* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Protein Function
Polyclonal B cell response wikipedia , lookup
Expression vector wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Magnesium transporter wikipedia , lookup
Ligand binding assay wikipedia , lookup
Ultrasensitivity wikipedia , lookup
Enzyme inhibitor wikipedia , lookup
Gene expression wikipedia , lookup
Point mutation wikipedia , lookup
Clinical neurochemistry wikipedia , lookup
Amino acid synthesis wikipedia , lookup
Interactome wikipedia , lookup
Oxidative phosphorylation wikipedia , lookup
Lipid signaling wikipedia , lookup
Phosphorylation wikipedia , lookup
Biochemical cascade wikipedia , lookup
Photosynthetic reaction centre wikipedia , lookup
Protein structure prediction wikipedia , lookup
Biosynthesis wikipedia , lookup
Nuclear magnetic resonance spectroscopy of proteins wikipedia , lookup
Protein purification wikipedia , lookup
Evolution of metal ions in biological systems wikipedia , lookup
Paracrine signalling wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Western blot wikipedia , lookup
Metalloprotein wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Two-hybrid screening wikipedia , lookup
Signal transduction wikipedia , lookup
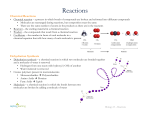
Protein Function Proteins and Ligands • Proteins work by interacting with other molecules. This means that each protein must bind to the specific molecules it interacts with. – Any molecule a protein binds to is called a ligand for that protein. Ligands can be small molecules or macromolecules, including other proteins. • The binding can be weak or strong: weak binding means only brief interactions, while strong binding means long term interactions, such as are found in large multi-subunit structures like ribosomes – The binding affinity of a protein for its ligand is measured by the dissociation constant, KD. Binding Sites • Ligands bind to specific regions of the proteins called binding sites. • Binding sites are usually cavities on the surface of a protein. • The R groups of various amino acids lining the cavity interact with the ligand by electrostatic forces, hydrogen bonds, and Van der Waals interactions. • • The amino acids involved in binding the ligand are very precisely placed, and they are very conserved in evolution. The other amino acids of the protein serve as a scaffold to help position the ligandbinding amino acids. They are often less conserved in evolution. Aligned Protein Sequences • These are the NTPase domains of 4 MutS proteins from the Plasmodium malaria parasite. • Note: internal gaps, ragged ends, highly conserved regions, semiconserved regions, regions with little homology. Antibody Binding Sites • Antibodies are part of the immune system. They are proteins that bind tightly to specific ligands. – Antibody ligands are called antigens. • The world produces vast numbers of possible antigens so the immune system must be capable of producing billions of different antibodies. – Disease organisms often disguise themselves by trying to have a non-antigenic surface. • The antigen binding sites are formed by the variable regions of both the heavy chain and the light chain. • Enzymes are proteins that catalyze chemical reactions. Each cell contains thousands of different enzymes. Each enzyme catalyzes a single reaction. • The ligands for enzymes are called substrates, and the enzyme converts a substrate molecule into a product molecule. • Enzymes are named for the reaction they catalyze: DNA polymerase polymerizes DNA from nucleotides, for example. • There are several general mechanisms by which enzymes work: – Putting two substrate molecules into the precise alignment needed for the reaction to occur – Stablizing a transition state between suubstrate and product – Providing a microenvironment that helps the reaction. For example, a highly acidic or a very reducing environment will encourage many reactions – Straining the conformation of a substrate to encourage it to transition to the product. Enzymes Example Enzyme: Lysozyme • • Lysozyme cuts the polysaccharide chains in the peptidoglycan coat of bacteria. This causes the bacterial cells to lyse due to osmotic pressure. Lysozyme performs a hydrolysis reaction: it adds water to a chemical bond to break it. – One product gets the –H from the water, and the other products gets the –OH. • The lysozyme reaction is energetically favorable: the free energy of the products is less than the free energy of the substrate. Thus, lysozyme