* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Chromatin Modifications
Gene nomenclature wikipedia , lookup
History of genetic engineering wikipedia , lookup
X-inactivation wikipedia , lookup
Minimal genome wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Genome (book) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Gene desert wikipedia , lookup
Point mutation wikipedia , lookup
Behavioral epigenetics wikipedia , lookup
Epigenetics of cocaine addiction wikipedia , lookup
Genome evolution wikipedia , lookup
Genomic imprinting wikipedia , lookup
Microevolution wikipedia , lookup
Gene expression programming wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Ridge (biology) wikipedia , lookup
Gene expression profiling wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Transcription factor wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenetics of depression wikipedia , lookup
Designer baby wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Epigenetics wikipedia , lookup
Epitranscriptome wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Primary transcript wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Epigenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Nutriepigenomics wikipedia , lookup
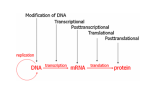
Chromatin Modifications Vered Fishbain Reading Group in Computational Molecular Biology 21/12/2006 Some Definitions… Chromatin is the complex of DNA and proteins found inside the nuclei of eukaryotic cells. Nucleosomes are the fundamental repeating subunits of all eukaryotic chromatin. They are made up of DNA and protein core, which is the histone core. The histone core is composed by two copies of the following set of proteins, called histones: H2A, H2B, H3 and H4. 147 bp in each nucleosome. Heterochromatin is condensed chromatin, includes inactive genes and untranscribed regions (like the centromer). Euchromatin is non-condensed chromatin, includes active and repressed genes. The Histone Core Chromatin Modifications Chromatin modifications are covalent modifications that can effect transcription. Acetylation Methylation Phosphorylation Ubiquitination Sumoylation Adenosine-diphosphate ribosylation Histone Acetylation 1. 2. Associated with transcription activation. Influence gene expression in (at least) two ways: Neutralize Lysine’s positive charge, which can weaken DNA-histone contacts, or histone-histone contacts. Acetyl-Lysine is bound by a specific protein domain that is found in many transcription factors and calls bromodomain. Rapidly reversible, and can turn over rapidly in vivo. Histone Methylation Characterized mainly for histone 3-lysin 4 (H3K4). The Lysine can be mono-, di- or tri-methylated. Doesn’t change the Lysine charge (naturally positive). methyl-Lysine can be bound by a methyl-lysin binding domain, such as chromodomain, WD40 domain, Tudor domain, etc. Long-lived. Research Challenges Absence of sufficient verified data. Contradictory evidences. The available data is in a low resolution. Outline TAF1 as an acetyltransferase (HAT). TAF1 and Gcn5 – is there a redundancy? TAF1 and other HATs in yeast (Durant and Pugh). Acetylation and methylation across promoters and ORFs (Pokholok et al.) High resolution mapping of acetylation and methylation (Liu et al.) Identifying two major groups with similar modification patterns within. Summary (Millar and Grunstein) Genome-Wide Relationships between TAF1 and Histone Acetyltransferases in Saccharomyces cerevisiae Melissa Durant and B. Franklin Pugh Molecular and Cellular Biology, April 2006 The transcription machinery assembles at promoters via two complexes, TFIID and SAGA, which have a compensatory function (Inna’s lecture…). Both complexes contain subunits (TAF1 and Gcn5) that harbor bromodomain and acetyltransferase (HAT) activity. In Saccharomyces cerevisiae, the bromodomains appear on the TFIID-interacting protein Bdf1. Do TAF1 and Gcn5 play redundant role in yeast? H3 Lysines: Gcn5, and not TAF1, is important for bulk H3 acetylation levels. Promoter vs. Non-promoters regions • TAF1 is not a major H3K9, H3K14 acetyltransferase (HAT). • Gcn5 is a HAT at most yeast promoters. Acetylation and Transcription A strong correlation between H3 K9, K14 in W.T and without transcription (without PolII). A little REAL biology… Acetylation