* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download BICH/GENE 431 KNOWLEDGE OBJECTIVES Chapter 9 – Mutations
Comparative genomic hybridization wikipedia , lookup
Nutriepigenomics wikipedia , lookup
DNA profiling wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Frameshift mutation wikipedia , lookup
SNP genotyping wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
DNA vaccination wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Holliday junction wikipedia , lookup
Genomic library wikipedia , lookup
Primary transcript wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Oncogenomics wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Epigenomics wikipedia , lookup
Microsatellite wikipedia , lookup
Molecular cloning wikipedia , lookup
Non-coding DNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Homologous recombination wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
DNA polymerase wikipedia , lookup
DNA supercoil wikipedia , lookup
Genome editing wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
History of genetic engineering wikipedia , lookup
Point mutation wikipedia , lookup
Microevolution wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Helitron (biology) wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Deoxyribozyme wikipedia , lookup
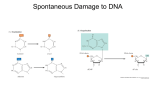
BICH/GENE 431 KNOWLEDGE OBJECTIVES Chapter 9 – Mutations and DNA Repair Three major causes for mutations in DNA: replication errors, chemical or environmental damage, insertion of transposable elements Replication errors at frequency of about 1 in 100,000; accuracy increased about 100fold by proofreading; DNA repair improves accuracy another 100-1000-fold DNA mismatch repair to catch replication errors - MutS, MutL, MutH proteins in E. coli; know this repair pathway - How is incorrect base (strand) recognized in E. coli? - Eukaryotic mismatch repair: MSH is homolog of MutS, interacts with sliding clamp, repairs strand with nick before ligation, mutations in genes lead to higher incidence of colon cancer Spontaneous DNA damage (in water) - deamination: know examples, 5-methylC becomes T (mutagenic hotspot) - depurination Chemical mutagens - alkylating agents (DMS, nitrosamines, MNNG); common product is O6methylguanine - reactive oxygen species (hydrogen peroxide, hydroxide radicals); common product is oxoG UV light causes pyrimidine dimers, such as thymine dimers Ionizing radiation (x rays, gamma rays) cause ds DNA breaks Bleomycin (anti cancer drug) causes ds breaks Base analogs – what are they? A common example is 5-bromouracil (can base pair sometimes with G) Intercalating agents – know examples; insert between bases in DNA to cause insertions or deletions during replication Direct reversal of damage - DNA photolyase to remove thymine dimers (plants, bacteria, not humans) - Methyltransferase enzyme to repair O6-methylguanine (single turnover) Base excision repair – removes base, leaving AP site - glycosylase enzymes are specific for altered base - know pathway of repair - base flipping mechanism used by glycosylases - fail-safe glycosylase for removing A replicated opposite unrepaired oxoG Nucleotide excision repair – removes nucleotide(s) containing abnormal base - UvrABCD pathway in E. coli - XP proteins, plus many others, in humans (XP denotes Xeroderma pigmentosum, a genetic disease caused by defects in nucleotide excision repair) - transcription and nucleotide excision repair are coupled in order to direct repair to genes that are being expressed – TFIIH in eukaryotes is a general transcription factor and a nucleotide excision repair enzyme Double-strand break repair – two major pathways - recombination repair mainly used in bacteria and lower eukaryotes (yeast): details covered in next chapter; uses homologous recombination - NHEJ (nonhomologous end joining) is major pathway in higher eukaryotes – by nature it is mutagenic; identifier proteins are Ku70 and Ku80 that bind to broken ends of DNA Translesion DNA synthesis - bypass synthesis when replication fork approaches unrepaired DNA damage - highly mutagenic, so a method of last resort for cells - know basic mechanism in E. coli - special DNA polymerases in Y family (bacteria to humans) - SOS response in E. coli used to induce TLS enzymes and other DNA repair