* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Assembling and Annotating the Draft Human Genome
Point mutation wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Human genetic variation wikipedia , lookup
Gene therapy wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Copy-number variation wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Genomic imprinting wikipedia , lookup
Gene expression programming wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Ridge (biology) wikipedia , lookup
Primary transcript wikipedia , lookup
Gene desert wikipedia , lookup
Genetic engineering wikipedia , lookup
Oncogenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Gene expression profiling wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Public health genomics wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Microevolution wikipedia , lookup
Transposable element wikipedia , lookup
Pathogenomics wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genomic library wikipedia , lookup
Genome (book) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Non-coding DNA wikipedia , lookup
Designer baby wikipedia , lookup
Helitron (biology) wikipedia , lookup
Human genome wikipedia , lookup
History of genetic engineering wikipedia , lookup
Minimal genome wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome editing wikipedia , lookup
Spaghetti Code, Soupy Logic
Quic kTime™ and a
TIFF (Unc ompres sed) dec ompres sor
are needed to see this pic ture.
Steaming fresh modules in
sourceforge.net
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
Combinatorical assembly of
transcription factors in cell.
Jim Kent - University of California Santa Cruz
A Challenge Every Speaker
Faces:
• Who is the audience?
• Bioinformaticians:
– Biologists with bigger, better databases?
– Geeks trading bits for bases?
– Leading edge interdisciplinary super scientists?
Top 5 Reasons Biologists Go Into
Bioinformatics
5 - Microscopes and biochemistry are so
20th century.
Top 5 Reasons Biologists Go Into
Bioinformatics
5 - Microscopes and biochemistry are so 20th
century.
4 - Got started purifying proteins, but it turns
out the cold room is really COLD.
Top 5 Reasons Biologists Go Into
Bioinformatics
5 - Microscopes and biochemistry are so 20th
century.
4 - Got started purifying proteins, but it turns
out the cold room is really COLD.
3 - After 23 years of school wanted to make
MORE than $23,000/year as a postdoc.
Top 5 Reasons Biologists Go Into
Bioinformatics
5 - Microscopes and biochemistry are so 20th
century.
4 - Got started purifying proteins, but it turns
out the cold room is really COLD.
3 - After 23 years of school wanted to make
MORE than $23,000/year as a postdoc.
2 - Like to swear, @ttracted to $_ Perl #!!
Top 5 Reasons Biologists Go Into
Bioinformatics
5 - Microscopes and biochemistry are so 20th
century.
4 - Got started purifying proteins, but it turns
out the cold room is really COLD.
3 - After 23 years of school wanted to make
MORE than $23,000/year as a postdoc.
2 - Like to swear, @ttracted to $_ Perl #!!
1 - Getting carpel tunnel from pipetting
Top 5 Reasons Computer People
go into Bioinformatics
5 - Bio courses actually have some females.
Top 5 Reasons Computer People
go into Bioinformatics
5 - Bio courses actually have some females.
4 - Human genome more stable than Windows XP
Top 5 Reasons Computer People
go into Bioinformatics
5 - Bio courses actually have some females.
4 - Human genome more stable than Windows XP
3 - Having mastered binary trees, quad trees, and parse
trees ready for phylogenic trees.
Top 5 Reasons Computer People
go into Bioinformatics
5 - Bio courses actually have some females.
4 - Human genome more stable than Windows XP
3 - Having mastered binary trees, quad trees, and parse
trees ready for phylogenic trees.
2 - Missing heady froth of the internet bubble.
Top 5 Reasons Computer People
go into Bioinformatics
5 - Bio courses actually have some females.
4 - Human genome more stable than Windows XP
3 - Having mastered binary trees, quad trees, and parse
trees ready for phylogenic trees.
2 - Missing heady froth of the internet bubble.
1 - Must augment humanity to defeat evil artificial
intelligent robots.
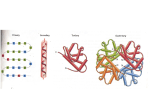
The Paradox of Genomics
How does a long, static, one dimensional string
of DNA turn into the remarkably complex,
dynamic, and three dimensional human body?
GTTTGCCATCTTTTG
CTGCTCTAGGGAATC
CAGCAGCTGTCACCA
TGTAAACAAGCCCAG
GCTAGACCAGTTACC
CTCATCATCTTAGCT
GATAGCCAGCCAGCC
ACCACAGGCATGAGT
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
The Analogy of the Code of Life
• DNA is popularly considered the code of life.
• Computer programs are complex systems that
ultimately are built up of 0’s and 1’s, perhaps they
are a model for a genome built of A,C,G and T?
