* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Genome-wide RNAi screening in Caenorhabditis elegans
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Transposable element wikipedia , lookup
Gene nomenclature wikipedia , lookup
Metagenomics wikipedia , lookup
Gene desert wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Non-coding RNA wikipedia , lookup
X-inactivation wikipedia , lookup
Public health genomics wikipedia , lookup
Essential gene wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Nutriepigenomics wikipedia , lookup
History of genetic engineering wikipedia , lookup
Pathogenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Genomic imprinting wikipedia , lookup
Gene expression programming wikipedia , lookup
Genomic library wikipedia , lookup
Ridge (biology) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Microevolution wikipedia , lookup
Minimal genome wikipedia , lookup
Designer baby wikipedia , lookup
Genome (book) wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Epigenetics of human development wikipedia , lookup
RNA silencing wikipedia , lookup
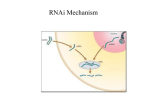
Genome-wide RNAi screening in Caenorhabditis elegans Ravi S. Kamath & Julie Ahringer What is RNAi? A cellular mechanism to regulate the expression of genes, mutant gene products and the replication of viruses (some) History of RNAi •1984: Stout & Caskey show antisense RNA can be used to silence gene expression in Mammalian tissue cultures •1990: Fire & Moerman show antisense RNA can disrupt myofilament protein encoding genes •1995: Guo & Kemphues accidentally discover that sense RNA can is as effective as antisense RNA in gene silencing •1998: Mello & Fire illustrate that dsRNA is the agent that leads to potent and specific genetic interference…not ssRNA •2001: Fraser et al. complete RNAi screen of 90% of chromosome I •2003: Ahringer & Kamath unveil the results of a genome-wide RNAi screen How does this stuff work? The cool movie revisited How do you get dsRNA into C.elegans? Microinjection Soaking in dsRNA Feeding bacteria expressing dsRNA Advantages of feeding for highthroughput RNAi screening •Fast •Cheap •Less labor intensive Disadvantage: Lots of molecular biology work to clone a fragment of a gene into a feeding vector and then transform it into an appropriate bacterial strain Aim of this paper provide research community with a rapid screening tool describe methods for bacterial feeding library construction Identify new gene functions Methods Cloning: Conveniently, Genepairs primers commercially available - optimized for max. overlap with coding region - amplify 1000-1500bp fragments at 5’ end of gene Problem: How do you rapidly get PCR products into vector for high-throughput analysis? How do you screen >19,000 genes rapidly and efficiently for RNAi phenotypes? Construction of feeding library: Making the construct Need dsRNA to yield effective RNAi phenotypes: Used L4440 (pPD129.36) MCS Construction of feeding library: Making the construct • Cut L4440 once with EcoRV and religated •Cut L4440 with EcoRV to create blunt ends for 3’ ddTTP addition by TdT • Recircularized to eliminate non-tailed products • Ligated PCR A-tailed PCR products directly into MCS of vector Construction of feeding library: Suitable Bacterial strain Transformed RNAi constructs into HT115(DE3) •RNase III-deficient strain •Tetracycline resistant •Increased transformation efficiency using TSS Construction of feeding library Plate positive clones onto NGM + Carb + IPTG plates High-throughput phenotype screening 7-10 worms Clone 3 adult worms to 3 separate wells High-throughput phenotype screening: Timeline High-throughput phenotype screening: Analysis of phenotypes Interlude Drawbacks Some genes hard to target 1. Genes whose protein product has a long ½ life 2. Nervous system genes difficult to target Variability in phenotypes 1. Inconsistency between animals 2. Phenotypes can resemble hypomorph rather than amorph Silencing of related genes 1. Genes with close homologs can often be abated in addition to target How rapid is this screen? • Once the operation is a well-oiled machine you can screen 200 genes/day with 3 people • Can screen entire genome in 3 months • Most labor is in manipulating worms & scoring How could you speed up this assay? Results of genome-wide screen & library construction 1. Identified novel gene functions for ~10% of the ~19000 genes screened using N2 worms 2. Created a functional, rapid means to perform a large-scale RNAi screen 3. Now a mutant analysis tool is available to the whole worm community….at a cost $$$ Follow-up Screen •Simmer et al. (2003) used rrf-3 , an RNAi-hypersensitive strain to re-assay the RNAi feeding library 1. Found additional loss-of-function phenotypes for 393 genes 2. In replicates of experiments, found consistent false-positives RNAi screen for novel muscle mutants Muscle ‘Expressome’ Microarray SAGE + Screen for Disorganized Sarcomeres Feed Myo-3::GFP worms RNAi clones RNAi Clone Library Normal myo3 localization Abnormal myo-3 localization Repeat RNAi screen of positive genes to confirm validity Characterize mutants obtained in RNAi screen