of H4K8 is dependant on Elp3, a HAT that is associated with PolII during elongation, while acetylation in other sites in H4 might be less PolII dependent. Same Acetylation Decrease in levelK8in mutant acetylation. and WT. Gcn5 and TAF1 contribution to Gene Expression Recent studies: changes in gene expression for about 25% were observed only when both Gcn5 and TAF1 are eliminated. If Gcn5 and TAF1 each make independent contributions to transcription, the loss of both should be equivalent to the multiplicative result (additive on a log scale) of losing each individually. If the two are functionally redundant, the double mutant should result in an effect that is substantially greater than the multiplicative effects of the individual mutants. Gcn5 and TAF1 contribution to Gene Expression TAF1 and Gcn5 make independent contribution to gene expression No redundancy in TAF1 and Gcn5 function. TAF1 redundancy with other HATs Sas3 Elp3 Hpa2 Their is no (or a very little) Hat1 redundancy between TAF1 and each Esa1 of the 5 tested HATs. Some Other HATs and Acetylation Why there is no effect of any HAT mutant on acetylation? (i) Having highly selective gene targets. (ii) Having Lysine specificities other than those tested. (iii) Making transient contributions. (iv) Being highly redundant with other HATs. TAF1 and Esa1 Esa1 is the main HAT for H4 acetylation of K5, K8, K12. Conclusions Taf1 and Gcn5 have no redundancy. In fact, Taf1 may not be a HAT in yeast. Transcription depends upon acetylation, but acetylation doesn’t depend upon transcription. Gcn5 and Esa1 have a major gene regulatory HATs, but not Hat1, Elp3, Hpa2 and Sas3. Genome-wide Map of Nucleosome Acetylation and Methylation in Yeast Dmitry K. Pokholok, Christopher T. Harbison, Stuart Levine, Megan Cole, Nancy M. Hannett, Tong Ihn Lee, George W. Bell, Kimberly Walker, P. Alex Rolfe, Elizabeth Herbolsheimer, Julia Zeitlinger, Fran Lewitter, David K. Gifford, and Richard A. Young Cell, August 2005 Global Nucleosome Occupancy Nucleosome occupancy at the promoter of CPA1, a gene encoding an amino acidbiosynthetic enzyme. A composite profile of histone occupancy at 5,324 genes. …Surprise! Differential enrichment of intergenic and genic regions also occurred in control experiments lacking antibody. After normalization to the control: No substantial differences in the relative levels of intergenic vs. genic DNA at the average gene, but 40% of the promoters have lower level of histones than their transcribed genes. Is there a correlation between gene expression and nucleosome occupancy? The genes were divided into five classes of transcription level. Before Normalization After Normalization Nucleosome occupancy is reduced maximally at the promoters of active genes. Histone Acetylation Two HATs were checked: Gcn5, which acetylates H3K9 and H3K14, and Esa1, which acetylates the four residues of H4. The acetylation level were measured relative to the histones level. Histone Acetylation – results: Histone Acetylation – Conclusion: There is a positive association between Gcn5, the modifications known to be catalyzed by Gcn5, and transcriptional activity. There is also a positive association between Esa1, the modifications known to be catalyzed by Esa1, and transcriptional activity, although the association is not as strong as that observed for Gcn5. Three interesting trimethylation patterns were observed 1 (Will be discusses later to details…) 2 3 Histone Methylation - conclusions There is a positive correlation between H3K4 trimethylation near the 5’ end of transcribed gene and transcription rate. There is also a positive correlation between H3K36 trimethylation near the 3’ end of transcribed gene, and transcription rate. Somewhat correlation exists between H3K79 trimethylation and transcription rate. http://web.wi.mit.edu/young/nucleosome/ Single-Nucleosome Mapping of Histone Modifications in S. cerevisiae Chih Long Liu, Tommy Kaplan, Minkyu Kim, Stephen Buratowski, Stuart L. Schreiber, Nir Friedman, Oliver J. Rando PLoS Biology, October 2005 For the first time, high-resolution measurement of histone modifications. Acetylation of H4K16 Transcription start site Genes Methylation of H3K4: Gradient from trimethylion in 5’, to dimethylation, and then to mono-metylation on the 3’. Transcriptionindependent