BUT….
• Human genome lacks documentation, has
accumulated 3 billion years of cruft, and does not
believe in local variables.
• Therefore we must look to less than
straightforward software programs as guides.
Bioperl CORBA module
sub new {
my ( $class, @args) = @_;
my $self = $class->SUPER::new(@args);
my ( $idl, $ior, $orbname ) = $self->_rearrange( [ qw(IDL IOR ORB
@args);
$self->{'_ior'} = $ior || 'biocorba.ior';
$self->{'_idl'} = $idl || $ENV{BIOCORBAIDL} || 'biocorba.idl';
$self->{'_orbname'} = $orbname || 'orbit-local-orb';
$CORBA::ORBit::IDL_PATH = $self->{'_idl'};
my $orb = CORBA::ORB_init($orbname);
my $root_poa = $orb->resolve_initial_references("RootPOA");
$self->{'_orb'} = $orb;
$self->{'_rootpoa'} = $root_poa;
return $self;
}
Obfuscated C
#define c(n,s)case n:s;continue
char x[]="((((((((((((((((((((((",w[]=
"\b\b\b\b\b\b\b\b\b\b\b\b\b\b\b\b\b\b\b\b\b\b";char r[]={92,124,47},l[]={2,3,1
,0};char*T[]={" |"," |","%\\|/%"," %%%",""};char d=1,p=40,o=40,k=0,*a,y,z,g=
-1,G,X,**P=&T[4],f=0;unsigned int s=0;void u(int i){int n;printf(
"\233;%uH\233L%c\233;%uH%c\233;%uH%s\23322;%uH@\23323;%uH \n",*x-*w,r[d],*x+
,r[d],X,*P,p+=k,o);if(abs(p-x[21])>=w[21])exit(0);if(g!=G){struct itimerval t=
{0,0,0,0};g+=((g<G)<<1)-1;t.it_interval.tv_usec=t.it_value.tv_usec=72000/((g>>
3)+1);setitimer(0,&t,0);f&&printf("\e[10;%u]",g+24);}f&&putchar(7);s+=(9-w[21]
)*((g>>3)+1);o=p;m(x);m(w);(n=rand())&255||--*w||++*w;if(!(**P&&P++||n&7936)){
while(abs((X=rand()%76)-*x+2)-*w<6);++X;P=T;}(n=rand()&31)<3&&(d=n);!d&&--*x<=
*w&&(++*x,++d)||d==2&&++*x+*w>79&&(--*x,--d);signal(i,u);}void e(){signal(14,
SIG_IGN);printf("\e[0q\ecScore: %u\n",s);system("stty echo -cbreak");}int main
(int C,char**V){atexit(e);(C<2||*V[1]!=113)&&(f=(C=*(int*)getenv("TERM"))==(
int)0x756E696C||C==(int)0x6C696E75);srand(getpid());system("stty -echo cbreak"
);h(0);u(14);for(;;)switch(getchar()){case 113:return 0;case 91:case 98:c(44,k
=-1);case 32:case 110:c(46,k=0);case 93:case 109:c(47,k=1);c(49,h(0));c(50,h(1
));c(51,h(2));c(52,h(3));}}
Reverse Engineering Microsoft
mouse
blue screen
of death
keyboard
network
elaborate proprietary process
Looks like ‘code’ not enough,
must study actual cells & DNA
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
How DNA is Used by the Cell
Promoter Tells Where to Begin
Different promoters activate different genes in
different parts of the body.
A Computer in Soup
Idealized promoter for a gene involved in making hair.
Proteins that bind to specific DNA sequences in the
promoter region together turn a gene on or off. These
proteins are themselves regulated by their own promoters
leading to a gene regulatory network with many of the
same properties as a neural network.
Genes can be transcription factors that activate
or repress other genes, leading to regulatory networks
such as this one from the development of the central
nervous system. (Image from D’Haeseleer Somogyi 1999)
The Decisions of a Cell
• When to reproduce?
• When to migrate and where?
• What to differentiate into?
• When to secrete something?
• When to make an electrical signal?
The more rapid decisions usually are via the cell
membrane and 2nd messengers. The longer
acting decisions are usually made in the nucleus.
Nucleus Used to Appear Simple
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
• Cheek cells stained with basic dyes. Nuclei are
readily visible.
