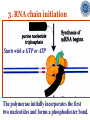
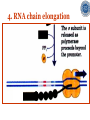
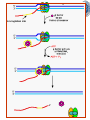
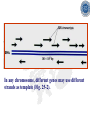
* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Transcription
Transfer RNA wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
X-inactivation wikipedia , lookup
Genetic code wikipedia , lookup
History of genetic engineering wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Metagenomics wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Epigenomics wikipedia , lookup
Human genome wikipedia , lookup
Transposable element wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
DNA supercoil wikipedia , lookup
Microevolution wikipedia , lookup
Point mutation wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Long non-coding RNA wikipedia , lookup
RNA interference wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Messenger RNA wikipedia , lookup
Transcription factor wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
DNA polymerase wikipedia , lookup
Non-coding DNA wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Polyadenylation wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
RNA silencing wikipedia , lookup
Deoxyribozyme wikipedia , lookup
History of RNA biology wikipedia , lookup
Epitranscriptome wikipedia , lookup
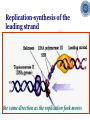
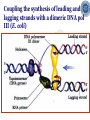
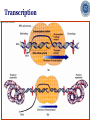
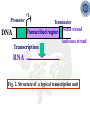
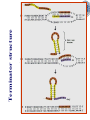
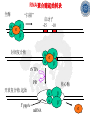
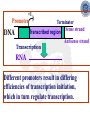
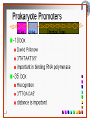
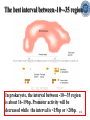
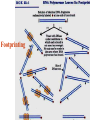
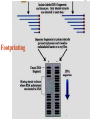
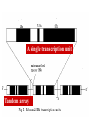
Chapter 5 Transcription A. Transcription in prokaryotes 5.1 Basic principles of transcription An overview, the process of RNA synthesis ( initiation, elongation, termination) 5.2 Escherichia coli RNA polymerase Properties, a subunit, b subunit, b’ subunit, sigma (s) factor 5.3 The E. coli s70 promoter Promoter, s70 size, -10 sequence, -35 sequence, transcription start site, promoter efficiency 5.4 transcription process. Promoter binding, unwinding, RNA chain initiation, elongation, termination (r factor) 5.1: Basic principles of transcription 1. Transcription: an overview (comparison with replication) 2. The process of RNA synthesis: initiation, elongation, termination 5.1-1: Transcription: an overview Key terms defined in this section (Gene VII) +1 upstream Gene X downstream Primary transcript m7Gppp AAAAAn mRNA Coding strand of DNA has the same sequence as mRNA. Downstream identifies sequences proceeding further in the direction of expression; for example, the coding region is downstream of the initiation codon. Upstream identifies sequences proceeding in the opposite direction from expression; for example, the bacterial promoter is upstream from the transcription unit, the initiation codon is upstream of the coding region. Transcription unit is the distance between sites of initiation and termination by RNA polymerase; may include more than one gene. Promoter is a region of DNA involved in binding of RNA polymerase to initiate transcription RNA Terminator is a sequence of DNA, represented at the end of the transcript, that causes RNA polymerase to terminate transcription. RNA polymerases are enzymes that synthesize RNA using a DNA template (formally described as DNA-dependent RNA polymerases). Primary transcript is the original unmodified RNA product corresponding to a transcription unit. Replication: synthesis of two DNA molecules using both parental DNA strands as templates. Duplication of a DNA molecule. 1 DNA molecule 2 DNA molecules Transcription: synthesis of one RNA molecule using one of the two DNA strands as a template. 1 DNA molecule 1 RNA molecule Replication-synthesis of the leading strand the same direction as the replication fork moves Replication- Synthesis of the Okazaki fragments Opposite to the replication fork movement Coupling the synthesis of leading and lagging strands with a dimeric DNA pol III (E. coli) Transcription 1. RNA synthesis occurs in the 5’3’ direction and its sequence corresponds to the sense strand (coding strand). 