just has to lower the activation energy of the reaction to get it to proceed. • • • • Lysozyme lowers the activation energy by straining the bond and by stabilizing the transition state (by allowing a temporary covalent bond between the sugar and the enzyme molecule). Also, in the microenvironment on the reaction site, note that glutamic acid is in the –COOH form and aspartic acid is in the –COO- form. This implies a pH of about 4.0, quite different from the pH of the cell, which is around 7.4. To add water to the bond, the polysaccharide needs to be in a distorted position, with its electron distribution altered. This is called the transition state. Getting into the transition state requires a lot of energy, which is difficult to achieve just from random collisions with water molecules. It is achieved by binding the substrate to the enzyme and holding it there during the reaction. – The lysozyme molecule has a long groove on its surface that holds 6 sugars in the peptidoglycan polysaccharide molecule. The enzyme breaks the bond between the 4th and 5th sugars. Lysozyme Lysozyme Action • At the active site, the C-O-C bonds between the sugars is strained. • A nearby glutamic acid (Glu35) in the –COOH form attacks the bond by donating its H+ to the connecting oxygen atom. (ES in the diagram) • This leaves the carbon from the upstream sugar (sugar D) with a strong partial + charge. (ES) • This carbon atom is then attacked by the –COO- from aspartic acid (Asp52), forming a (temporary) covalent bond between the sugar and the enzyme. (Transition State) • The combination of the –COO- on Glu35 and the unstable sugar-enzyme covalent bond splits a water molecule into H+ and OH-. The H+ goes to Glu35, regenerating its uncharged carboxylic acid state (-COOH). The OH- gets attached to the sugar carbon, leaving Glu35 in its original –COO- state. • The two halves of the polysaccharide are then released, and the lysozyme can catalyze another reaction. (EP) Coenzymes • Enzymes often use tightly bound small molecules, called coenzymes, to perform the chemical reaction. – Some coenzymes are metal ions: Zn, Ni, Co, Cu, etc. – Other coenzymes are small organic molecules, some of which are vitamins: thiamin, riboflavin, retinal, etc. Others, such as heme, are synthesized from other molecules. • Examples: – Retinal changes its conformation when struck by a photon. The photoreceptor protein rhodopsin is bound to retinal and detects the changed conformation – The iron atom in heme reversibly binds oxygen molecules – Biotin transfers –COOH groups by forming a transient covalent bond with the –COOH – The zinc (Zn) atom in carboxypeptidase forms a covalent bond with the peptide bond that carboxypeptidase cleaves. Some Co-enzymes Heme, with and without O2 bound Retinal: cis-trans change after absorbing a photon Biotin reversibly binds a –COOH group, then transfers it to another molecule. The Na-K Pump • Many proteins perform important non-enzymatic functions. One of these is membrane transport: for most small molecules, going across the membrane requires them to pass through a protein channel. (this is a big subject in BIOS 303) • Some molecules need to use ATP energy to actively pump them up the concentration gradient. The sodium-potassium pump (also called Na-K ATPase) is a very important case: it keeps the cell in a state of low sodium and high potassium inside the cell, relative to the outside environment. • Each cycle of the Na-K pump pumps 3 Na+ ions out and 2 K+ ions in. – An important consequence is that the cell has a lower internal potential than the outside: the interior is at about -70 millivolts. For most cells, the Na-K pump uses about 20% of all ATP. In neurons, it uses up to 2/3 of the cell’s energy. How the Pump Works • The pump forms a channel through the membrane. • The pump has 2 conformational states: open to the interior of the cell, or open to the exterior. • When it is open to the interior, 3 sodium ions bind to sites in the channel. – This just happens by diffusion—there is always lots of Na+ around. • The pump binds to an ATP molecule and hydrolyzes it. This phosphorylates an aspartic acid on the pump protein, and releases ADP. • Phosphorylation of Asp changes the conformation of the pump, so it is now open to the exterior. • The 3 Na+ ions diffuse away, and 2 K+ ions bind to the channel. • The binding of the K+ ions causes the phosphate group on the Asp to be released. • The unphosphorylated pump changes back to the original conformation: open to the