Mammalian Nuclei Stained in Various Ways
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
Image from Tom Misteli lab
Artist’s rendition of nucleus
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
Image from nuclear protein database
Chromatin
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
Turning on a gene:
• Getting DNA into the right compartment of the
nucleus (may involve very diffuse signals in DNA
over very long distances)
• Loosening up chromatin structure (this involves
activator and repressors which can act over
relatively long distances)
• Attracting RNA Polymerase II to the transcription
start site (these involve relatively close factors
both upstream and downstream of transcription
start).
Methods for Studying Transcription
• Genetics in model organisms
• Promoters hooked to reporter genes
• Gel shifts and DNAse footprinting.
• Phylogenic footprinting
• Motif searches in clusters of coregulated
genes.
Drosophila Genetics
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
normal
antennapedia
mutant
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
Reporter Gene Constructs
promoter to study
easily seen gene
Qui ckTime™ and a
TIFF (U ncompr essed) decompressor
are needed to see thi s pi cture.
Drosophila embryo transfected with ftz promoter hooked
up to lacz reporter gene, creating stripes where ftz promoter
is active.
Biochemical Footprinting
Assays
Gel showing selective
QuickTime™ and a
protection of DNA fromTIFF (Uncompressed)
decompressor
are needed to see this picture.
nuclease digestion
where transcription
factor is bound.
Txn factor
footprint
Comparative Genomics
Webb Miller
Comparative Genomics at BMP10
Conservation of Gene Features
100%
95%
90%
85%
80%
75%
70%
65%
60%
55%
50%
aligning
identity
Conservation pattern across 3165 mappings of human RefSeq mRNAs
to the genome. A program sampled 200 evenly spaced bases across
500 bases upstream of transcription, the 5’ UTR, the first coding exon,
introns, middle coding exons, introns, the 3’ UTR and 500 bases after
polyadenylatoin. There are peaks of conservation at the transition from
one region to another.
Detail Near Translation Start
100%
95%
90%
85%
80%
75%
70%
65%
60%
-15 -14 -13 -12 -11 -10 -9 -8 -7 -6 -5 -4 -3 -2 -1
1
2
3
4
5
6
7
8
9 10 11 12 13 14 15
Note the relatively conserved base 3 before translation
Start (constrained to be a G or an A by the Kozak
Consensus sequence, and the first three translated bases
(ATG).
Normalized eScores
Conservation Levels of
Regulatory Regions in
Human/Mouse Alignments
Conservation in Multiple
Alignments
• As you add more species the phylogenic
footprint gets sharper.
• Currently genome.ucsc.edu shows multiple
alignments between 8 species using Webb
Miller’s multiz program on chained
pairwise alignments.
• The phylogenic tree has to be considered
when calculating conservation levels.
Simple human/rodent tree
human
mouse
rodent
rat
• Mutations that occur in rodent ancestor must be
counted only once
• Ideally should take into consideration varying
mutation rates across species.
• Conservation track at genome.ucsc.edu is based on
Adam Siepel’s PhyloHMM
PhyloHMM on Drosophila
QuickTime™ and a
TIFF (Uncompressed) decompressor
are needed to see this picture.
• Drosophila proteasome alpha 7-1. In many genes
like this one phylogenic footprint suggests promoter
actually is downstream of transcription start site.
Genome Evolution
• Duplication, deletion, and rearrangement is as important to
genome evolution as base-level mutations.
• Much of this is driven by transposons
– Transposon relics are ~50% of genome
– Reverse transcriptase activity from transposons
encourages processed pseudogene formation as well.
– Transposons seed out of place recombination leading to
tandem and segmental duplications, non-processed
pseudogenes.
• Only ~5% of human genome seems functional.
• This messiness provides opportunities for the development
of new genes, but makes understanding the genome a
challenge.
Pseudogene Data from Robert Baertsch, UCSC Grad Student
QuickTime™ and a
TIFF (LZW) decompressor
are needed to see this picture.
Mouse/Human
Rearrangement Statistics
Number of rearrangements of given type per megabase
excluding known transposons.
Chaining Alignments
• Chaining bridges the gulf between syntenic blocks
and base-by-base alignments.
• Local alignments tend to break at transposon
insertions, inversions, duplications, etc.
• Global alignments tend to force non-homologous
bases to align.
• Chaining is a rigorous way of joining together
local alignments into larger structures.
Chains join together related local alignments
Protease Regulatory Subunit 3
Affine penalties are too harsh for long gaps
Log count of gaps vs. size of gaps in mouse/human
alignment correlated with sizes of transposon relics. Affine
gap scores model red/blue plots as straight lines.
