* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Gene Tagging with Transposons
Deoxyribozyme wikipedia , lookup
DNA vaccination wikipedia , lookup
Primary transcript wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Epigenomics wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Public health genomics wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Oncogenomics wikipedia , lookup
Genomic imprinting wikipedia , lookup
Molecular cloning wikipedia , lookup
Gene nomenclature wikipedia , lookup
Metagenomics wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Gene therapy wikipedia , lookup
Copy-number variation wikipedia , lookup
Genetic engineering wikipedia , lookup
Gene desert wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Human genome wikipedia , lookup
Point mutation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Pathogenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Microsatellite wikipedia , lookup
Gene expression programming wikipedia , lookup
Non-coding DNA wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Minimal genome wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Microevolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genome evolution wikipedia , lookup
Genome editing wikipedia , lookup
Designer baby wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genomic library wikipedia , lookup
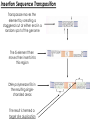
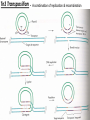
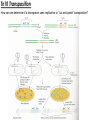
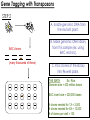
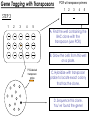
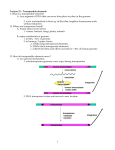
Transposon Introduction • Transposable elements are stretches of DNA that can move to new locations in a genome • These elements can contain genes or be non-coding • Large portions of higher eukaryotes’ genomes are composed of either inert or active transposons (often as repetitive DNA) • Transposons are thus important evolutionarily • Transposons can also be used to isolate genes or introduce foreign genes into cells Bacterial Transposon Discovery • First transposons characterized, these are the simplest • Detected because experimental lac- strains kept reverting back to wild type (ie, colonies kept turning blue) • lac- mutants were due to transposons which then moved back out of the gene Agar w/X-Gal transposon Agar w/X-Gal Transposition event lac gene lac gene lac gene Bacterial Transposons • Can occur in bacterial genome or in plasmid, and can move between these two • Consist of two major types: Insertion Sequences (IS) small, <2500 bp Composite Transposons large, flanked by two IS elements Insertion Sequences • Consists of a pair of inverted terminal repeats at each end (cannot be mutated without loss of transposition activity) • Between this is a stretch of DNA, often containing the gene for transposase – the enzyme that catalyzes transposition • Flanking the terminal repeats are a pair of direct repeats that result from the transposition process Insertion Sequence Transposition Transposase moves the element by creating a staggered cut at either end in a random spot of the genome The IS element then moves then inserts into this region DNA polymerase fills in the resulting singlestranded areas The result is termed a target site duplication Composite Transposons • Denoted Tn • Created when two IS elements insert near each other • The elements can be either in inverse or direct orientation to each other • These two then move together and transpose the sequence between them (often carrying genes) • Movement of these large elements is how bacteria become antibiotic resistance (often using viral intermediates) Composition Transposon Transposition • Involves both IS elements • Two types: Replicative Transposon transposon is copied & moved (ie, a copy remains in place) Non-Replicative Tranposon (aka “cut & paste”) the whole element is moved (no copy remains) • Similar to conserved & semi-conservative DNA replication Tn3 Transposition • A combination of replication & recombination Tn10 Transposition How can we determine if a transposon uses replicative or “cut and paste” transposition? Gene Tagging with Transposons • We can use transposons to tag and isolate genes • Ex: let’s say we want to isolate a blue flower color gene STEP 1 transposon A. Transform plant with a vector containing a transposon. (Thousands of plants needed, depending on genome size) transposon Blue color gene B. Grow progeny from that plant and pick out the mutant phenotype (in this case, an uncolored flower). C. This plant should have the transposon inserted somewhere in the gene. Gene Tagging with Transposons STEP 2 A. Isolate genomic DNA from the mutant plant. BAC clones (many thousands of these) B. Make genomic DNA library from this sample (ex: using BAC vectors). C. Pool clones of the library into 96-well plate. THE MATH Ex: Rice Genome size = 400 million bases BAC insert size = 200,000 bases # clones needed for 1X = 2,000 # clones needed for 6X = 12,000 # of clones per well = 125 Gene Tagging with Transposons PCR w/ transposon primers 1 2 3 4 5 STEP 3 1 2 3 4 5 A. Find the well containing the BAC clone with the transposon (use PCR). B. Grow the cells from this well on a plate. P32-labeled transposon probe C. Hybridize with transposon probe to locate exact colony that has the clone. D. Sequence this clone. You’ve found the gene!