* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Distinct Roles for Drosophila Dicer-1 and Dicer
Gene desert wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Genome evolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Gene therapy wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
X-inactivation wikipedia , lookup
Gene nomenclature wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome (book) wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Nutriepigenomics wikipedia , lookup
History of RNA biology wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Designer baby wikipedia , lookup
Polyadenylation wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Gene expression programming wikipedia , lookup
Gene expression profiling wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Microevolution wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Non-coding RNA wikipedia , lookup
Messenger RNA wikipedia , lookup
Primary transcript wikipedia , lookup
Epitranscriptome wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
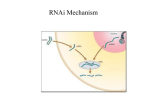
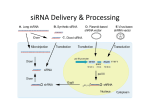
Distinct Roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA Silencing Pathways Lee, S.Y., Nakahara, K., Phan, J.W., Kim, K., He, Z., Sontheimer, E.J., Carthew, R.W. (2004). Cell 117: 69-81 What is RNAi? A cellular mechanism to regulate the expression of genes, mutant gene products and the replication of viruses (some) History of RNAi •1984: Stout & Caskey show antisense RNA can be used to silence gene expression in Mammalian tissue cultures •1990: Fire & Moerman show antisense RNA can disrupt myofilament protein encoding genes •1995: Guo & Kemphues accidentally discover that sense RNA can is as effective as antisense RNA in gene silencing •1998: Mello & Fire illustrate that dsRNA is the agent that leads to potent and specific genetic interference…not ssRNA •2003: Ahringer & Kamath unveil the results of a genome-wide RNAi screen How does it work? • Long dsRNA fragments enter the cell & are diced into small (21-23 nt) fragments known as siRNA by Dicer • siRNA associate with RNAinduced silencing complex (RISC); become unwound + activated • Bind complementary mRNA •mRNA is cleaved by RISC Gratuitous RNAi Movie Another class of small RNA’s can inhibit gene expression: miRNA • Processed from stemloop RNA precursors • Play roles in growth and development • Block translation of mRNA into protein •Act as a guide for a multiprotein complex, miRISC Dicer, an Rnase III, processes both dsRNA and pre-miRNA • Dicer makes staggered cuts (3’ overhangs) to form siRNA • Processes miRNA • Dicer mutants are defective for both transcript degradation and translational repression Implies a dual role for Dicer in miRNA & dsRNA processing Dicer Dicer also required downstream of dsRNA processing depletion of dicer results in reduced effectiveness of injected siRNA Dicer binds to components of RISC (R2D2) & binds tightly to siRNA Role of Dicer in siRISC is not well characterized… The authors took a genetic approach to study Dicer function in Drosophila Screen for RNAi mutants • Engineered Drosophila strain to express an eye-specific hairpin dsRNA corresponding to an exon of White Red eyes to pale orange • Screened for enhanced (white) or suppressed (red) phenotypes • Identified 15 loci that led to stronger pigmentation phenotype • 1 locus had a complementation group of 39 alleles • Mapped locus to Dicer-2 Dicer-2 homozygous suppressor Dicer-2 functions uptstream & downstream of siRNA production •Dcr-2 mutants show marked decrease in siRNA levels Implies dcr-2 required for dsRNA processing • Injected eggs with dsRNA corresponding to bicoid Implies dcr-2 is required for siRNA dependent mRNA degradation • Rapid degradation of bicoid mRNA in wt, but not in dcr-2 mutants Dcr-1 is required for miRNA & siRNA-dependent gene silencing • Dicer-1 mutants have normal levels of wIR siRNAs (dsRNA) • Injected dicer-1 mutant eggs with either dsRNA or siRNA complementary to bicoidmeasured bicoid expression Dcr-1 eggs showed impaired RNAi response to dsRNA and siRNA Dcr-1 is required for miRNA & siRNA-dependent gene silencing • Dcr-1 is required for the formation of mature miRNAs No detectable mature miRNA Dicer-1 is primarily for miRNA production, whereas Dicer-2 is critical for siRNA production RNAi Pathway: To date Dicer 2 What role does dcr-1 and dcr-2 play in the formation of a functional siRISC complex? Dicer R2D2 Dicer R2D2 Dcr-1/2 are required for siRISC formation R1: Dicer-2 and R2D2 proteins bound to siRNAs R2: intermediate complex that links R1 to R3 R3: siRISC complex that is capable of cleaving cognate mRNA *material too large for the gel to resolve How do the Dicers recognize their targets? • Dicer-1 processes pre-miRNA Imperfect basepairing? Low abundance? PAZ domain vs. DExH domain? • Dicer-2 processes dsRNA Perfect basepairing? High abundance? DExH (helicase domain) vs. PAZ domain? Discussion • Dicer-1 is not required to form a stable R1 complex, but is required for the intermediate R2 complex synthesis •Dicer-2 is required for the formation of a stable siRISC complex • Dicer-1 is required for unwinding dsRNA in the siRISC complex (no helicase domain in Dicer-2) • Dicer-2 may cleave siRNA/mRNA duplex Conclusion • Dcr-1, not Dcr-2, is required for gene silencing by miRNAs • Dcr-2 is required for efficient dsRNA processing (but weak phenotype) • Dcr-2 and Dcr-1 may have some redundant properties • Loss of Dcr-1 has effects on growth and patterning; derepresses miRNA target genes Future Directions 1. Make a double mutant 2. Assay mRNA/siRNA cleavage capabilities of Dcr-2 3. Does this complex function analogously in other systems?