* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download BIOL 112 – Principles of Zoology
Population genetics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Holliday junction wikipedia , lookup
DNA vaccination wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
History of genetic engineering wikipedia , lookup
Epigenomics wikipedia , lookup
Molecular cloning wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Genetic code wikipedia , lookup
SNP genotyping wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Non-coding DNA wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Genome editing wikipedia , lookup
Oncogenomics wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Helitron (biology) wikipedia , lookup
DNA polymerase wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Microsatellite wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Microevolution wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
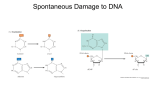
Mutations & DNA Repair I. II. What are mutations? Mutagenesis: Process of producing a mutation III. Repair of mutations Mutations can cause changes in the shape of a protein which alters its function. I. What are mutations? A. Classes of mutations: Spontaneous mutation - occurs in nature without the addition of a mutagen Induced mutation – caused by a mutagen Point mutation – change of 1 nucleotide Insertion/Deletion – base added or deleted Frameshift mutation – loss or addition of a nucleotide alters the codon reading frame Forward mutation – converts wild type to mutant Reverse mutation – converts mutant back to wild type Loss of function mutation Gain of function mutation Base substitutions/Point mutations Transitions Purine replaced by a purine… or pyrimidine replaced by a pyrimidine Transversions Purimidine replaced by a purine, or vise versa Less common… why? T&C A & G Additional mutational categories: Lethal mutation – results in death Conditional mutation – expression depends on the environment i.e. temperature sensitive mutations Somatic mutation – not transmitted to future generations Germinal/gametic mutation – transmitted to offspring Mutational outcomes 1) 2) 3) Silent substitution – function of the protein product of gene is unaltered Missense mutation – alters codon so that it encodes a different amino acid Nonsense mutation – alters gene so that it creates a nonsense codon (no normal tRNA exists) causing termination of translation Spontaneous mutations Spontaneous mutations arise from replication errors & base modifications DNA Replication errors Replication slippage – one strand loops out and becomes displaced during replication DNA pol stuttering Occurs frequently in repeat regions: Hot Spots for DNA mutation Spontaneous mutation rate various among organisms (table 15.2) II. Mutagenesis: Process of producing a mutation Ways Mutations can occur: -Replication Error -Breaks in DNA strands -Damage to nucleic acids -Mobile elements Induced Mutation Mechanisms 1) Base replacement 2) Base alteration 3) Base damage 1. Base replacement Base analogs (chemicals that are similar to nucleotides) substitute themselves for the nucleotide Result = improper base pairing Examples: a) Tautomeric Shifts - Tautomerization – isomerization of a nitrogen base to an alternative H-bonding condition b) Chemicals: 5-Bromocuracil (T analog), 2-Aminopurine (A analog) Tautomerization – Known as a tautomeric shift “rare” forms result in mispairing, . Mispairing results in replication errors – the wrong bases are incorporated into the daughter strands 5 BU (derivative of uracil) behaves as a thymine analog, if 5 BU is incorporated it will base pair with guanine, after 1 round of replication an A-T to G-C transition results 2. Base alteration Chemicals cause the shape of the nucleotide to change, resulting in improper base pairing Depurination (loss of nitrogenous base) & Deamination (amino group converted to keto group) Alkylation – addition of alkyl group (CH3 or CH3CH2) to bases • EMS (ethylmethane sulfonate) Intercalation – planer molecules that mimic base pairs and slip themselves between the stacked nitrogen bases at the core of the helix • • • Ethidium bromide Proflavin Acridine orange EMS alkylates the keto groups of G and T, base pairing is altered and a transition results Intercalating agents slip between the nitrogenous bases, which can lead to insertion/deletions. Frameshift mutations result – generated at gaps produced in DNA during replication 3. Base damage Chemicals, oxidation, radiation cause the nucleotide to become modified in such a way that it can no longer base pair UV light • Results in pyrimidine dimers Radiation • • • Causes ionization of molecules Creates substitutions Breaks phosphodiester bonds Spontaneous Mutations– arise due to natrual biological/chemical processes: DNA replication errors DNA replication errors = each of the bases can appear in one of several forms called tautomers (isomers) Spontaneous lesions Depurination - Apurinic sites can’t specify a base complementary to the original Deamination = ie deamination of C yields U, which will pair w/A leading to a GC to AT transition Oxidative damage – superoxide radicals (byproducts of metabolism) alter bases to cause mispairing… 8oxidG or GO pairs with A Transposable elements significant part of the genome consists of “nomadic” DNA sequences that are present at different locations III. Repair of mutations 1. 2. 3. 4. 5. 6. Direct Reversal of damage Excision repair Proofreading Mismatch repair Post-replication repair & SOS Double-strand break repair 1. Direct reversal of damage Photoreactivation repair: reversal of UV damage Photolyase splits Thymine dimers, restoring DNA to its original condition Photolyase works with cofactor folic acid The two bind together in dark to T-dimer When light shines on cell –folic acid absorbs the light & uses the energy to break the covalent bond between T’s O6-mGua DNA methyltransferase Alkyltransferase – one time repair enzyme that removes ethyl or methyl groups from guanine 2. Excision repair Involved in repair of deamination and depurination Enzymes recognize an abnormal base and cleave the bond between in and the sugar in the DNA backbone. 1) Uracil N-glycosylase 2) AP endonuclease 3) removes uracil cuts 5’ side of damaged site on apurinic bases Phosphodiesterase Removes sugar-phosphate residue deamination AGTGACTTAG TCACTGAATC AGTGACTTAG TCAUTGAATC Ligase Uracil N-clycosylase U AGTGACTTAG TCA TGAATC AGTGACTTAG TCA TGAATC Pol I 3. Proofreading DNA Pol III error rate: 10-5 Proofreading ability: Pol II can recognize mismatched base pairs, determine which base is the incorrect one excise the wrong base and carry out repair synthesis 3’ to 5’ exonuclease ability, lowering the error rate to 10-7 4. Mismatch repair Mismatch repair – after proofreading, mismatches identified, improper base excised and replaced w/correct base Adenine methylase recognizes parent strand and adds methyl group to A’s Unmethylated daughter strand recognized by repair enzyme Mismatch repairImportant to recognize difference between old strand & new strand: If mutated base excised, the wild type is restored, but if the original wild type is excised, the mutant sequence becomes fixed. 5. Post-replicational repair & SOS Post-replicational repair (aka recombination repair ): Damaged DNA cause Pol III to “stutter” and skip past damaged site Replication restarts downstream and a gap is left Gap is repaired by retrieving sequence from the normal copy and then the subsequent gap is repaired SOS response Severe damage due to alkylating agents or cross-linking agents (UV radiation best studied) triggers this response Translesional polymerases (POL II, IV, V) can replicate over damaged regions Has very high error rate: 10-2, however allows for survival under extremely bad conditions 6. Double strand break repair Repairs DSBs by reannealing the two DNA segments - protein aligns the broken ends of DNA for rejoining Recombination repair mechanism Homologous recombination repair – damaged DNA replaced by homologous DNA section from sister chromatid Nonhomologous recombination repair – uses non-homologous region for replacement Errors in direct joining may be a cause of translocations