* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Exercises Biological databases PART ensembl
Long non-coding RNA wikipedia , lookup
Human genetic variation wikipedia , lookup
Transposable element wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Oncogenomics wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Genomic imprinting wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Minimal genome wikipedia , lookup
Genetic engineering wikipedia , lookup
Metagenomics wikipedia , lookup
Non-coding DNA wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genomic library wikipedia , lookup
Point mutation wikipedia , lookup
X-inactivation wikipedia , lookup
Gene therapy wikipedia , lookup
Copy-number variation wikipedia , lookup
The Selfish Gene wikipedia , lookup
Gene expression profiling wikipedia , lookup
Gene expression programming wikipedia , lookup
Human genome wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Public health genomics wikipedia , lookup
Gene nomenclature wikipedia , lookup
Gene desert wikipedia , lookup
Human Genome Project wikipedia , lookup
Pathogenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Genome editing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Helitron (biology) wikipedia , lookup
Microevolution wikipedia , lookup
Designer baby wikipedia , lookup
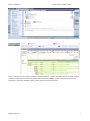
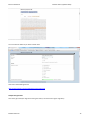
Exercise databases Bioinformatics (updated 2015) EXERCISES BIOLOGICAL DATABASES PART ENSEMBL Exercises Biological databases PART ensembl.......................................................................................................... 1 Discovering genome projects WITH ENSEMBL ............................................................................................................... 1 location VIEW ................................................................................................................................................................. 2 genome view............................................................................................................................................................... 3 Chromosome view ...................................................................................................................................................... 3 Region overview ......................................................................................................................................................... 3 Region in detail ........................................................................................................................................................... 4 comparative genomics ................................................................................................................................................ 8 gene view ...................................................................................................................................................................... 13 Transcript level ............................................................................................................................................................. 16 DISCOVERING GENOME PROJECTS WITH ENSEMBL http://www.ensembl.org/index.html for more tutorials also look at the following sites http://www.ensembl.org/info/website/tutorials/index.html (what is assembly, what is genome annotation) Kathleen Marchal 1 Exercise databases Bioinformatics (updated 2015) In this exercise we will discover more about ensemble, next to ncbi one of the most important databases in molecular biology (the European counterpart of NCBI). Go the the ensembl home page “The Ensembl project produces genome databases for vertebrates and other eukaryotic species, and makes this information freely available online.” Check for which species information is available in ensemble (click view the list of all ensemble species) Search for the human pax 6 gene We find one entry with and ensemble accessionnumber ENSG00000007372. For this gene we have a location view, gene view and transcript view. Study each of the views in detail. LOCATION VIEW Kathleen Marchal 2 Exercise databases Bioinformatics (updated 2015) GENOME VIEW Look at whole genome. This gives you very general information on the assembly and genome build that was used. CHROMOSOME VIEW The chromosome view gives you an overview of the chromosome in which pax 6 is located. REGION OVERVIEW Kathleen Marchal 3 Exercise databases Bioinformatics (updated 2015) REGION IN DETAIL The location view provides a view of the gene on the human chromosome. On which chromosome is the gene? What is the start and stop position of the gene? A detailed region comparison shows a 1 MB region (500000 bp upstream and downstream of the gene pax6). The dark/light blue alternating bars are the contigs that make up the assembly of the genome. You can observe the genes neighbouring pax 6 (> indicates forward strand; < indicates the gene is on the reverse strand). Yellow bars represent coding genes, blue genes represent processed RNAs (so non coding elements), grey genes are pseudogenes. Kathleen Marchal 4 Exercise databases Bioinformatics (updated 2015) The lower panel shows a zoom in on the pax6 gene. Splice variants or alternative transcripts are shown: Open boxes are UTRs, dark boxes are exons and lines are introns. Red or orange are protein coding genes. Blue are unprocessed transcripts. Click on the orange transcript A popup window appears showing details on the transcript. It says that the transcript is confirmed by both ensemble and Havana annotation, so it is a highly relevant transcript. Green transcripts are referred to as resulting from the consensus coding sequence project and they are confirmed by Havana, ensemble, NCBI and USCS genome