2. The template of RNA synthesis is the antisense strand (template strand). 3. Phosphodiester bonds: same as in DNA 4. Necessary components: RNA polymerase, transcription factors, rNTPs, promoter & terminator/template DNA 非模板链(编码链), (+)正链 或正义链) DNA 双螺旋 5’-CGCTATAGCGTTTGCAGGCGTTCACGGC-3’ 3’-GCGATATCGCAAACGTCCGCAAGTGCCG-5’ 转录 模板链,负链(-) 或反义链 mRNA (RNA transcript) 5’-CGCUAUAGCGUUUGCAGGCGUUCACGGC-3’ 5.1-2: The process of RNA synthesis 1.initiation 2.elongation 3.termination +1 Promoter DNA Terminator Sense strand Transcribed region Transcription Antisense strand RNA Fig. 2. Structure of a typical transcription unit Before initiation: RNA pol recognizes promoter (RNA聚合酶识别启动子) 1. RNA聚合酶结合到启动子上游附近的 双链DNA模板; 2. 沿双链DNA滑动,找到启动子 (promoter)序列; 3. 负责识别启动子的是RNA聚合酶的σ 亚基; 4. 真核生物的转录在启动子(promoter) 序列处首先结合转录起始因子; 8-18 Initiation (template recognition) 1. Binding of an RNA polymerase to the dsDNA 2. Slide to find the promoter 3. Unwind the DNA helix 4. Synthesis of the RNA strand at the start site (initiation site), this position called position +1 Link Elongation • Covalently adds ribonucleotides to the 3’-end of the growing RNA chain. • The RNA polymerase extend the growing RNA chain in the direction of 5’ 3’ • The enzyme itself moves in 3’ to 5’ along the antisense DNA strand. Link Termination • Ending of RNA synthesis: the dissociation of the RNA polymerase and RNA chain from the template DNA at the terminator site. • Terminator: often contains selfcomplementary regions which can form a stem-loop or hairpin structure in the RNA products Terminator structure 5.2 Escherichia coli RNA polymerase 1. E. coli RNA polymerase 2. a subunit 3. b subunit 4. b’ subunit 5.sigma (s) factor 5.2-1 E. coli RNA polymerase Synthesis of single-stranded RNA from DNA template. RNA polymerase (NMP)n + NTP (NMP)n+1 + PPi 1. Requires no primer for polymerization 2. Requires DNA for activity and is most active with a double-stranded DNA as template. 3. 5’ 3’ synthesis 4. Require Mg2+ for RNA synthesis activity 5. lacks 3’ 5’ exonuclease activity, and the error rate of nucleotides incorporation is 10-4 to 10-5. Is this accuracy good enough for gene expression?? 6. usually are multisubunit enzyme. E. coli polymerase 1. E. coli has a single DNA-directed RNA polymerase that synthesizes all types of RNA. 2. One of the largest enzyme in the cells 3. Consists of at least 5 subunits in the holoenzyme, 2 alpha (a), and 1 of beta (b), beta prime (b’), omega (w) and sigma (s) subunits 4. Shaped as a cylindrical channel that can bind directly to 16 bp of DNA. The whole polymerase binds over 60 bp. 5. RNA synthesis rate: 40 nt per second at 37oC E. coli RNA polymerase 155 KD 36.5 KD 11 KD 36.5 KD 70 KD 151 KD Both initiation & elongation Initiation only E.coli RNA 聚合酶的亚基性质和功能 亚基 基因 相对分子量 亚基数 a rpoA 4.0104 2 core b rpoB 1.51105 1 core b’ w s rpoC ? rpoD 1.55105 11104 7.0104 1 1 1 功能 core assemble, promoter recognition b and b’combined together to form catalyzed center core core unknown s factor different factors recognize different 8-28 promoters 可解离的sigma亚基赋予RNA聚合酶 对原核启动子的特异性 a a b b’ + w s a s b a b’ w 核心酶 Core enzyme 全酶 Holoenzyme 非特异性结合启动 子,并且结合紧密 特异性结合启动子, 结合程度较弱 8-29 RNA聚合酶起始转录 全酶 “扫描” a s b a b’ 封闭复合物 rNTPs 启动子 -35 -10 a s b a b’ PPi 开放复合物; 起始 5’pppA 核心酶 a a mRNA b b’ s 8-30 RNA聚合酶的主要功能 ①识别和结合DNA链上的启动子; ②能沿DNA双链作单向运动; ③解开DNA双螺旋,转录后又恢复双螺旋; ④能同时结合单链DNA和转录产物RNA; ⑤按DNA反义链为模板选择正确的底物NTP, 以5’ → 3’方向催化磷酸二酯键的形成,合 成RNA链; ⑥识别转录的终止信号; ⑦能够与转录因子相互作用,调节转录; ⑧能在转录受到阻遏时进行自我调整,借助 辅助因子,恢复和维持RNA的合成。 8-31 The polymerases of bacteriophage T3 and T7 are smaller single polypeptide chains, they synthesize RNA rapidly (200 nt/sec) and recognize their own promoters which are different from E. coli promoters. RNA polymerase differs from organism to organism 5.2-2: a subunit E. coli polymerase: a subunit • • • • Two identical subunits in the core enzyme Encoded by the rpoA gene Required for assembly of the core enzyme Plays a role in promoter recognition. Experiment: When phage T4 infects E. coli, the α subunit is modified by ADP-ribosylation of an arginine. The modification is associated with a reduced affinity for the promoters formerly recognized by the holoenzyme. • plays a role in the interaction of RNA polymerase with some regulatory factors 大肠杆菌RNA聚合酶: a 亚基 1. 