* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download video slide - Course
Transcription factor wikipedia , lookup
History of RNA biology wikipedia , lookup
Transposable element wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Human genome wikipedia , lookup
DNA vaccination wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Oncogenomics wikipedia , lookup
Messenger RNA wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Minimal genome wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Epigenomics wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Genome (book) wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Epitranscriptome wikipedia , lookup
Non-coding RNA wikipedia , lookup
Gene expression profiling wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
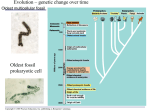
Genome evolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Non-coding DNA wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Point mutation wikipedia , lookup
Helitron (biology) wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Chapter 19 Eukaryotic Genomes: Organization, Regulation, and Evolution PowerPoint Lectures for Biology, Seventh Edition Neil Campbell and Jane Reece Lectures by Chris Romero Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Overview: How Eukaryotic Genomes Work and Evolve • In eukaryotes, the DNA-protein complex, called chromatin – Is ordered into higher structural levels than the DNA-protein complex in prokaryotes Figure 19.1 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Both prokaryotes and eukaryotes – Must alter their patterns of gene expression in response to changes in environmental conditions Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Concept 19.1: Chromatin structure is based on successive levels of DNA packing • Eukaryotic DNA – Is precisely combined with a large amount of protein • Eukaryotic chromosomes – Contain an enormous amount of DNA relative to their condensed length Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Nucleosomes, or “Beads on a String” • Proteins called histones – Are responsible for the first level of DNA packing in chromatin – Bind tightly to DNA • The association of DNA and histones – Seems to remain intact throughout the cell cycle Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • In electron micrographs – Unfolded chromatin has the appearance of beads on a string • Each “bead” is a nucleosome – The basic unit of DNA packing 2 nm DNA double helix Histones Histone tails Histone H1 Linker DNA (“string”) Nucleosome (“bad”) (a) Nucleosomes (10-nm fiber) Figure 19.2 a Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 10 nm Higher Levels of DNA Packing • The next level of packing – Forms the 30-nm chromatin fiber 30 nm Nucleosome (b) 30-nm fiber Figure 19.2 b Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The 30-nm fiber, in turn – Forms looped domains, making up a 300-nm fiber Protein scaffold Loops 300 nm (c) Looped domains (300-nm fiber) Figure 19.2 c Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Scaffold • In a mitotic chromosome – The looped domains themselves coil and fold forming the characteristic metaphase chromosome 700 nm 1,400 nm (d) Metaphase chromosome Figure 19.2 d Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • In interphase cells – Most chromatin is in the highly extended form called euchromatin Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Concept 19.2: Gene expression can be regulated at any stage, but the key step is transcription • All organisms – Must regulate which genes are expressed at any given time • During development of a multicellular organism – Its cells undergo a process of specialization in form and function called cell differentiation Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Differential Gene Expression • Each cell of a multicellular eukaryote – Expresses only a fraction of its genes • In each type of differentiated cell – A unique subset of genes is expressed Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Many key stages of gene expression – Can be regulated in eukaryotic cells Signal NUCLEUS Chromatin Chromatin modification: DNA unpacking involving histone acetylation and DNA demethlation DNA Gene available for transcription Gene Transcription RNA Cap Exon Primary transcript Intron RNA processing Tail mRNA in nucleus Transport to cytoplasm CYTOPLASM mRNA in cytoplasm Degradation of mRNA Translation Polypetide Cleavage Chemical modification Transport to cellular destination Active protein Degradation of protein Figure 19.3 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Degraded protein Regulation of Chromatin Structure • Genes within highly packed heterochromatin – Are usually not expressed Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Histone Modification • Chemical modification of histone tails – Can affect the configuration of chromatin and thus gene expression Chromatin changes Transcription RNA processing mRNA degradation Translation Protein processing and degradation Histone tails DNA double helix Amino acids available for chemical modification Figure 19.4a (a) Histone tails protrude outward from a nucleosome Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Histone acetylation – Seems to loosen chromatin structure and thereby enhance transcription Unacetylated histones Figure 19.4 b Acetylated histones (b) Acetylation of histone tails promotes loose chromatin structure that permits transcription Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings DNA Methylation • Addition of methyl groups to certain bases in DNA – Is associated with reduced transcription in some species Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Epigenetic Inheritance • Epigenetic inheritance – Is the inheritance of traits transmitted by mechanisms not directly involving the nucleotide sequence Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Regulation of Transcription Initiation • Chromatin-modifying enzymes provide initial control of gene expression – By making a region of DNA either more or less able to bind