements. As mentioned earlier, more than half of the spontaneous mutations in Drosophila result from transposable-element insertion. Insertions of IS elements near the promoter region of the bgl operon, which encodes proteins participating in β-glucosidase metabolism, activates transcription in this normally “cryptic” operon. There are numerous other examples of insertions that turn on gene expression. Insertion elements provide portable regions of homology that can serve as substrates for recombination enzymes, creating deletions, duplications, and inversions. There is evidence that transposable element-mediated rearrangements may have played an important role in generating the genomes of different organisms, as well as contributing to new functions by stimulating gene duplication. Transposable elements have accumulated during evolution to the point where they constitute much of the genome of higher cells. For instance, in humans, approximately 30 percent of the genomic DNA consists of transposable elements. The evolution of different antibiotic-resistance-carrying microorganisms is influenced by transposable elements. Recall the transposons that can generate different combinations of resistance-encoding genes on R plasmids that are then transferred to different bacteria. Uses of transposable elements Transposable elements have many uses. In prokaryotes, the antibiotic-resistance marker carried by different transposons serves as a convenient marker. For instance, 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements derivatives of Tn10 are often used to create insertions in the bacterial chromosome. After selection for tetracycline-resistant (Tetr) cells, each of which contains an insertion somewhere in the gene, the position of the insert can be mapped genetically, by following the Tetr phenotype, or physically, by using primers matching the ends of Tn10 and then matching the sequence of the adjacent bacterial DNA with that from the known sequence of the entire genome (in S. typhimurium and E. coli for example). With the use of linked transposons, genes can be transferred from one strain to another easily and can be cloned by selecting for the antibioticresistance marker that is either inserted into or very near the gene of interest. In eukaryotes, transposable elements are also used for generating insertion mutations, mapping them, and facilitating both the cloning of genes and the generation of transgenic organisms. P elements The P element in Drosophila is one of the best examples of exploiting the properties of transposable elements in eukaryotes. This element, shown in Figure 20-37, is 2907 bp long and features a 31-bp inverted repeat at each end. The element encodes a transposase. Although the transposase is required for transposition, it can be supplied by a second element. Therefore, P elements with internal deletions can be mobilized and then remain fixed in the new position in the absence of the second element; thus the P element can serve as a convenient marker. P elements do not utilize an RNA intermediate during transposition and can insert at many different positions in the Drosophila chromosome. The transposition of a P element is controlled by repressors encoded by the element. P elements have been developed as tools for Drosophila much in the same way that transposons such as Tn10 have for bacteria. Namely, P elements can be used to create mutations by insertion, to mark the position of genes, and to facilitate the cloning of genes. P elements can be inserted into genes in vivo, and different phenotypes can be selected. Then, the interrupted gene can be cloned, with the use of P element segments as a probe, a method termed transposon tagging. Primers matching the 31-bp sequence at each end can be used to sequence chromosomal regions adjacent to P element insertion sites. Using P elements to insert genes Gerald Rubin and Allan Spradling showed that P element DNA can be used as an effective vehicle for transferring donor genes into the germ line of a recipient fly. Rubin and Spradling devised the following experimental procedure (Figure 20-38). The recipient genotype is homozygous for the rosy (ry−) mutation, which confers a characteristic eye color. From this strain, embryos are collected at the completion of about nine nuclear divisions. At this stage, the embryo is one multinucleate cell, and the nuclei destined to form the germ cells are clustered at one end. (P elements mobilize only in germ-line cells.) Two types of DNA are injected into embryos of this type. The first is a bacterial plasmid carrying a deleted P element into which the ry+ 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements gene has been spliced. This deleted element is not able to transpose, owing to the deletion; so, as mentioned earlier, a helper plasmid bearing a complete element also is injected. Flies developing from these embryos are phenotypically still rosy mutants, but their offspring include a large proportion of ry+ flies. These ry+ descendants show Mendelian inheritance of the newly acquired ry+ gene, suggesting that it is located on a chromosome. This location was confirmed by in situ hybridization, which shows that the ry+ gene, together with the deleted P element, has been inserted into one of several distinct chromosome locations. None appears exactly at the normal locus of the rosy gene. These new ry+ genes are found to be inherited in a stable fashion. A variation of this method, described in Chapter 13, uses P elements to make transgenic Drosophila by transferring foreign genes into the germ line and then monitoring their expression pattern. Figure 20-29. The life cycle of a retrovirus. Viral RNA is shown in red; DNA, in blue. