* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Protein moonlighting wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Point mutation wikipedia , lookup
Transposable element wikipedia , lookup
Copy-number variation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Genetic engineering wikipedia , lookup
Oncogenomics wikipedia , lookup
Non-coding DNA wikipedia , lookup
Gene nomenclature wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Human genome wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Public health genomics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Gene desert wikipedia , lookup
Essential gene wikipedia , lookup
Genomic library wikipedia , lookup
Metagenomics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Gene expression programming wikipedia , lookup
Helitron (biology) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Pathogenomics wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Genomic imprinting wikipedia , lookup
Ridge (biology) wikipedia , lookup
Genome (book) wikipedia , lookup
Microevolution wikipedia , lookup
Designer baby wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Genome evolution wikipedia , lookup
Minimal genome wikipedia , lookup
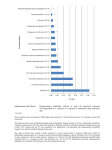
What The Aspergillus Genomes Have Told Us William C. Nierman The Institute for Genomic Research Rockville, MD 5.0 Sp Af Sc Electrophoretic Karyotyping 5 day run 5.7 4.6 1x 4.0 3.5 3.5 1.8 CHEF DRII 1.2% CGA, 1x TAE, 14C, 1.8 V/cm: 2200 s, 48 h; 2200-1800 s, 68 h sizes in Mb Aspergillus fumigatus karyotype 1,789 Kb 3,779 Kb 2,021 Kb 3,992 Kb 4,018 Kb 4,834 Kb 4,891 Kb 3,933* Kb Optical Analysis • Molecule maps generated from images of single DNA molecule digested with NheI • Resolution (avg fragment size) 8.28kb • Total coverage: 8,987 Mbase, or 300x • Total of 8 chromosomes • Total size: 29.189 Megabases A. fumigatus chr5-7 contig placement Aspergillus fumigatus Chromosomes 1 2 1.8 3 4.9 Mb 2.7 2.2 4.8 Mb 3.0 1.3 2.8 4.0 Mb rRNA 4 0.7 0.4 2.5 3.9 Mb 2.6 3.9 Mb 0.3 5 1.2 6 1.3 3.6 Mb 2.5 Mitochondrion 7 0.7 1.3 2.0 Mb 32 Kb 8 0.8 1.0 Presumed centromeric area 1.8 Mb Telomere Nuclear Genome Size (Mb) 29.2 GC Content 49.9% Gene Number 10,034 Mean Gene Length (bp) 1,431 Percent Coding 50.1 Percent Genes with Introns 77.0 Mitochondrial Genome Size (bp) 31,892 GC Content 25.41% Gene Number 16 Mean Gene Length (bp) 1,189 Percent Coding 44.1% Percent Genes with Introns 6.2 tRNA Number 10 Synteny Map of A. fumigatus, A. nidulans, A. oryzae Aspergillus fumigatus Synteny AF293 vs CEA10 TIGR Autoannotation vs Sanger Curated Annotation • • • • • • • • • Status Total Sanger genes analyzed Same gene structure Different gene structure Sanger missing in TIGR annotation Sanger matches multiple TIGR annotations Sanger, TIGR annotations opposite strands TIGR missing in Sanger annotation TIGR matches multiple Sanger annotations Count 360 137 177 37 2 7 12 9 Using Ortholog Clusters to Identify Potential Annotation Problems Using Ortholog Clusters to Identify Potential Annotation Problems Different exon number due to annotation discrepancy In some cases, differences in exon number are real We need to be able to distinguish annotation inconsistencies from real, interesting phenomena Expression profiling analysis to study • Pathogenesis • Response to fungicidal drugs • Temperature-dependent gene expression - A. fumigatus is an environmental species can grow at temperatures as high as 55ºC can survive at temperatures up to 70ºC. - It is commonly isolated from metabolically heated compost heaps The Beast: Microarray Robot from Intelligent Automation <http://www.ias.com> Microarray data analysis Software freely available at, < http://www.tigr.org/software > Hybridization Reference sample Scanning Query sample Multi-experiment comparison Obtain signal intensity values from images Data Normalization and analysis Example hybs (A flip-dye set) Temperature shift experiments • Two shift experiments – 30ºC to 37ºC shift – 30ºC to 48ºC shift • Design – A. fumigatus was grown in a rich medium at 30ºC for two days from conidia, and shifted to 37ºC or to 48ºC. – Samples were taken throughout a time course. • Samples were prepared in Greg May’s Lab 1. A number of genes of various functional roles express differentially at each temperature. 