the transcription machinery Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Organization of a Typical Eukaryotic Gene • Associated with most eukaryotic genes are multiple control elements – Segments of noncoding DNA that help regulate transcription by binding certain proteins Enhancer (distal control elements) Poly-A signal Termination sequence region Proximal control elements Exon Intron Exon Intron Exon DNA Downstream Upstream Promoter Chromatin changes Transcription Exon Primary RNA 5 transcript (pre-mRNA) Intron Exon RNA processing: Cap and tail added; introns excised and exons spliced together Transcription Intron RNA RNA processing mRNA degradation Cleared 3 end of primary transport Coding segment Translation Protein processing and degradation Poly-A signal Intron Exon mRNA G P Figure 19.5 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings P P 5 Cap 5 UTR (untranslated region) Start codon Stop codon Poly-A 3 UTR (untranslated tail region) The Roles of Transcription Factors • To initiate transcription – Eukaryotic RNA polymerase requires the assistance of proteins called transcription factors Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Enhancers and Specific Transcription Factors • Proximal control elements – Are located close to the promoter • Distal control elements, groups of which are called enhancers – May be far away from a gene or even in an intron Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • An activator – Is a protein that binds to an enhancer and stimulates transcription of a gene Distal control element Activators Enhancer 1 Activator proteins bind to distal control elements grouped as an enhancer in the DNA. This enhancer has three binding sites. 2 A DNA-bending protein brings the bound activators closer to the promoter. Other transcription factors, mediator proteins, and RNA polymerase are nearby. Promoter Gene TATA box General transcription factors DNA-bending protein Group of Mediator proteins RNA Polymerase II Chromatin changes 3 The activators bind to certain general transcription factors and mediator proteins, helping them form an active transcription initiation complex on the promoter. Transcription RNA processing mRNA degradation Figure 19.6 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings RNA Polymerase II Translation Protein processing and degradation Transcription Initiation complex RNA synthesis • Some specific transcription factors function as repressors – To inhibit expression of a particular gene • Some activators and repressors – Act indirectly by influencing chromatin structure Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Coordinately Controlled Genes • Unlike the genes of a prokaryotic operon – Coordinately controlled eukaryotic genes each have a promoter and control elements • The same regulatory sequences – Are common to all the genes of a group, enabling recognition by the same specific transcription factors Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Mechanisms of Post-Transcriptional Regulation • An increasing number of examples – Are being found of regulatory mechanisms that operate at various stages after transcription Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings RNA Processing • In alternative RNA splicing – Different mRNA molecules are produced from the same primary transcript, depending on which RNA segments are treated as exons and which as introns Chromatin changes Transcription RNA processing mRNA degradation Translation Protein processing and degradation Exons DNA Primary RNA transcript RNA splicing Figure 19.8 mRNA Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings or mRNA Degradation • The life span of mRNA molecules in the cytoplasm – Is an important factor in determining the protein synthesis in a cell – Is determined in part by sequences in the leader and trailer regions Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • RNA interference by single-stranded microRNAs (miRNAs) – Can lead to degradation of an mRNA or block its translation 1 The microRNA (miRNA) precursor folds back on itself, held together by hydrogen bonds. 22 An enzyme called Dicer moves along the doublestranded RNA, cutting it into shorter segments. 3 One strand of each short doublestranded RNA is degraded; the other strand (miRNA) then associates with a complex of proteins. 4 The bound miRNA can base-pair with any target mRNA that contains the complementary sequence. 55 The miRNA-protein complex prevents gene expression either by degrading the target mRNA or by blocking its translation. Chromatin changes Transcription RNA processing mRNA degradation Translation Protein processing and degradation Protein complex Dicer Degradation of mRNA OR miRNA Target mRNA Figure 19.9 Hydrogen bond Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Blockage of translation Initiation of Translation • The initiation of translation of selected mRNAs – Can be blocked by regulatory proteins that bind to specific sequences or structures of the mRNA • Alternatively, translation of all the mRNAs in a cell – May be regulated simultaneously Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Protein Processing and Degradation • After translation – Various types of protein processing, including cleavage and the addition of chemical groups, are subject to control Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Proteasomes – Are giant protein complexes that bind protein molecules and degrade them 3 Enzymatic components of the 1 Multiple ubiquitin molChromatin changes ecules are attached to a protein by enzymes in the cytosol. 