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-30. Transposition by a retrovirus. By making an RNA transcript that is reverse transcribed into double-stranded DNA, retroviruses effectively transpose on insertion of the new DNA into a different point in the chromosome. Figure 20-31. A schematic representation of a viral retrotransposon. Such elements have two LTRs (long terminal repeats that flank a central region that encodes specific protein functions). (From H. Lodish, D. Baltimore, A. Berk, S. L. Zipursky, P. Matsudaira, and J. Darnell, Molecular Cell Biology, 3d ed., p. 329. Copyright © 1995 by Scientific American Books.) Figure 20-32. The structure of a yeast transposable element. The Ty1 sequence appears approximately 35 times in the yeast genome. It contains two copies of the delta (δ) sequence in direct orientation at each end. Delta appears approximately 100 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements times in the yeast genome. Figure 20-33. Demonstration of transposition through an RNA intermediate. A Ty element is altered by adding a promoter that can be activated by the addition of galactose. Activation of the promoter will increase transcription through the Ty element. Then an intron from another gene is inserted into the Ty element. Because the final product of transposition contains no intron, the intron must have been spliced out of an RNA transcript (see Chapter 10). This splicing must have taken place as shown here, where the primary transcript contains the intron but the final processed mRNA does not. This RNA is then copied by reverse transcriptase and integrated into the chromosomal DNA. (After H. Lodish, D. Baltimore, A. Berk, S. L. Zipursky, P. Matsudaira, and J. Darnell, Molecular Cell Biology, 3d ed., p. 332. Copyright © 1995 by Scientific American Books.) 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-34. Copia-like elements carry long direct terminal repeats. Each repeat makes up about 5 percent of the length of the element. These repeats are shown on an expanded scale below the element to illustrate the presence of short, imperfect inverted repeats (→) at the ends of each long direct repeat and the presence of a few base pairs of duplicate target sequence ( ) flanking the element after insertion. The different genomic copies of the family elements are very similar in structure to one another. (From G. Robin, in J. A. Shapiro, ed., Mobile Genetic Elements, pp. 329–361. Copyright 1983 by Academic Press.) Figure 20-35. A comparison of the genes of integrated retrovirus DNA and the yeast Ty elements and Drosophila copia elements. The four functions encoded by the retroviral DNA have counterparts in the yeast and Drosophila elements shown here. The LTRs are represented by the colored ends of the elements. (From H. Lodish, D. Baltimore, A. Berk, S. L. Zipursky, P. Matsudaira, and J. Darnell, Molecular Cell Biology, 3d ed., p. 329. Copyright © 1995 by Scientific American Books.) 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Figure 20-36. Repetitive elements found in the human gene (HGO) encoding homogentisate 1,2-dioxygenase, the enzyme whose deficiency causes alkaptonuria. The first line diagrams the position of the HGO exons. The location and direction of Alus (blue), SINEs (purple), LINEs (orange), retrotransposon-derived sequences (LTRs, red), and short-sequence repeats (SSRs, maroon) in the HGO sequence are indicated by color. (After B. Granadino, D. Beltrán-Valero de Bernabé, J. M. Fernández-Cañón, M. A. Peñalva, and S. Rodríguez de Córdoba, “The Human Homogentisate 1,2-Dioxygenase (HGO) Gene,” Genomics 43, 1997, 115.) Figure 20-37. P element structure. DNA sequence analysis of the 2.9-kb element reveals a gene, composed of four exons and three introns, that encodes transposase. There is a perfect 31-bp inverted repeat at each terminus. (From G. Robin, in J. A. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Shapiro, ed., Mobile Genetic Elements, pp. 329–361. Copyright 1983 by Academic Press.) Figure 20-38. P element-mediated gene transfer in Drosophila. The rosy+ (ry+) eye-color gene is inserted into a deleted P element (∆P) carried on a bacterial plasmid. At the same time, a helper plasmid bearing an intact P element is used. Both are injected into a ry− embryo, where ry+ transposes with the ∆P element into the chromosomes of the germ-line cells. Review of transposable elements in eukaryotes Let's examine some of the essential points about eukaryotic transposable elements: 1. Transposable elements exist in all cells. Elements in yeast, Drosophila, and maize have been well studied, as have retroviruses in mammalian cells. 2. Some transposable elements can be used as tools for cloning and gene manipulation. For instance, the P elements of Drosophila can be employed to transfer genes into the germ line of a recipient fly. As another example, the T-DNA segment of Ti plasmids (described in Chapter 13) can be used to introduce cloned genes into certain plants. 