* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download PSC - University of Pittsburgh
Craniometry wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
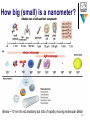
Neurogenomics wikipedia , lookup
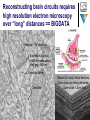
Types of artificial neural networks wikipedia , lookup
Donald O. Hebb wikipedia , lookup
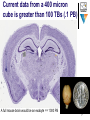
Functional magnetic resonance imaging wikipedia , lookup
Neuroesthetics wikipedia , lookup
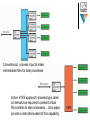
Neural engineering wikipedia , lookup
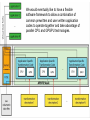
Human multitasking wikipedia , lookup
Haemodynamic response wikipedia , lookup
Aging brain wikipedia , lookup
Development of the nervous system wikipedia , lookup
Human brain wikipedia , lookup
Selfish brain theory wikipedia , lookup
Activity-dependent plasticity wikipedia , lookup
Neuroeconomics wikipedia , lookup
Neural correlates of consciousness wikipedia , lookup
Artificial general intelligence wikipedia , lookup
Neurolinguistics wikipedia , lookup
Time series wikipedia , lookup
Brain Rules wikipedia , lookup
Cognitive neuroscience wikipedia , lookup
Neuroplasticity wikipedia , lookup
Neurophilosophy wikipedia , lookup
Nervous system network models wikipedia , lookup
Brain morphometry wikipedia , lookup
Neuropsychology wikipedia , lookup
Mind uploading wikipedia , lookup
Neuropsychopharmacology wikipedia , lookup
Holonomic brain theory wikipedia , lookup
History of neuroimaging wikipedia , lookup
Neuroanatomy wikipedia , lookup
Metastability in the brain wikipedia , lookup
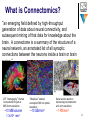
An active processing virtual filesystem for manipulating massive electron microscopy datasets required for connectomics research Art Wetzel - Pittsburgh Supercomputing Center National Resource for Biomedical Supercomputing [email protected] 412-268-3912 www.psc.edu and www.nrbsc.org Source data from … R. Clay Reid, Jeff Lichtman, Wei-Chung Allen Lee Harvard Medical School, Allen Institute for Brain Science Center for Brain Science, Harvard University Davi Bock HMMI Janelia Farm David Hall and Scott Emmons Albert Einstein College of Medicine Aug 30, 2012 Comp Sci Connectomics Data Project Overview 1 What is Connectomics? “an emerging field defined by high-throughput generation of data about neural connectivity, and subsequent mining of that data for knowledge about the brain. A connectome is a summary of the structure of a neural network, an annotated list of all synaptic connections between the neurons inside a brain or brain region.” DTI “tractography” Human Connectome Project at MRI 2 mm resolution ~10 MB/volume 1.3x106 mm3 “Brainbow” stained neuropil at 300 nm optical resolution ~10 GB/mm3 Serial section electron microscopy reconstruction at 3-4 nm resolution ~1 PB/mm3 2 An infant human brain contains ~80 billion neurons. A typical human cortical neuron makes more than 10,000 connections Smaller brains with ~500,000 neurons 3 How big (small) is a nanometer? Below ~10 nm its not anatomy but lots of rapidly moving molecular detail 4 Reconstructing brain circuits requires high resolution electron microscopy over “long” distances == BIGDATA Vesicles ~30 nm diam. A synaptic junction >500 nm wide with cleft gap ~20 nm Dendritic spine Dendrite www.coolschool.ca/lor/BI12/unit12/U12L04.htm Recent ICs have 32nm features 22nm chips are being delivered. Gate oxide 1.2nm thick 5 A10 Tvoxel dataset aligned by our group was an essential part of the March 2011 Nature paper with Davi Bock, Clay Reid and Harvard colleagues Now we are working on two datasets of 100TB each and expect to reach PBs in 2-3 years. 6 Current data from a 400 micron cube is greater than 100 TBs (.1 PB) A full mouse brain would be an exabyte == 1000 PB 7 The CS project is to test a virtual filesystem concept to address common problems with connectomics and other massive datasets. The most important aim is reducing unwanted data duplication as raw data are preprocessed for final analysis. The virtual filesystem addresses this by replacing redundant storage by on-the-fly computing. The second aim is to provide a convenient framework for efficient on-the-fly computation on multidimensional datasets within high performance parallel computing environments using both CPU and GPGPU processing. We are also interested in the image warping and other processes required for neural circuit reconstruction. The Filesystem in User Space mechanism (FUSE) provides a convenient implementation basis that can work on a variety of systems. There are many existing FUSE codes that serve as useful examples. (i.e. scriptfs) 8 One very useful transform is on-the-fly image warping… This example from http://davis.wpi.edu/~matt/courses/morph/2d.htm 9 Conventional: process input to make intermediate files for later processes Active VVFS approach: processing is done on demand as required to present virtual file contents to later processes… Unix pipes provide a restricted subset of this capability 10 We would eventually like to have a flexible software framework to allow a combination of common prewritten and user written application codes to operate together and take advantage of parallel CPU and GPGPU technologies. 11 Multidimensional data structures to provide efficient random and sequential access analogous to the 1D representations provided by standard filesystems will be part of this work. Students will have access to PSC linux machines which access our datasets along with the compilers and other tools required. Basic end-to-end functionality with simple transforms can likely be achieved and may be extended as time permits. Ideally students would have good C/C++, data structures, graphics and OS skills. (biology not required but could be useful) 12