2 The ubiquitin-tagged protein is recognized by a proteasome, which unfolds the protein and sequesters it within a central cavity. proteasome cut the protein into small peptides, which can be further degraded by other enzymes in the cytosol. Transcription RNA processing mRNA degradation Proteasome and ubiquitin to be recycled Ubiquitin Translation Proteasome Protein processing and degradation Protein to be degraded Ubiquinated protein Protein entering a proteasome Figure 19.10 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Protein fragments (peptides) • Concept 19.3: Cancer results from genetic changes that affect cell cycle control • The gene regulation systems that go wrong during cancer – Turn out to be the very same systems that play important roles in embryonic development Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Types of Genes Associated with Cancer • The genes that normally regulate cell growth and division during the cell cycle – Include genes for growth factors, their receptors, and the intracellular molecules of signaling pathways Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Oncogenes and Proto-Oncogenes • Oncogenes – Are cancer-causing genes • Proto-oncogenes – Are normal cellular genes that code for proteins that stimulate normal cell growth and division Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • A DNA change that makes a proto-oncogene excessively active – Converts it to an oncogene, which may promote excessive cell division and cancer Proto-oncogene DNA Translocation or transposition: gene moved to new locus, under new controls Gene amplification: multiple copies of the gene New promoter Normal growth-stimulating protein in excess Normal growth-stimulating protein in excess Figure 19.11 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Point mutation within a control element Point mutation within the gene Oncogene Oncogene Normal growth-stimulating protein in excess Hyperactive or degradationresistant protein Tumor-Suppressor Genes • Tumor-suppressor genes – Encode proteins that inhibit abnormal cell division Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Interference with Normal Cell-Signaling Pathways • Many proto-oncogenes and tumor suppressor genes – Encode components of growth-stimulating and growth-inhibiting pathways, respectively Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The Ras protein, encoded by the ras gene – Is a G protein that relays a signal from a growth factor receptor on the plasma membrane to a cascade of protein kinases 1 Growth factor MUTATION Ras GTP 3 (a) Cell cycle–stimulating pathway. This pathway is triggered by 1 a growth factor that binds to 2 its receptor in the plasma membrane. The signal is relayed to 3 a G protein called Ras. Like all G proteins, Ras is active when GTP is bound to it. Ras passes the signal to 4 a series of protein kinases. The last kinase activates 5 a transcription activator that turns on one or more genes for proteins that stimulate the cell cycle. If a mutation makes Ras or any other pathway component abnormally active, excessive cell division and cancer may result. P Ras P P P P P GTP 4 2 Receptor G protein Hyperactive Ras protein (product of oncogene) issues signals on its own Protein kinases (phosphorylation cascade) NUCLEUS 5 Transcription factor (activator) DNA Gene expression Protein that stimulates the cell cycle Figure 19.12a Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The p53 gene encodes a tumor-suppressor protein – That is a specific transcription factor that promotes the synthesis of cell cycle–inhibiting proteins (b) Cell cycle–inhibiting pathway. In this pathway, 1 DNA damage is an intracellular signal that is passed via 2 protein kinases and leads to activation of 3 p53. Activated p53 promotes transcription of the gene for a protein that inhibits the cell cycle. The resulting suppression of cell division ensures that the damaged DNA is not replicated. Mutations causing deficiencies in any pathway component can contribute to the development of cancer. 2 UV light Protein kinases 3 1 DNA damage in genome Active form of p53 DNA Protein that inhibits the cell cycle Figure 19.12b Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings MUTATION Defective or missing transcription factor, such as p53, cannot activate transcription • Mutations that knock out the p53 gene – Can lead to excessive cell growth and cancer (c) Effects of mutations. Increased cell division, possibly leading to cancer, can result if the cell cycle is overstimulated, as in (a), or not inhibited when it normally would be, as in (b). EFFECTS OF MUTATIONS Protein overexpressed Cell cycle overstimulated Figure 19.12c Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Protein absent Increased cell division Cell cycle not inhibited The Multistep Model of Cancer Development • Normal cells are converted to cancer cells – By the accumulation of multiple mutations affecting proto-oncogenes and tumorsuppressor genes Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • A multistep model for the development of colorectal cancer Colon Colon wall Normal colon epithelial cells 1 Loss of tumorsuppressor gene APC (or other) 4 Loss of tumor-suppressor gene p53 2 Activation of ras oncogene Small benign growth (polyp) 3 Loss of tumorsuppressor gene DCC Figure 19.13 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Larger benign growth (adenoma) 5 Additional mutations Malignant tumor (carcinoma) • Certain viruses – Promote cancer by integration of viral DNA into a cell’s genome Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Inherited Predisposition to Cancer • Individuals who inherit a mutant oncogene or tumor-suppressor allele – Have an increased risk of developing certain types of cancer Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Concept 19.4: Eukaryotic genomes can have many noncoding DNA sequences in addition to genes • The bulk of most eukaryotic genomes – Consists of noncoding DNA sequences, often described in the past as “junk DNA” • However, much evidence is accumulating – That noncoding DNA plays important roles in the cell Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings The Relationship Between Genomic Composition and Organismal Complexity • Compared with prokaryotic genomes, the genomes of eukaryotes – Generally are larger – Have longer genes – Contain a much greater amount of noncoding DNA both associated with genes and between genes Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Now that the complete sequence of the human genome is available – We know what makes up most of the 98.5% that