interior. • The K+ ions diffuse away and the cycle starts again. Motor Proteins • Motor proteins can move in a directional manner. Used for things like movement of the cell, movement of chromosomes in cell division, muscle movement, etc. • The best studied motor protein system is myosin and actin in muscle cells. This system also moves materials from place to place within the cell, and causes cells to move in an amoeba-like fashion. • Actin is composed on 2 strands wrapped around each other. The strands are composed of many globular subunits. – In non-muscle cells, the actin filaments are called microfilaments, and they make up the cytoskeleton. – Actin is found in all eukaryotic cells, and its amino acid sequence is highly conserved Stained microfilaments in fibroblasts. More Myosin • Myosin is the actual motor protein: it uses ATP energy to pull itself along an actin fiber. – Many different forms in the cell – Composed of a head region that uses ATP, a flexible neck, and a long tail that interacts with other myosin molecules or with cargo materials. • Myosin in the muscle consists of 2 chains wrapped around each other, with 2 head groups. – Myosin filaments are composed of several myosin molecules wrapped up together. – In muscle cells, the thick filaments are composed of 2 groups of myosin molecules joined tail-to-tail. Myosin Action • The myosin head group has 2 conformations: low energy and high energy (cocked). • In the high energy position, ADP and Pi (the products of ATP hydrolysis) are bound to myosin, and the myosin head attaches to an actin molecule. (1 on the next slide) • The power stroke occurs when ADP and Pi are released: this causes myosin to change configuration, pulling actin along with it. (2) • Then, a new ATP binds to the myosin head, causing it to release the actin. (3) • ATP hydrolysis occurs, returning the myosin head to its high energy or cocked position. (4) Myosin Action Kinesin • Kinesin is a protein that transports materials down microtubule pathways, from the center of the cell to the periphery. • It uses ATP, and it seems to walk done the microtubule, pulling its load behind. • The exact mechanism isn’t clear, but seems to involve winding and unwinding the twisted stalks. • Great video: https://www.youtube.com/watch?v=y-uuk4Pr2i8 Control of Protein Activity • To keep its metabolism running efficiently, a cell needs to have the proper amount of each enzyme. This requires control over the amount of enzyme present and how active each enzyme is. • Control at different levels: – How much of a specific protein is made: transcription and translation – How rapidly the protein is degraded – Confining the protein to a specific compartment in the cell: e.g. lysosome, mitochondria, etc. – Regulation of protein activity by small molecules or covalent modification (our current subject) Allosteric Regulation • Allosteric regulation occurs when a small molecule affects an enzyme’s activity by binding to a site separate from the enzyme’s catalytic site and changes the enzymes conformation. – Many enzymes are controlled by small molecules that have different structures from their substrates or products. – The enzymes have 2 different binding sites: one for the substrate, and another for the inhibitor. – When the inhibitor binds to the enzyme, the enzyme changes its conformation, which makes it less active. Aspartate transcarbamoylase, first step in pyrimidine biosynthesis, is inhibited by binding cytosine triphosphate (CTP). The enzyme changes its conformation, so the active site is closed off. Feedback Inhibition • Feedback inhibition is a form of allosteric inhibition that occurs when the regulatory molecule is part of the same biochemical pathway. – The feedback inhibitor comes from a later step in the biochemical pathway. If too much inhibitor builds up, it slows the earlier steps down. – Complex biochemical pathways often have several different feedback inhibition steps. • Negative regulation: when an increase in the regulatory molecule causes a decrease in the enzyme’s activity. • Positive regulation: when an increase in the regulatory molecule causes an increase in the enzyme’s activity. Sample Biochemical Pathway • Glycolysis and the Krebs cycle, plus some of the pathways leading to other chemical compounds. • Each step is catalyzed by a specific enzyme. • The black words and abbreviations are compounds, while the blue abbreviations are the enzymes. Covalent Modification • • • • • Protein conformation can be affected by attaching