browser. So they are consensus transcripts. In this view several tracks are shown and turned on by default: these are the %GC, regulatory regions (zoom at the legend, can you pinpoint promotor of pax6 and the transcription factor binding sites?), variants (variants that occur more than 1 % in the 1000 human genome project, variants that have been coupled to a phenotype). Zoom in on some variants (place a box around the variants and click jump to region. What type of variants do you observe? Kathleen Marchal 5 Exercise databases Bioinformatics (updated 2015) Jump back to the original region (e.g. by zooming out). About 400 different tracks are available and most have not been shown. To show more tracks click on the configure wheel. Look at the different tracks you can display. For instance display the tilepaths (these are the clones that were sequenced to make the assembly; they are part of clones and miscellaneous regions). Close the window and check whether they are added. Kathleen Marchal 6 Exercise databases Bioinformatics (updated 2015) Now try to display somatic mutations (what are somatic mutations, how are they different from the other variants?, are there somatic mutations known for pax6? Add the ones from COSMIC). Click on one variant and check the information. Look at the phenotypic data. In which organ the variant was found? Kathleen Marchal 7 Exercise databases Bioinformatics (updated 2015) To save your image, select All information shown in the course can be revised through the following online tutorial http://www.ensembl.org/info/website/tutorials/index.html COMPARATIVE GENOMICS Kathleen Marchal 8 Exercise databases Bioinformatics (updated 2015) Alignment image of the pax 6 region in 12 species or in 8 primates: white regions are unaligned and usually correspond to introns or intergenic regions. Corresponding multiple alignment view Kathleen Marchal 9 Exercise databases Bioinformatics (updated 2015) Region comparison Shows the extent to which the gene neighbourhood of pax6 is aligned. You can select your species in the configure view Kathleen Marchal 10 Exercise databases Bioinformatics (updated 2015) At this moment Human is compared with zebrafish, rat and gorilla. The genomic context of human pax 6 is most similar with the one of gorilla. Kathleen Marchal 11 Exercise databases Bioinformatics (updated 2015) View the detailed alignment; Green regions correspond to aligned regions. The human pax 6 is in the middle panel as it is the reference. Despite its lower conservation of the genomic context, the pax6 gene itself is very conserved at the exon level. Synteny View between mouse and human: shows on which different mouse chromosomes the regions of the chromosome containing the pax6 gene are located (human chromosome is scattered over 2 mouse chromosomes). The human chromosome is in the middle. The small red box indicates the location of pax6. Kathleen Marchal 12 Exercise databases Bioinformatics (updated 2015) Change the species to view the synteny with chimp. This is much higher than with mus. GENE VIEW The gene is identified by a unique Ensembl ID and its common name (HGNCID). Its location on the genome is indicated (Contig Z95332.1.1.20874.). Annotation was performed using an automatic pipeline and manually curated. How many different splice variants are available You view a table and when you scroll down the page you have a graphical overview. Geta detailed view on the splice variants. Kathleen Marchal 13 Exercise databases Bioinformatics (updated 2015) Show the supporting evidence: these are the individual sequences that were used to construct the different gene models and splice variants View sequence (sequendary structure). This provides you a view on the sequence with the exons indicated in red. If you now click “export your sequence from the ensemble browser” you can download the sequence. Kathleen Marchal 14 Exercise databases Bioinformatics (updated 2015) You can select the features you want to down load. Also look at the following tutorial http://www.ensembl.org/info/website/tutorials/index.html Comparative genomics Look at the gene tree (the alignment at the gene level (in contrast to the region alignment) Kathleen Marchal 15 Exercise databases Bioinformatics (updated 2015) The pax6 gene family is a very conserved gene family that did not undergo significant gene losses or gains. How many paralogs has the gene. What are paralogs. What is the difference with orthologs. TRANSCRIPT LEVEL Click on one of the transcripts and explore the transcript centered view. View the schematic view of the transcript. Kathleen Marchal 16 Exercise databases Bioinformatics (updated 2015) Check the supporting evidence Yellow and green are exon based evidence while purple is EST based evidence. The est based evidence supports the untranslated regions while the exon based evidence supports the translated regions View the ontology information Kathleen Marchal 17 Exercise databases Bioinformatics (updated 2015) Gene ontology provides a structured annotation of the gene. It is subdivided in 3 classes, molecular function, localization and biological process. The latter one is the most interesting. View it in a graph structure and try to understand in what functions pax6 is involved. View the presence of protein domains Note on Havana sequences: At the Sanger Institute, automated prediction provided by Ensembl are subjected to high quality manual curation. Finished genomic sequence is analysed on a clone by clone basis using a combination of similarity searches (against various DNA and protein databases) and ab initio gene predictions (Genscan, Fgenes). A sizable proportion of the human genome has been manually annotated by the Havana group. The results of this project are released in the VEGA database. Kathleen Marchal 18