在核心酶中两个a亚基是相同的; 2. a亚基是由rpoA 基因编码; 3. 对于RNA聚合酶核心蛋白的组装是 必需的; 4. 在启动子识别上可能起着重要的作 用; 5.2-3&4: b and b’ subunit 1. b is encoded by rpoB gene, and b’ is encoded by rpoC gene . 2. Make up the catalytic center of the RNA polymerase 3. Their sequences are related to those of the largest subunits of eukaryotic RNA polymerases, suggesting that there are common features to the actions of all RNA polymerases. 4. The b subunit can be crosslinked to the template DNA, the product RNA, and the substrate ribonucleotides; mutations in rpoB affect all stages of transcription. Mutations in rpoC show that b’ also is involved at all stages. b subunit may contain two domains responsible for transcription initiation and elongation •Rifampicin (利福平):has been shown to bind to the β subunit, and inhibit transcription initiation by prokaryotic RNA pol. Mutation in rpoB gene can result in rifampicin resistance. •Streptolydigins(利迪链菌素):resistant mutations are mapped to rpoB gene as well. Inhibits transcription elongation but not initiation. b’ subunit • Binds two Zn 2+ ions and may participate in the catalytic function of the polymerase • Hyparin (肝素):binds to the b’ subunit and inhibits transcription in vitro. • Hyparin competes with DNA for binding to the polymerase. 2. b’ subunit may be responsible for binding to the template DNA . 大肠杆菌RNA聚合酶: b 亚基 1. β亚基由rpoB 基因编码; 2. RNA聚合酶的催化中心; Rifampicin (利福平):与β亚基结合可以抑 制转录的起始,rpoB 基因突变导致对利福平 的抗性; Streptolydigins(利迪链菌素):抗性突变也 定位于 rpoB 基因,它抑制转录延伸,但不抑 制起始; 3. β亚基可能含有两个结构域负责转录的起始和 延伸; 大肠杆菌RNA聚合酶: b’亚基 1. b 亚基由rpoC 基因编码; 2. 与两个Zn2+ 离子结合参与RNA聚合酶的 催化功能; • Heparin (肝素):在体外与 β’ 亚基结合 并抑制转录; • Heparin 与DNA竞争结合RNA聚合酶; 3. β’ 亚基可能是负责与DNA模板的结合; 5.2-5: Sigma (s) factor 1. Many prokaryotes contain multiple s factors to recognize different promoters. The most common s factor in E. coli is s70. 2. Binding of the s factor converts the core RNA pol into the holoenzyme. 3. s factor is critical in promoter recognition, by decreasing the affinity of the core enzyme for non-specific DNA sites (104) and increasing the affinity for the corresponding promoter 4. s factor is released from the RNA pol after initiation (RNA chain is 8-9 nt) 5. Less amount of s factor is required in cells than that of the other subunits of the RNA pol. 大肠杆菌RNA聚合酶: s亚基 1. 在启动子的识别上σ 因子至关重要,对 于非特异性DNA位点核心酶的亲和力 会降低(104),而对于相应的启动子 亲和力会增加; 2. 当起始后(RNA chain is 8-9 nt) ,σ 因子 会从RNA聚合酶中释放出来; 3. 在细胞中σ 因子的需要量比RNA聚合酶 其他亚基的需要量少; 5.3: The E. coli s70 promoter 1. Promoter 2. s70 size 3. -10 sequence 4. -35 sequence 5. transcription start site 6. promoter efficiency 5.3-1: Promoter 1. The specific short conserved DNA sequences: 2. upstream from the transcribed sequence, and thus assigned a negative number (location) 3. required for specific binding of RNA Pol. and transcription initiation (function) 4. Were first characterized through mutations that enhance or diminish the rate of transcription of gene +1 Promoter DNA Terminator Transcribed region Sense strand Transcription Antisense strand RNA Different promoters result in differing efficiencies of transcription initiation, which in turn regulate transcription. 5.3-2,3&4: s70 promoter G TTGACA-----16-18 bp------- TATAAT ---5-8 bp--- C T A -35 sequence -10 sequence +1 • Consists of a sequence of between 40 and 60 bp • -55 to +20: bound by the polymerase • -20 to +20: tightly associated with the polymerase and protected from nuclease digestion by DNaseΙ(see the supplemental) • Up to position –40: critical for promoter function (mutagenesis analysis) • -10 and –35 sequence: 6 bp each, particularly important for promoter function in E. coli -10 sequence (Pribonow box) 1. The most conserved sequence in s70 promoters at which DNA unwinding is initiated by RNA Pol. 2. A 6 bp sequence which is centered at around the –10 position, and is found in the promoters of many different E. coli gene. 3. The consensus sequence is TATAAT. The first two bases (TA) and the final T are most highly conserved. 