modifications Transcriptiondependent modifications Nucleosomes Correlation between modification the matrix of correlations between the 12 modifications shows that there are two groups of strongly correlated acetylations: Tri-methylation of H3K4 correlates with the larger group. Mono- and dimethylation orrelates with the smaller group. Principal Component Analysis PCA 81% of the variance in histone modification patterns is captured by these two principal components. Nucleosomes have continuous variation, both in the total level of acetylation, and in the relative ratio of the two groups of modifications, but they do not show much complexity beyond these two axes. Principal Component Analysis PCA Component #1: Overall level of histone modification. Component #2: Relative levels of two groups of histone modification - the “Transcription dependent modifications” that occur in 5’ to 3’ gradients over coding regions, and the “Transcription - independent modifications” that characterized by short hypo-acetyl domains surrounding TSS. Association Between Chromosomal Location and Histone Modification Promoter Coding region In the PCA plot, it is easy to distinguish between the promoters nucleosomes and the genic nucleosomes. 5’ end Middle 3’ end Moreover, it is possible to distinguish between the promoters nucleosomes and different coding regions (5’, middle and 3’). Conclusion Specific genomic regions are characterized by distinct modification patterns, with little overlap in modification types between the different regions. But… This correlation is imperfect, and it might be due to the different expression level of the genes. Is there a better correlation while separate genes according to the PolII activity level? Highly Transcribed Genes Poorly Transcribed Genes 5’ coding region nucleosomes High PolII activity level Correct classification: 75.4% Medium PolII activity level Low PolII activity level Is there a difference between TSS proximal nucleosomes and TSS distal nucleosomes? TSS proximal Correct nucleosomes classification: 72.8% Modifications occur proximal to transcribed gene contain data about transcription level. TSS distal Correct nucleosomes classification: 58.4% Modifications occur distal to transcribed gene can’t help predict transcription level. Association Between Modifications and Transcription Factor Domains Modification Boundaries Tri-methylation for nucleosome N-1 Tri-methylation for nucleosome N Example of “punctate” nucleosome Conclusions For the first time, modifications mapping in a single-nucleosome resolution. Two distinct groups of acetylation modifications. The modification patterns can be explained by only two principle components. There is no “Histone Code”. Genome-wide patterns of histone modifications in yeast Catherine B. Millar and Michael Grunstein Nature, September 2006 Histone Modification Enzymes Substrate preference: In yeast, all known HMT methylate only one substrate. HATs and HDACs act on several sites, but have distinct preferences. Enzyme targeting: Specific targeting – recruitment by a transcription factor/repressor. This can result in a class-specific modification. Global – function over large regions, irrespective of promoters and coding regions, and without TFs. Global targeting thought to be independent on transcription status. Histone Modification Enzymes – cont. Some HATs function as subunits in a few complexes, one of them has a speciofic targeting and the other has a global targeting. Some HATs have a large but limited region – usually enzymes that are involved in heterochromation formation. No specific HMTs are known to interact with TFs, but some do recruit specifically to coding regions. Histone Modification Enzymes HAT HDAC HMT HDM Gradient of histone modifications in Active Genes Patterns of multiple histone modification K-means clustering – identified groups of at least 20 promoters that have a similar acetylation state at 11 different sites, 53 clusters were defined (kurdistany et al.). The promoters within 55% of these clusters share DNA-sequence motifs, whereas 26% bind similar transcription factors, and 23% of clusters contain promoters that lie upstream of genes that belong to the same functional category. Histone modifications in two different clusters Thanks for your listening, and !!!חנוכה שמח