does not code for proteins, rRNAs, or tRNAs Exons (regions of genes coding for protein, rRNA, tRNA) (1.5%) Repetitive DNA that includes transposable elements and related sequences (44%) Alu elements (10%) Figure 19.14 Simple sequence DNA (3%) Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Repetitive DNA unrelated to transposable elements (about 15%) Introns and regulatory sequences (24%) Unique noncoding DNA (15%) Large-segment duplications (5-6%) Transposable Elements and Related Sequences • The first evidence for wandering DNA segments – Came from geneticist Barbara McClintock’s breeding experiments with Indian corn Figure 19.15 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Movement of Transposons and Retrotransposons • Eukaryotic transposable elements are of two types – Transposons, which move within a genome by means of a DNA intermediate – Retrotransposons, which move by means of an RNA intermediate Transposon DNA of genome Transposon is copied New copy of transposon Insertion Mobile transposon (a) Transposon movement (“copy-and-paste” mechanism) Retrotransposon New copy of retrotransposon DNA of genome RNA Reverse transcriptase Figure 19.16a, b (b) Retrotransposon movement Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Insertion Sequences Related to Transposable Elements • Multiple copies of transposable elements and sequences related to them – Are scattered throughout the eukaryotic genome • In humans and other primates – A large portion of transposable element– related DNA consists of a family of similar sequences called Alu elements Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Other Repetitive DNA, Including Simple Sequence DNA • Simple sequence DNA – Contains many copies of tandemly repeated short sequences – Is common in centromeres and telomeres, where it probably plays structural roles in the chromosome Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Genes and Multigene Families • Most eukaryotic genes – Are present in one copy per haploid set of chromosomes • The rest of the genome – Occurs in multigene families, collections of identical or very similar genes Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • Some multigene families – Consist of identical DNA sequences, usually clustered tandemly, such as those that code for RNA products RNA transcripts DNA Non-transcribed spacer Transcription unit DNA 18S 5.8S 28S rRNA Figure 19.17a Part of the ribosomal RNA gene family 28S 18S Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings 5.8S • The classic examples of multigene families of nonidentical genes – Are two related families of genes that encode globins -Globin Heme Hemoglobin -Globin -Globin gene family -Globin gene family Chromosome 16 Chromosome 11 Figure 19.17b The human -globin and -globin gene families Embryo 1 2 1 2 Fetus and adult Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings G A Embryo Fetus Adult • Concept 19.5: Duplications, rearrangements, and mutations of DNA contribute to genome evolution • The basis of change at the genomic level is mutation – Which underlies much of genome evolution Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Duplication of Chromosome Sets • Accidents in cell division – Can lead to extra copies of all or part of a genome, which may then diverge if one set accumulates sequence changes Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Duplication and Divergence of DNA Segments • Unequal crossing over during prophase I of meiosis – Can result in one chromosome with a deletion and another with a duplication of a particular gene Transposable element Gene Nonsister chromatids Crossover Incorrect pairing of two homologues during meiosis and Figure 19.18 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Evolution of Genes with Related Functions: The Human Globin Genes • The genes encoding the various globin proteins – Evolved from one common ancestral globin gene, which duplicated and diverged Ancestral globin gene Duplication of ancestral gene Mutation in both copies Transposition to different chromosomes Further duplications and mutations Figure 19.19 2 2 1 1 -Globin gene family on chromosome 16 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings G A -Globin gene family on chromosome 11 • Subsequent duplications of these genes and random mutations – Gave rise to the present globin genes, all of which code for oxygen-binding proteins Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • The similarity in the amino acid sequences of the various globin proteins – Supports this model of gene duplication and mutation Table 19.1 Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Evolution of Genes with Novel Functions • The copies of some duplicated genes – Have diverged so much during evolutionary time that the functions of their encoded proteins are now substantially different Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Rearrangements of Parts of Genes: Exon Duplication and Exon Shuffling • A particular exon within a gene – Could be duplicated on one chromosome and deleted from the homologous chromosome Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings • In exon shuffling – Errors in meiotic recombination lead to the occasional mixing and matching of different exons either within a gene or between two nonallelic genes EGF EGF EGF Epidermal growth factor gene with multiple EGF exons (green) EGF Exon shuffling F F F Exon duplication F Fibronectin gene with multiple “finger” exons (orange) F EGF K K Plasminogen gene with a “kfingle” exon (blue) Figure 19.20 Portions of ancestral genes Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings Exon shuffling TPA gene as it exists today K How Transposable Elements Contribute to Genome Evolution • Movement of transposable elements or recombination between copies of the same element – Occasionally generates new sequence combinations that are beneficial to the organism • Some mechanisms – Can alter the functions of genes or their patterns of expression and regulation Copyright © 2005 Pearson Education, Inc. publishing as Benjamin Cummings