4. This hexamer is separated by between 5 and 8 bp from position +1, and the distance is critical. -35 sequence: enhances recognition and interaction with the polymerase s factor • A conserved hexamer sequence around position –35 • A consensus sequence of TTGACA • The first three positions (TTG) are the most conserved among E. coli promoters. • Separated by 16-18 bp from the –10 box in 90% of all promoters It can increase to recognize RNA Polyσ 核心启动子区包括 Sextama Box ; -35 区 RNA聚合酶松散结合位点 RNA聚合酶的识别位点 (R to site) It is crucial loose DNA helix TTGAC (Sextama Box) Pribnow Box ; -10 区 RNA聚合酶紧密结合位点 (B site) TATAAT (pribnow Box) Initiation site ; +1 RNA transcriptional startpoint (I site) A/G RNA PolyRNA Poly -35 (R) -10 (B) RNA Poly +1 (I) RNA 8-53 The best interval between -10~-35 region In prokaryote, the interval between -10~-35 region is about 16~19bp. Promoter activity will be decreased while the interval is <15bp or >20bp. 8-54 RNA聚合酶在DNA链上的构型变化 1.全酶与DNA接触时占据 长度为75~80bp,从-55 ~ +20; 2. 进入延伸阶段, 伴随σ 因子的释放,构象发生变 化,覆盖的长度为60 bp; 3. 当新生RNA链聚合15~20bp时,构象进一步发生 转变,形成延伸反应复合物,此时覆盖的长度为 30~40bp的DNA; Supplemental material RNA Polymerase Leaves Its FootPrint on a Promoter • Footprinting is a technique derived from principles used in DNA sequencing. It is used to identify the specific DNA sequences that are bound by a particular protein. Footprinting Footprinting 5.3-5: Transcription start site • Is a purine in 90% of all gene • G is more common at position +1 than A • There are usually a C and T on either side of the start nucleotide (i.e. CGT or CAT) The sequences of five E. coli promoters K3-6: promoter efficiency There is considerable variation in sequence between different promoters, and the transcription efficiency can vary by up to 1000-fold . 1. The –35 sequence constitutes a recognition region which enhances recognition and interaction with the polymerase s factor. 2. The -10 sequence is important for DNA unwinding. 3. The sequence around the start site influence initiation efficiency. 4. The sequence of the first 30 bases to be transcribed controls the rate at which the RNA polymerase clears the promoter, hence influences the rate of the transcription and the overall promoter strength. Weak promoters and activating factor Some promoter sequence are not sufficiently similar to the consensus sequence to be strongly transcribed under normal condition, thus activating factor is required for efficient initiation. Example: Lac promoter P lac requires activating protein, cAMP receptor protein (CRP ), to bind to a site on the DNA close to the promoter sequence in order to enhance polymerase binding and transcription initiation. 5.4 Transcription process 1. Promoter binding 2.DNA unwinding 3.RNA chain initiation 4.RNA chain elongation 5.RNA chain termination (r factor) 1. Promoter binding The searching process is extremely rapidly Closed complex: the initial complex of the polymerase with the base-paired promoter DNA) and –10 region Link The role of s factor in promoter binding • The RNA polymerase core enyzme, a2bb’w, has a general non-specific affinity for DNA, which is referred to as loose binding that is fairly stable. • The addition of s factor to the core enzyme markedly reduces the holoenzyme affinity for non-specific binding by 20 000-fold, and enhances the holoenzyme binding to correct promoter sites 100 times. • Overall, s factor binding dramatically increases the specificity of the holoenzyme for correct promoter-binding site. 2. DNA unwinding +1 The initial unwinding of the DNA results in formation of an open complex with the polymerase, and this process is referred to as tight binding Negative supercoiling & unwinding • It is necessary to unwind the DNA so that the antisense strand to become accessible for base pairing and RNA synthesis. • Negative supercoiling enhances the transcription of many genes, since it facilitates unwinding. Some promoters are not. • Exceptional example: promters for the enzyme subunits of DNA gyrase are inhibited by negative supercoiling, serving as an elegant feedback loop for DNA gyrase expression. 