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements 3. A similarity between eukaryotic transposable elements and their counterparts in prokaryotes is that transposition into a new site generates a short repeated sequence at the target site. 4. A difference between certain eukaryotic and prokaryotic transposable elements lies in the mechanism of transposition. Some eukaryotic transposable elements transpose through an RNA intermediate; prokaryotic elements do not use an RNA intermediate. Summary Nature has devised many different ways of changing the genetic architecture of organisms. We are now beginning to understand the molecular processes behind some of these phenomena. Gene mutation, recombination between chromosomes, and transposition can all be reasonably explained at the DNA level. Far from merely producing genetic waste, all these processes undoubtedly have important roles in evolution. This idea is strengthened through the knowledge that the processes themselves are to a large extent under genetic control: there are genes that affect the efficiency of mutation, recombination, and transposition. Although different mechanisms of transposition are sometimes used, the analogies between the transposable elements of phages, bacteria, and eukaryotes are striking. At present, it is not known if transposons are elements that normally play a role in the day-to-day transactions of the genome, as originally proposed by Barbara McClintock in the 1950s, or if they are pieces of “selfish DNA” that exist for no purpose other than their own survival. Whatever the truth of this matter is, transposons certainly constitute a completely unexpected element of chaos in the genome, which geneticists have already harnessed into their team of analytical procedures. At the evolutionary level, transposons may be important in the sudden leaps that characterize the fossil record. Concept Map Draw a concept map interrelating as many of the following terms as possible. Note that the terms are listed in no particular order. Chapter Integration Problem In Chapter 11 we studied the operon model. Note that for the gal operon the order of transcription of the genes in the operon is E-T-K. Suppose we have five different mutations in galT:gal-1, gal-2, gal-3, gal-4, and gal-5. The following table shows the expression of galE and galK in mutants carrying each of these mutations: 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements In addition, the reversion patterns of these mutations with several mutagens we studied in Chapter 19 are shown in the following table. Here, a “+” indicates a high rate of reversion in the presence of each mutagen, a “−” depicts no reversion, and a “low” indicates a low rate of reversion. Which mutation is most likely to result from the insertion of a transposable element such as IS1 and why? Can you assign the other mutations to other categories? See answer Solution Transposable elements will cause polarity, preventing expression of downstream genes from the point of insertion, but not of upstream genes. Therefore, we would expect the insertion mutation to prevent expression of the galK gene. Three mutations are in this category, gal-1, gal-2, and gal-3. These could be frameshifts, nonsense mutations, or insertions, since each of these mutations can lead to polarity. If we examine the reversion data, however, we can distinguish among these possibilities. Transposable elements revert at low rates spontaneously, and this rate is not stimulated by base analogs, frameshift mutagens, alkylating agents, or UV. On the basis of these criteria, the gal-3 mutation is most likely to result from an insertion, since it reverts at a low rate that is not stimulated by any of the mutagens, gal-1 might be a frameshift, since it does not revert with 2-AP and EMS, but does with ICR191, a frameshift mutagen, and UV. (Refer to Chapter 16 for details of each mutagen.) Likewise, gal-2 is probably a frameshift, since it reverts only with ICR191. The gal-4 mutation is probably a deletion, since it is not stimulated to revert at all. The gal-5 mutation appears to be a base substitution, since it reverts with 2-AP, but not above the spontaneous background rate with ICR191. Solved Problem 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements Transposable elements have been referred to as “jumping genes” because they appear to jump from one position to another, leaving the old locus and appearing at a new locus. In light of what we now know concerning the mechanism of transposition, how appropriate is the term “jumping genes” for bacterial transposable elements? See answer Solution In bacteria, transposition takes place by two different modes. The conservative mode results in true jumping genes, because, in this case, the transposable element excises from its original position and inserts at a new position. A second mode is termed the replicative mode. In this pathway, transposable elements move to a new location by replicating into the target DNA, leaving behind a copy of the transposable element at the original site. When operating by the replicative mode, transposable elements are not really jumping genes, because a copy does remain at the original site. Problems 1. Suppose that you want to determine whether a new mutation in the gal region of E. coli is the result of an insertion of DNA. Describe two physical experiments that would allow you to demonstrate the presence of an insertion. 2. Explain the difference between the replicative and conservative modes of transposition. Briefly describe an experiment demonstrating each of these modes in prokaryotes. 3. Describe the generation of multiple drug-resistance plasmids. See answer R plasmids are the main carriers of drug resistance. They acquire these genes by transposition of drug-resistance genes located between IR (inverted repeat) sequences. Once in a plasmid, the transposon carrying drug resistance can be transferred on conjugation if it stays in the R plasmid or it can insert into the host chromosome. 