2. More genes are shifted to downregulation than up-regulation at 48˚C in comparison to 30˚C. 3. More genes are turned up at 37˚C when temperature was shifted from 30˚C. This suggests that the fungus has more variety of activities at 37˚C than does at the other temperatures, and it is least active at 48˚C. 30ºC to 37ºC 784 genes heat shock proteins 30ºC to 48ºC 257 genes heat shock proteins Putative virulence genes 1. More heat shock and stress-responsive genes (ex. those coding for heat shock proteins and chaperons) are highly expressed at 48˚C than are at lower temperatures, indicating that the fungus is under heat stress. 2. More putative virulence genes (ex. those coding for the proteins responsive to oxidative stress and host immune system and for toxin production) are highly expressed at 37˚C, although there is no contact with host cells. While predicted function from each gene should be experimentally verified, we suggest from this study that temperature is a key environmental signal for the organism that triggers gene regulation cascades that may ultimately lead to adaptation to a specific new environment. Many transposases, especially those of Mariner-4 type, are highly expressed at 48˚C. It will be interesting to see if the high expression of the transposases actually leads to the transposition events of the transposons. Af ut 1I G A yp fu t2 sy -I G -1yp I_ sy AF G -2yp I sy _ AF 3 G yp - I_ sy AF -4 - I_ AF M I-1 ar in _AF M er-1 ar in _AF M er-2 ar in _AF M er-3 ar in _AF e M ar r-4 _A in M er-4 F ar in a_A e M r-4 F ar b_ in A M er-5 F ar in _AF M er-6 ar in _AF er -7 _ TE AF co _na pi a- me 1I_ AF Transposons in A. fumigatus 25 20 15 10 5 0 Dispersed in the genome Overview – comparative statistics The ortholog was computed by performing an all vs. all BlastP of the three proteomes with a cut-off of 1 x e-15 (no length requirement). The mutual best hits were then organized into clusters based on shared protein nodes. COG A. fumigatus A. Oryzae A. nidulans avg_pctid avg_coverage num_cogs 3 member + + + 70% 86% 5899 + + 65% 84% 967 + 61% 79% 533 + 61% 80% 936 2 member + + #genes included in COG percent of predicted proteome A. fumigatus 7507 79% A. nidulans 7429 75% A. Oryzae 7988 57% Total 22924 68%(22924/33552) Species Aspergillus fumigatus Unique Genes • Vast majority are hypothetical • Includes – – – – – Several transcriptional regulators A chaperonin An hsp 70 related protein ArsC, arsenate reductase Teichoic Acid Biosynthetic Protein Comparative Genomic Hybridization (CGH) • Competitive hybridization between two genomic DNA • Uses microarray to score the presence of genes relative to the reference on the microarray • Provides a quick and easy way of comparing the gene content of a reference organism relative to an unsequenced CLOSE relative A. fumigatus vs. A. fischerianus • Within same cluster by large subunit rRNA analysis • Average DNA identity of ~ 90% based on 4X contigs of A. fischerianus • A. fischerianus rarely identified as a pathogen • A. fischerianus possesses a known sexual cycle A. fumigatus vs. A. fischerianus • Relative to A. fumigatus, A. fischerianus is missing 700 genes – 13 Secondary metabolite genes – 28 Transcription regulators and protein kinases – 21 Transporters – 199 Metabolic and other proteins – 400 Hypothetical proteins CGH between A. fumigatus and A. fischerianus A. fumigatus vs. A. fischerianus Secondary Metabolite Gene Summary • Relative to A. fumigatus, A. fischerianus is missing • 3 of 7 DMAT genes • 6 of 14 PKS genes • 1 of 15 NRPS genes Additional related genomic projects underway or soon to be initiated • Comparative analysis of Aspergillus fumigatus AF293 and CEA10 • Sequencing of Aspergillus flavus • Sequencing of Aspergillus terreus • Sequencing of Aspergillus clavatus • Sequencing of Aspergillus fischerianus • CGH of Neosartorya fennelliae with A. fumigatus • CGH across multiple A. fumigatus strains Aspergillus fumigatus AF293 David Denning Michael Anderson Arnab Pain Goeff Robson Javier Arroyo Goeff Turner David Archer Joan Bennett Matt Berriman Jean Paul Latge Paul Dyer Paul Bowyer Neil Hall Aspergillus nidulans – James Galagan Aspergillus oryzae – Masayuki Machida TIGR Sequencing and Closure Tamara Feldblyum Hoda Khouri Microarray H. Stanley Kim Dan Chen Annotation Jennifer Wortman Jiaqi Huang Resham Kulkarni Natalie Fedrova Claire Fraser NIAID and Dennis Dixon