3. RNA chain initiation Starts with a GTP or ATP +1 The polymerase initially incorporates the first two nucleotides and forms a phosphodiester bond. Abortive initiation The first 9 nt are incorporated without polymerase movement along the DNA. Afterward, there is a significant probability that the chain will be aborted. • The RNA pol. goes through multiple abortive initiations before a successful initiation, which limits the overall rate of transcription • The minimum time for promoter clearance is 1-2 seconds (a long event, the synthesis is 40 nt/ sec) 4. RNA chain elongation • Promoter clearance: when initiation succeeds, the enzyme releases s factor and forms a ternary complex of polymerase-DNA-nascent RNA, causing the polymerase to progress along the DNA to allow the re-initiation of transcription. Transcription bubble: 1. containing ~ 17 bp of unwound DNA region and the 3’-end of the RNA that forms a hybrid helix about 12 bp. 2. moves along the DNA with RNA polymerase which unwinds DNA at the front and rewinds it at the rear. 3. The E. coli polymerase moves at an average rate of ~ 40 nt per sec, depending on the local DNA sequence. Transcription bubble 5. RNA chain termination 1.Termination occurs at terminator DNA sequences. 2.The most common stop signal is an RNA hairpin (self-complement structure) commonly GC-rich to favor the structure stability 3. Rho-dependent and Rho-independent Termination. Terminator A specific DNA sequence where the transcription complex dissociate Rho protein (r) independent terminator contains: (1) self-complementary region that is G-C rich and can form a stem-loop or hairpin secondary structure. GC-rich favouring the base pairing stability and causing the polymerase to pause. (2) a run of adenylates (As) in the template strand that are transcribed into uridylates (Us) at the end of the RNA, resulting in weak RNAantisense DNA strand binding. A model for rindependent termination of transcription in E. coli. The A-U base-pairing is less stable that favors the dissociation Rho protein (r) dependent terminator 1. 2. 3. 4. Contains only the self-complementary region Requires r protein for termination r protein binds to specific sites in the singlestranded RNA r protein hydrolyzes ATP and moves along the nascent RNA towards the transcription complex then enables the polymerase to terminate transcription RNA polymerase/transcription and DNA polymerase/replication RNA pol DNA pol dsDNA Require primer dsDNA is better than ssDNA No Initiation promoter origin elongation 40 nt/ sec 900 bp/sec terminator Synthesized RNA Template DNA Template Yes In any chromosome, different genes may use different strands as template (Fig. 25-2). B. Transcription in Eukaryotes 原核与真核基因转录的差异 1)只有一种RNA聚合酶参与原核基因的转录, 真核生物有3种以上的RNA聚合酶负责不同 类型的基因转录。 2)转录产物差别很大,以多顺反子形式存在, 真核转录产物是以单顺反子形式存在成熟 的mRNA只占初始转录产物的一小部分。 3)真核的转录产物需要经过剪接,加工成熟, 而原核转录产物几乎不需要加工。 4)原核转录产物为多顺反子,大多数真核 生物的mRNA是单顺反子; 5)在原核生物细胞中,转录产物可以直接 作为蛋白质合成的模板,转录mRNA与 蛋白质的翻译相互偶联。真核生物细胞 的转录是在细胞核进行的成熟后通过核 孔进入细胞质在此翻译出蛋白质。 5.5 The three RNA Polymerases: characterization and function 5.6 RNA Pol I genes: the ribosomal repeat 5.7 RNA Pol III genes: 5S and tRNA transcription 5.8 RNA Pol II genes: promoters and enhancers 5.9 General transcription factors and RNA Pol II initiation 5.5 The three RNA Polymerases: characterization and function 1. Eukaryotic RNA polymerases 2. RNA polymerase subunits 3. Eukaryotic RNA polymerase activities 4. The CTD of RNA Pol II 5.5-1: Eukaryotic RNA polymerases Main Features of eukaryotic transcription 1. The mechanism of eukaryotic transcription is similar to that in prokaryotes. 2.A lot more proteins are associated with the eukaryotic transcription machinery, which results in the much more complicated transcription. 3. Three eukaryotic polymerases transcribe different sets of genes. The activities of these polymerases are distinguished by their sensitivities to the fungal toxin a-amanitin (鹅膏菌素, 或鹅膏蕈碱). 4. In addition, eukaryotic cells contain additional RNA Pols in mitochondria and chloraplasts. Three eukaryotic polymerases Type Location Substrate a-amanitin RNA Pol I Nucleoli Insensitive Most rRNAs gene RNA Pol II Nucleoplasm All protein-coding Very genes and some sensitive snRNA genes RNA Pol III tRNAs, 5S rRNA, U6 snRNA and other small RNAs Nucleoplasm Moderately sensitive 真核RNA聚合酶活性 Eukaryotic RNA polymerase activity 1)Similarities to that in prokaryotic cells 1. Don’t require a primer 2. Synthesize RNA in a 5’ to 3’ direction. 3. RNA complementary to the antisense template strand. 2) Difference to bacterial polymerases 1. There are 3 RNA polymerases 2. Require more accessory factors for binding promoter DNA & initiating transcription. 