* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download 슬라이드 1
Behavioral epigenetics wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Human genetic variation wikipedia , lookup
Neuronal ceroid lipofuscinosis wikipedia , lookup
Public health genomics wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Oncogenomics wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Copy-number variation wikipedia , lookup
Epigenetics of depression wikipedia , lookup
Metagenomics wikipedia , lookup
Pathogenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Primary transcript wikipedia , lookup
Point mutation wikipedia , lookup
Epigenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genomic library wikipedia , lookup
Transposable element wikipedia , lookup
Human genome wikipedia , lookup
Non-coding DNA wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Genetic engineering wikipedia , lookup
Gene expression programming wikipedia , lookup
Gene nomenclature wikipedia , lookup
Gene desert wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
History of genetic engineering wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Gene therapy wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Genome evolution wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Microevolution wikipedia , lookup
Genome editing wikipedia , lookup
Designer baby wikipedia , lookup
Helitron (biology) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
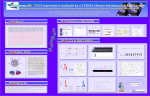
Human LTR promoter in NOS3 gene: Structure, Expression, Methylation and Evolution Hong-seok HA, Jae-Won HUH, Dae-Soo KIM1, Myung-JinJOO2 and Heui-Soo KIM * Division of Biological Sciences, College of Natural Sciences, Pusan National University. 1PBBRC, Interdisciplinary Research Program of Bioinformatics, College of Natural Sciences, Pusan National University. HTTP://WWW.PRIMATE.OR.KR 2Department of Psychiatry, Hyung Ju Hospital. INTRODUCTION ABSTRACT LTR: Long Terminal Repeat Endothelial nitric oxide synthase (NOS3) is a key enzyme in the regulation of vascular wall homeostasis and regulation of vasomotor tone, which has been identified to consist of 26 exons spanning 21 kb of genomic DNA and encoding an mRNA of 4052 nucleotides which is translated into a 1203 amino acids. Here we found new transcript variant that derived from LTR10A belonging to HERV-I family on human NOS3 gene. The LTR10A element located on the upstream of the original promoter elements (Sp1 and GATA motifs) of NOS3 gene seems to be inserted into primate genome approximately 33 Myr ago. The LTR10A-derived promoter transcripts by RTPCR amplification are detected in placenta tissue only. Methylation study using the sodium bisulfied DNA sequencing indicated that LTR10A element of placenta tissue showed hypomethylated pattern. Reporter gene assay of LTR10A element on NOS3 gene indicated good promoter activity of in human colon carcinoma cells (HCT116). These findings suggest that the LTR10A element acquired the role of placentaspecific regulation of NOS3 gene during primate evolution. . MATERIALS & METHODS Bioinformatics Luciferase assay Transfac 6.0 RT-PCR Bisulfite Sequencing PCR Genomic DNA PCR & Gene cloning LINE: Long Integrated nuclear element SINE: Short Integrated nuclear element SINE 13% Other region 16% Retroelement HERV: Human Endogenous Retro-Virus RNA intermediate LINE 20% - LTR element + LTR element Retroposon - env Gene-related Sequence 36% HERV element 8% Retrotransposon - RT DNA element 3% Pseudogene Coding sequence 1% + RT LTR SINE ORF1 ORF2 LTR Yeast Ty1/copia/truncated HERVs 3% LTR P LTR Human THE1 Poly(A) Human Alu Retroelements have been subjected to many amplification and transposition events resulting in a widespread distribution of complete or partial retroviral sequences throughout the human genome. The human genome comprises approximately 8% of the human endogenous retroviruses (HERVs) and other long terminal repeat (LTR)–like elements . Most HERVs seem to have entered the genome between 10 and 50 million years ago, and they comprise over 200 distinct groups and subgroups . Expression of retroelements can influence the outcome of infections in different ways that can be either beneficial or detrimental to the host. A function of the multiple copy families, scattered throughout the genome, has been reported regulatory functions on the gene expression of nearby located genes . A small minority of such sequences has acquired a role in regulating gene expression, and some of these may be related to differences between individuals, and to expression of disease. . + env Retrovirus LINE LTR P ORF1 ORF2 L1 Poly(A) gag pol env LTR Full-length HERVs/exogenous retrovirus REFERENCES TTTTT 1. J. Boeke, J. Stoye, Retrotransposons, endogenous retroviruses, and the evolution of retroelements, In Retroviruses (1997) 343–435. 2. J. Jurka, Repbase update: a database and an electronic journal of repetitive elements, Trends Genet. 16 (2000) 418-420. RESULTS & DISCUSSION Fig. 1. The genomic structure of NOS3 gene including LTR10A element. Exons were represented by solid box with the exon numbers. Arrows indicate the primer location. The LTR10A element was integrated into the NOS3 gene with the antisense orientation on human chromosome 7q36.1. Fig. 3. Luciferase reporter gene assay for LTR10A-derived promoter of NOS3 gene in transient transfected HCT116 and COS7 cells (A) and sequence analysis of LTR10A-derived promoter of NOS3 gene for the prediction of transcription factor binding sites (B). Relative activity of luciferase assay for pGL2-hNOS3-LTR10A in forward and reverse orientation or the pGL2 basic vector was indicated as schematic diagram. Results are expressed as ratios of the luciferase activity to that of the promoterless pGL2 reporter plasmid. The average and standard error of the mean are presented as error bars. Putative transcription binding sites, AREB6, MEF-2, NF-Y, AP-1, FOXO4, are indicated. The transcription start sites (GAAAC) are represented by an arrow. Fig. 5. Nucleotide sequence alignment of LTR10A-derived promoter region of NOS3 gene from human (NOS3HU), chimpanzee (NOS3-CH), gorilla (NOS3-GO), orangutan (NOS3-OR), gibbon (NOS3-GI), Japanese monkey (NOS3-JM), crab-eating monkey (NOS3-CR), and rhesus monkey (NOS3-RM). The dot represents the same nucleotides as the human NOS3-HU sequences. TSS indicates transcription start sites of NOS3 gene. TSD (AATTT) represents target sites duplication of the boundary of LTR10A element of NOS3 gene. Specific mutations of Old world monkeys are highlighted by the solid box. Binding sites for the transcription factors are also indicated. Fig. 4. PCR analysis for the presence of LTR10A-derived promoter region of NOS3 gene using the various primate genomic DNAs. Hominoids and Old World monkeys showed PCR products that were cloned and sequenced (see figure 5). Fig. 2. RT-PCR analysis of LTR10A derived transcript (A) and methylation analysis (B) from different human tissues. Methylation state of all cytosines in the CpG sequences was analyzed by the bisulfite-modified DNA sequencing method. Each nucleotide position is symbolized by a circle representing the results of seven clones analyzed. Black sectors indicate the percentage of methylated cytosine. Fig. 6. Luciferase reporter gene assay for LTR10A-derived promoter of NOS3 gene derived from the crab-eating monkey in transient transfected HCT116 and COS7 cells. Relative activity of luciferase assay for pGL2-mNOS3LTR10A in forward and reverse orientation or the pGL2 basic vector was indicated as schematic diagram. Results are expressed as ratios of the luciferase activity to that of the promoterless pGL2 reporter plasmid. The average and standard error of the mean are presented as error bars.