Before and After Chaining
Chaining Algorithm
• Input - blocks of gapless alignments from blastz
• Dynamic program based on the recurrence relationship:
score(Bi) = max(score(Bj) + match(Bi) - gap(Bi, Bj))
j<i
• Uses Miller’s KD-tree algorithm to minimize which parts
of dynamic programming graph to traverse. Timing is O(N
logN), where N is number of blocks (which is in hundreds
of thousands)
Netting Alignments
• Commonly multiple mouse alignments can
be found for a particular human region,
particularly for coding regions.
• Net finds best match mouse match for each
human region.
• Highest scoring chains are used first.
• Lower scoring chains fill in gaps within
chains inducing a natural hierarchy.
Net Focuses on Ortholog
Net highlights rearrangements
A large gap in the top level of the net is filled by an
inversion containing two genes. Numerous smaller
gaps are filled in by local duplications and processed
pseudo-genes.
Useful in finding pseudogenes
Ensembl and Fgenesh++ automatic gene predictions
confounded by numerous processed pseudogenes.
Domain structure of resulting predicted protein must
be interesting!
Other tools to cybernetically enhance
your mind at genome.ucsc.edu
Quick Time™a nd a TIFF ( Uncomp res sed) deco mpre ssor are n eede d to s ee this picture .
QuickTi me™ and a T IFF (Uncom pressed) decom pressor are needed to see t his pict ure.
UCSC Gene Sorter
Quick Time™a nd a TIFF ( Uncomp res sed) deco mpre ssor are n eede d to s ee this picture .
• Swiss army knife for dealing with gene sets.
• Presents functional data on genes including
microarray expression information.
• Hilights relationships and connections
between genes.
• Powerful data mining tool.
UCSC Gene Sorter
QuickTime™ and a TIFF (LZW) decompressor are needed to see this picture.
Expression and other information on genes in a big sorted, linked table
A Big Bioinformatics Web Site
• genome.ucsc.edu gets > 100,000 hits by >
5000 scientists each day.
• Involves 570,000 lines of C code, bits of
awk, perl, bash, tcsh, java, r and tcl.
• 1200 CPUs and 12 Terabytes of disk
• 12 full time staff, 18 part time, grad student
and post-doc.
Site Architecture
• 8 web servers running Apache and MySQL
• CGI’s written in C access genome data and
user interface settings in MySQL.
• Genome database is bottleneck, and is
replicated on each server.
• Cluster of 1000 CPUs, and smaller clusters
of faster CPUs create annotation files which
are loaded into database.
Site Sociology
• 1/3 of group telecommutes.
• Thursdays are devoted to reading and testing
each other’s code and if necessary a one or
two hour meeting.
• We develop very incrementally, and do a new
release once a week.
• 1/4 of group is dedicated to quality assurance,
I’m wanting to increase this to 1/3.
• User support is shared by everyone.
Parasol and Kilo Cluster
• UCSC cluster has 1000 CPUs
running Linux
• 1,000,000 BLASTZ jobs in 25
hours for mouse/human
alignment
• We wrote Parasol job
scheduler to keep up.
– Very fast and free.
– Jobs are organized into batches.
– Error checking at job and at
batch level.
Conclusions
• Spaghetti code is not so helpful in understanding
the genome.
• Human genome suggests that trial and error
development is likely to yield a robust version of
windows within 3 billion years.
• Understanding the flow of control in the genome
is a problem that fascinates biologists and
computer scientists alike.
Further Acknowledgements
Individuals
Chuck Sugnet, Angie Hinrichs, Fan
Hsu, Terry Furey, Heather
Trumbower, Kate Rosenbloom,
Hiram Clawson, Brian Raney,
Rachel Harte, Bob Kuhn, Mathieu
Blanchette, Donna Karolchik, David
Haussler
John Sulston, Richard Gibbs, Eric
Lander, Francis Collins,
Roderic Guigo, Michael Brent,
Olivier Jaillon, David Kulp, Victor
Solovyev, Ewan Birney, Greg
Schuler, Deanna Church, Scott
Schwartz, Ross Hardison, and
everyone else!
Institutions
NHGRI, The Wellcome Trust,
HHMI, NCI, Taxpayers in the US
and worldwide.
Baylor, Sanger, Wash U,
Whitehead, Stanford, JGI/ DOE,
Vancouver GSC, UW and the
international sequencing centers.
UCSC, NCBI, EBI, Ensembl,
Genoscope, MGC, Intel, TIGR,
Jackson Labs, Affymetrix,
SwissProt.
THE END