3. The C-terminus of RNA Pol II largest subunit contains a stretch of heptapeptide repeats, named as carboxyl terminal domain (CTD) • Amino acid sequence: Tyr-Ser-Pro-Thr-SerPro-Ser. Repeated 26 x (yeast) & 52x in mouse • Involved in polymerase phosphorylation during elongation. 4. The CTD is unphosphorylated at transcription initiation, and phosphorylation occurs during transcription elongation as the RNA Pol II leaves the promoter (In vitro results). 5. Because it transcribes all eukaryotic protein-coding gene, RNA Pol II is the most important RNA polymerase for the study of differential gene expression. The CTD is an important target for differential activation of transcription elongation. 5.5-2: RNA polymerase subunits Each eukaryotic polymerase contains 12 or more subunits. – the two largest subunits are similar to each other and to the b’ and b subunits of E. coli RNA Pol. – There is one other subunit in all three RNA Pol homologous to a subunit of E. coli RNA Pol. – Five additional subunits are common to all three polymerases. – Each RNA Pol contain additional four or seven specific subunit. 5.5-3: RNA polymerase activities 1. Transcription mechanism is similar to that of E. coli polymerase (How?) 2.Different from bacterial polymerasae, they require accessory factors for DNA binding. 5.5-4: The CTD of RNA pol II 1. The C-terminus of RNA Pol II contains a stretch of seven amino acids that is repeated 52 times in mouse enzyme and 26 times in yeast. 2. The heptapeptide sequenc is: Tyr-SerPro-Thr-Ser-Pro-Ser 3. This repeated sequence is known as carboxyl terminal domain (CTD) 4. The CTD sequence may be phosphorylated at the serines and some tyrosines 5. The CTD is unphosphorylated at transcription initiation, and phosphorylation occurs during transcription elongation as the RNA Pol II leaves the promoter (In vitro results). 6. Because it transcribes all eukaryotic protein-coding gene, RNA Pol II is the most important RNA polymerase for the study of differential gene expression. The CTD is an important target for differential activation of transcription elongation. 5.6 RNA Pol I genes: the ribosomal repeats 1-2: Structure of the rRNA genes 1. Ribosomal RNA genes 2. Role of the necleolus 3-6:RNA Pol I promoters & binding factors 3. RNA Pol I promoters 4. Upstream binding factor (UBF) 5. Selectivity factor 1 6. TBP and TAFIs 7: Other rRNA genes 5.6-1&2: Structure of the rRNA genes 1. Ribosomal RNA genes 2. Role of the necleolus Ribosomal RNA Genes & nucleolus 1. A copy of 18S, 5.8S and 28S rRNA genes is organized as a single transcription unit in eukayotes. A 45S rRNA transcript (~13 000 nt long) is produced during transcription, which is then processed into 18S, 5.8S and 28S rRNA. 2. Pre-rRNA transcription units are arranged in clusters in the genome as long tandem arrays separated by nontranscribed spacer squences. A single transcription unit Tandem array 3. Continuous transcription of multiple copies of rRNA genes by RNA Pol I is essential to produce sufficient rRNAs which are packaged into ribosomes. 4. The arrays of rRNA genes (rRNA cluster) loop together to form the nucleolus and are known as nucleolar organizer regions. 5. During active rRNA synthesis, the prerRNA transcripts are packaged along the rRNA genes, visualizing in the electronic microscope as “Christmas tree structures”. Christmas Tree Structures 5.6-3~6: RNA Pol I promoters & binding factors 3. RNA Pol I promoters 4. Upstream binding factor (UBF) 5. Selectivity factor 1 6. TBP and TAFIs RNA Pol I promoters 1. Generally consists of a bipartite sequence in the region preceding the start site, including core element and the upstream control elements (UCE). 2. RNA Pol I promoters in human cells are best characterized. • Core element: -45 to +20, sufficient for transcription initiatiation. • UCE: -180 to -107, to increase the transcription efficiency. • Both regions are rich in G:C, with ~85% identity. RNA Pol I promoters in human cells Two ancillary factors (UBF & SL1) of RNA Pol I & their roles in transcription initiation Upstream binding factor (UBF) • A specific DNA-binding protein that binds to UCE, as well as a different site in the upstream of the core element, causing the DNA to loop between the two sites. (two binding sites have no obvious similarity) • UBF is essential for high level of transcription, and low level of expression occurs in its absence. Selectivity factor 1 (SL1) 1. Does not bind to promoters by itself 2. Binds to and stabilizes the UBF-DNA complex. 