4. Briefly describe the experiment that demonstrates that the transposition of the Ty1 element in yeast takes place through an RNA intermediate. See answer Boeke, Fink, and their co-workers demonstrated that transposition of the Ty element in yeast requires an RNA intermediate. They constructed a plasmid by using a Ty element that had a promoter that could be activated by galactose and an intron inserted into its coding region. First, the frequency of transposition was greatly increased by the addition of galactose, indicating that an increase in transcription (and production of RNA) was correlated to rates of transposition. More importantly, after transposition, they found that the newly transposed Ty DNA lacked the intron sequence. Because intron splicing takes place only in RNA processing, there must have been an RNA intermediate in the transposition event 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements 5. Explain how the properties of P elements in Drosophila make possible gene-transfer experiments in this organism. See answer P elements are transposable elements found in Drosophila. Under certain conditions they are highly mobile and can be used to generate new mutations by random insertion and gene knockout. As such, they are a valuable tool to tag and then clone any number of genes. (See Problem 15 from Chapter 13 for a discussion on cloning by tagging.) P elements can also be manipulated and used to insert almost any DNA (or gene) into the Drosophila genome. P element–mediated gene transfer works by inserting the DNA of interest between the inverted repeats necessary for P element transposition and injecting this recombinant DNA along with helper intact P element DNA (to supply the transposase) into very early embryos and screening for (random) insertion among the injected fly's offspring. 6. When Rhoades took pollen from wholly pigmented anthers on plants of genotype a1/a1 ; Dt/Dt and used this pollen to pollinate a1/a1 ; dt/dt tested females, he found wholly pigmented kernels and, in addition, some dotted kernels. Explain the origin of both phenotypes. 7. In Drosophila, M. Green found a singed allele (sn) with some unusual characteristics. Females homozygous for this X-linked allele have singed bristles, but they have numerous patches of sn+ (wild-type) bristles on their heads, thoraxes, and abdomens. When these flies are mated with sn males, some females give only singed progeny, but others give both singed and wild-type progeny in variable proportions. Explain these results. See answer element. The best explanation is that the mutation is due to an insertion of a transposable 8. Crown gall tumors are found in many dicotyledonous plants infected by the bacterium Agrobacterium tumefaciens. The tumors are caused by the insertion of DNA from a large plasmid carried by the bacterium into the plant DNA. Suppose that a tobacco plant of type A (there are many types of tobacco plants) is infected, and it produces tumors. You remove tumor tissue and grow it on a synthetic medium. Some of these tumor cultures produce aerial shoots. You graft one of these shoots onto a normal tobacco plant of type B, and the graft grows to an apparently normal A-type shoot and flowers. a. You remove cells from the graft and place them in synthetic medium, where they grow like tumor cells. Explain why the graft appears to be normal. b. When seeds are produced by the graft, the resulting progeny are normal A-type plants. No trace of the inserted plasmid DNA remains. Propose a possible explanation for this “reversal.” a. and b. The soil bacterium Agrobacterium tumefaciens contains a See answer large plasmid called the Ti (tumor-inducing) plasmid. When this bacterium infects a plant, a region of the Ti plasmid called the T-DNA is transferred and inserted randomly into the plant's genome. The T-DNA directs the synthesis of plant hormones that cause uncontrolled growth (a tumor) and also directs the plant's synthesis of compounds called opines. (These compounds cannot be metabolized by the plant but are used by the bacterium.) 免疫学信息网 http://www.immuneweb.com Transposable Genetic Elements When a piece of “normal” plant tissue is cultured with appropriate nutrients and growth hormones, cells are stimulated to divide in a disorganized manner, forming a mass of undifferentiated cells called a callus. These cells will differentiate only into shoots (or roots) if the levels of growth hormones are carefully adjusted. The T-DNA causes undifferentiated growth because it directs the unbalanced synthesis of these same hormones. The fact that some of the infected cultures produced shoots suggests that these cells “lost” the ability to overproduce these hormones. This would be consistent with the loss of the T-DNA (similar to the loss of other transposable elements that is observed in many species). Thus, the A graft would grow normally, and seeds produced by the graft would have no trace of the T-DNA. The fact that cells from the A graft grow like tumor cells when placed on synthetic medium suggests that the medium supplies the high levels of hormones necessary for undifferentiated growth even in the absence of T-DNA. 9. Consider two maize plants: a. Genotype C/cm ; Ac/Ac+ where cm is an unstable allele caused by Ds insertion b. Genotype C/cm, where cm is an unstable allele caused by Ac insertion What phenotypes would be produced and in what proportions when (1) each plant is crossed with a base-pair-substitution mutant c/c and when (2) the plant in part a is crossed with the plant in part b? Assume that Ac and c are unlinked, that the chromosome breakage frequency is negligible, and that mutant c/C is Ac+. 免疫学信息网 http://www.immuneweb.com