various groups to the amino acid side chains. The most common example of covalent modification is phosphorylation, where a phosphate group is attached to the –OH group in serine, threonine, or tyrosine. Many complex events in the cell are regulated by phosphorylation: for example, signaling pathways that are activated by extracellular signals and end by increasing transcription of certain genes. Also, cell division. Phosphorylation is a reversible reaction: a protein kinase takes a phosphate group from ATP and attaches it to the protein, and a protein phosphatase removes the phosphate group (dephosphorylation. Depending on the system, phosphorylation can either increase or decrease enzyme activity. Cell Signalling Example • When a specifc growth factor (GF) binds to a receptor protein on the cell surface, the transcription factor eIF-4G is activated through a series of steps. • Many of the steps involve activating enzymes by phosphorylating them. Other Covalent Modifications • Other small molecules can be attached to specific amino acids on a protein. A few examples: – Adding the fatty acid palmitate to a cysteine causes anchors the protein to the membrane. – Adding the small polypeptide ubiquitin marks a protein for degradation. – Adding acetyl groups to lysines on histone proteins removes some of their positive charges and allows the chromatin to condense together. GTP-binding Proteins • • Another major form of protein regulation comes from the binding of guanosine triphosphate (GTP). Hydrolysis of GTP to GDP + Pi releases energy that is used to change the conformation of the protein and do useful work. Proteins that use GTP in this manner are called G-proteins. Steps: 1. 2. 3. 4. • GTP binds to a site on the protein, activating it. The protein slowly hydrolyzes the GTP to GDP + Pi. The protein becomes inactive. GDP is released. Protein still inactive. A new GTP binds to the protein. Very common in the first step of signal transduction: the receptor protein interacts with a GTP-binding protein. We will also see a lot of G-proteins in the processes of transcription and translation. GTP-binding Protein Example • • • • EF-Tu is a protein involved in translating messenger RNA into protein. Specifically, EF-Tu detects proper base pairing between the tRNA’s anticodon and the codon of the messenger RNA. If the tRNA’s anticodon matches the messenger RNA codon, EF-Tu hydrolyzes GTP to GDP and changes its conformation. This causes EF-Tu to release the tRNA and the tRNA to become fully bound to the ribosome. EF-Tu thus serves a proofreading function: it only releases the tRNA if there is a match between the tRNA and the messenger RNA codon. EF-Tu in blue, GTP in yellow, tRNA in red G-Protein Coupled Membrane Receptors • External signaling molecules are detected when they bind to a receptor protein embedded in the cell membrane. – The receptor protein changes its conformation, which is detected inside the cell. • Many receptors use a GTP-binding protein (Gprotein) to start the signal transduction process. – These G-protein coupled receptors all have 7 transmembrane alpha helices – There are about 800 different G-protein coupled receptors in the human genome. • These G-proteins have 3 subunits, alpha (α), beta (β), and gamma (γ). That is, the G-protein is a heterotrimer. The alpha and gamma subunits have covalently attached lipids that keep them bound to the inner surface of the membrane. External surface of a G-protein with its ligand (epinephrine) bound. G-Protein Activation • The alpha subunit interacts with GTP. • Normally, the G-protein is associated with the receptor protein, has GDP bound, and has all 3 subunits associated together. • When the membrane receptor binds to its ligand (in this case, the hormone epinephrine), GDP is released and GTP binds. • This leads to the dissociation of the alpha subunit (with GTP) from beta and gamma. G-Protein Effects • The activated alpha subunit (with bound GTP), activates the enzyme adenyl cyclase, which makes cyclic AMP (cAMP), a “second messenger” inside the cell that triggers the response of the cell to the hormone. – cAMP causes the activation of protein kinase A, followed by a whole cascade of events, including turning on some genes. • The alpha subunit slowly hydrolyzes GTP to GDP, which causes it to no longer stimulate adenyl cyclase and allows it to re-bind to the beta and gamma subunits • In some systems, the beta-gamma subunits, dissociated from alpha, activate further steps in the signaling pathway. Extra