3. Interacts with the free downstream part of the core element. 4. Recruit RNA Pol I to bind and to initiate the transcription. Subunits of SL1 SL1 consists of 4 proteins. 1. TBP (TATA-binding protein): a factor also required for initiation by RNA Pol II and III. A critical general factor in eukaryotic transcription that ensures RNA Pol to be properly localized at the startpoint. 2. Other three subunits are referred to as TBP-associated factors (TAFIs) that are specific for RNA Pol I transcription. The initiation complex assembles in three stages The initiation complex UBF UBF TAFIs proposed in your text book TAFIs TBP RNA Pol I TAFIs It is not known which representation one is more accurate. Other rRNA genes (simple) In a simple eukaryote, Acanthamoeba(棘阿米巴属 ), the rRNA genes have only one control element (promoter) around 12-72 bp upstream from the transcription start site. Simple initiation: TIF (homolog of SL-1) binds to the promoter RNA Pol I bind TIF remains bound and the RNA Pol I is released for elongation. 5. 7 RNA Pol III genes: 5S and tRNA transcription 1. RNA polymerase III 2. tRNA genes 3. 5S rRNA genes 4. Alternative RNA Pol III promoters 5. RNA Pol III termination 5.7-1. RNA Pol III 1. Contains at least 16 or more subunits 2. Is located in nucloplasm 3. Synthesizes the precursors of 5S rRNA, the tRNAs and other small nuclear and cytosolic RNAs Promoters for RNA polymerase III May consist of bipartite sequences downstream of the startpoint, with boxA separated from either boxC or boxB. Or they may consist of separated sequences upstream of the startpoint (Oct, PSE, TATA). 5.7-2. tRNA genes 1. The initial transcripts of tRNA genes need to be processed to produce the mature tRNA. 2. The transcription control regions of tRNA lies after the start site within the transcribed region. The two highly conserved control sequences are called A box (5’-TGGCNNAGTGG) and B box (5’-GGTTCGANNCC). • A box and B box also encode important sequences in the tRNA itself, the D-loop and TC-loop. • Therefore, the highly conserved sequence in tRNAs are also highly conserved promoter DNA sequences. 3. Two complex DNA-binding factors required for tRNA transcription initiation: • TFIIIC---binds to both the A and B boxes, an assembly factor for positioning TFIIIB. TFIIIB: (1) binds 50 bp upstream from the A box, but has no sequence specificity and the binding position is determined by the DNA bound TFIIIC. (2) consists of three subunits, one of which is TBP, the general initiation factor; the second is called BRF (TFIIB-related factor); and the third is called B”. TFIIIC: A and B boxes binding and a assembly factor to position TFIIIB TFIIIB: DNA binding and RNA Pol III recruiting 5.7-3 5S rRNA genes 1. Tandemly arranged in a gene cluster. (In human, there is a single cluster of around 2000 genes.) 2. Transcription control regions (promoters) are organized similar to those of tRNA, except that C box is in place of B box. C box: +81-99 bp; A box: +50-65 3. Transcription factors: (1) The C box acts as the binding site for TFIIIA. (2) TFIIIA acts as an assembly factor which allows TFIIIC to interact with the 5S rRNA promoter. (3) The A box may also stabilize TFIIIC binding. (4) TFIIIC is then bound to DNA site near +1. (5) TFIIIB and TFIIIC interact to recruit RNA Pol III to initiate transcription. TFIIIA TFIIIC TFIIIB Pol III 5.7-4 Alternative RNA Pol III promoters Many RNA Pol III genes also rely on upstream sequences for regulation of their transcription e.g. U6 snRNA and Epstein-Barr virus 1. Use only regulatory genes upstream from their transcription start sites. U6 snRNA 1. The coding region contains a characteristic A box that is not required for transcription. 2. The upstream sequence contains sequences typical of RNA Pol II promoters, including a TATA box at bases –30 to –23. 3. Shares several other transcription factor binding sequences with many U RNA genes which are transcribed by RNA Pol II Suggestion: common transcription factors can regulate both RNA Pol II and Pol III genes 5.7-5 RNA Pol III termination The RNA polymerase can terminate transcription without accessory factors. A cluster of A residue is often sufficient for termination. Xenopus borealis terminator: 5’-GCAAAAGC-3’ 5.8 RNA Pol II genes: promoters and enhancers 1. RNA Pol II 2.Cis-acting elements • Promoters • Upstream regulatory elements • Enhancers enhancer Initiation element TATA Inr Upstream element mRNA Down-stream element 5.8-1 RNA Pol II 1. located in nucleoplasm 2. catalyzing the synthesis of the mRNA precursors for all proteincoding genes. 3. RNA Pol Ⅱ-transcribed premRNAs are processed through cap addition, poly(A) tail addition and splicing. 5.8-2 Promoters • Most promoters contain a sequence called the TATA box around 25-35 bp upstream from the start site of transcription. It has a 7 bp consensus sequence 5’-TATA(A/T)A(A/T)-3’. •TBP binds to TATA box that includes an additional downstream bp. 转录起始点 • 转录起始位点没有广泛的序列同源性,但第一个 碱基为腺嘌呤,而两侧是嘧啶碱基。这个区域被 称为起始子(initiator, Inr),序列可表示为 PyPyANT/APyPy。Inr元件位于-2~+4。仅由Inr 元件组成的启动子是具有可被RNA聚合酶II识别的 最简单启动子形式 • 转录起始位点与TATA box一起组成核心启动子; • 上游调控元件具有促进转录作用 转录起始点与核心启动子 ( class II recognized by RNA polymerase II ) l Cap site ; initiation point (+1) Capping m7GpppA/G------70 ±---AUG---(in mRNA) 起始位点区 PyPyANT/(A)PyPy 1-4Kb -70 -30 GC island CAAC/T box Enhancer TATA box +1 Cap UPE core promoter Promoter (basic factor) •TATA box acts in a similar way to an E. coli promoter –10 sequence to position the RNA Pol II for correct transcription initiation. The spacing but not the sequence between the TATA box and the start site is important. Transcription starts with an adenine ~50% of the time. Some eukaryotic genes contain an initiator element instead of a TATA box. The initiator element is located around the transcription start site. Other genes have neither a TATA box nor an initiator element, and usually are transcribed at very low rates. 5.8-3 Upstream regulatory elements • The basal elements (the TATA box and initiator elements) : primarily determine the location of the startpoint, and sponsor initiation only at a rather low level. • Upstream regulatory elements (URE) such as the SP1 box and CCAAT boxes, greatly increase the frequency of initiation. URE is located within 100200 bp from the promoter, and plays an important role in ensuring efficient transcription. 5.8-4 Enhancers Sequence elements which can activate transcription from thousands of base pairs upstream or downstream. General characteristics of Enhancers • Exert strong activation of transcription of a linked gene from the correct start site. • activate transcription when placed in either orientation with respect to linked genes • Able to function over long distances of more than 1 kb whether from an upstream or downstream position relative to the start site. • Exert preferential stimulation of the closets of two tandem promoters 5.9 General transcription factors and RNA PolⅡ initiation 1. RNA Pol II basal transcription factors 2. TFIID (TBP) 3. TFIIA 4. TFIIB and RNA Pol binding 5. Factors binding after RNA Pol. 6. CTD phosphorylation by TFIIH 7. The initiator complex 1. TFIID: Multiprotein Complex, including TBP, other proteins are known as TAFIIs. TBP is the only protein binds to TATA box TBP: 1. a general transcription factor bound to DNA at the TATA box. 2. a general transcription required by all 3 RNA pol. TBP: 3. Has a saddle structure with an overall dyad symmetry. Outer surface (with ?) TBP DNA Inner surface (with ?) TBP 45o Kink TBP causes DNA bending 2. TFIIA • binds to TFIID • stabilizes TFIID-DNA complex • contains at least 3 subunits 3. TFIIB & RNA Pol binding • binds to TFIID •Binds to RNA Pol with TFIIF 4-1 TFIIE binding •Necessary for transcription 4-2 TFIIJ, TFIIH binding •Necessary for transcription 5. phosphorylation of the polymerase CTD by TFIIH Formation of a processive RNA polymerase complex and allows the RNA Pol to leave the promoter region. The initiator transcription complex For TATA-box lacking RNA Pol II promoters, TBP is recruited to the initiator element 0verlapping the start site by some DNA-binding proteins, TBP then recruit the other transcription factors and polymerase similar to TATA box gene transcription.