* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download L05v04.stamped_doc
Genome (book) wikipedia , lookup
Frameshift mutation wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
SNP genotyping wikipedia , lookup
Genetic engineering wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Oncogenomics wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
DNA vaccination wikipedia , lookup
Epigenomics wikipedia , lookup
Primary transcript wikipedia , lookup
Neocentromere wikipedia , lookup
X-inactivation wikipedia , lookup
Molecular cloning wikipedia , lookup
Genomic library wikipedia , lookup
Non-coding DNA wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Holliday junction wikipedia , lookup
DNA polymerase wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Microevolution wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
History of genetic engineering wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
DNA supercoil wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Helitron (biology) wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Homologous recombination wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Point mutation wikipedia , lookup
Genome editing wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
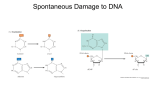
L05v04 [00:00:00.00] [00:00:02.18] PROFESSOR: Hi. In this video, we'll talk about three different ways that cells repair damaged DNA-- base excision repair, nucleotide excision repair, and double-strand break repair. [00:00:15.67] Method 1-- base excision repair. This occurs when a single base is affected. In the case shown here, it is a type of damage called deanimation. This is when a C base has an amino group that gets deaminated, leaving a carbonyl group in its place. And coincidentally, when this happens, the result is the base of cytosine turns into a base of uracil-- the base that is normally in RNA. [00:00:42.14] Now you will remember that uracil acts the same and base pairs the same as a T. And so this seems, to DNA, like a C-to-T mutation. But the cell also knows that there should be no uracil in DNA. [00:00:55.54] In fact, there are special enzymes called uracil DNA glycosylases that will scan the genome for places where a U is present. When it finds it, it will excise it, cut out the base, and then cut out the corresponding backbone groups-- a sugar and a phosphate group. Then DNA polymerase will come along and insert the correct base-- a C, to base pair with the G. And then the enzyme DNA ligase will seal the nick. [00:01:21.18] There's a very similar type of DNA damage called depurination, which happens to purines, which are Gs or As. This class of mutations cleave the bond between the sugar and the base, and the bases will be lost. So those types of mutations or DNA errors will use this repair mechanism as well. [00:01:40.59] These are both two common kinds of mutations, and they are repaired effectively by the cell, but not perfectly. Historically, we can actually see that, evolutionarily, there are more C-to-T mutations than other types of base-pair changes. So again, it's a very commonly occurring event. And the cell catches the vast majority, but not all of them. [00:02:02.92] A second type of DNA repair method is called nucleotide excision repair. In this case, two or more bases can be affected. And the type of damage shown here is a pyrimidine dimer, which is a more generic term for the type of damage we saw in the previous video, thymine dimers. [00:02:19.84] So this is a pyrimidine dimer. Instead of a T and T, a thymine dimer, we have a CT base pair, which can also form this dimer. And once again, there are proteins which constantly scan and surveil the genome. When it finds this kind of an error, it will cleave the backbone at some distance from the lesion site, remove the entire piece of DNA using the enzyme DNA helicase, and DNA polymerase will fill in the template, from three prime to five prime, and the ligase will seal the nick-- final nick. So again, this is an efficient mechanism for repairing thymine dimers. [00:02:55.26] Before we get to the third mechanism of repair, we're going to ask the question-when the cell encounters a mutation drawn mostly as a bubble, how does it know which strand to repair? This picture is not very realistic, because, really, the bubble is on both sides. This picture implies that the correct base is straight and has a normal backbone, and the incorrect base bubbles out a bit. But in reality, both sides will be affected. [00:03:23.15] In this case, you have a C-to-A mutation. After replication, a T will be inserted next to the A, and a C will properly be inserted next to the G. And going on, you would have 50% mutated cells and 50% nonmutated cells. [00:03:40.56] Now let's look at what happens if the cell knows somehow that the A is the incorrect base-- although we don't know how it does that, yet. It can correct the A and then it can put two G's there. And now you'll have 100% correct cells of the progeny, which will all have the correct genome. [00:03:59.66] But think about the consequences if the incorrect base is repaired-- if the G is cut out, and it gets converted to a T. Now you have 100% of the progeny cells having the mutation incorporated. [00:04:10.53] So how does the cell figure out which is the proper base to go at that position? The answer is quite ingenious. And what the cell does-- which gets it right most of the time, but not every time-- is it will, once it finds a mismatch, it will scan along the genome in both directions, looking for the closest nick in the backbone of the strand. [00:04:32.44] The cell then assumes that this is the most recently synthesized strand, the other strand, with no nicks, having stood the test of time, per se. And so it will decide to cut out the mutated region of the DNA that's on the strand that has the closest preexisting nick. And it will repair that with a polymerase and then a ligase sealing the final nick. [00:04:54.27] As you might guess, this all happens relatively quickly, right after DNA synthesis. It has to be quick because if a different enzyme happens to notice there's a nick present in the DNA it will repair the nick. Now the cell will not know which is the newly synthesized strand. So after synthesis, the DNA is basically a ticking time bomb, where the cell has to repair this bulge before another enzyme fixes this remote nick, if it's going to use this mechanism for pricking out which strand to repair. [00:05:26.99] Now the third DNA repair mechanism that we'll study is a mechanism called homologous recombination. And this is a way that cells repair, most, of the time, the most serious of DNA lesions-- double-strand breaks. When both strands are mutated or damaged, the cell needs to repair both strands, not just one. [00:05:46.34] One crude, brute-force method, shown here on the left, is nonhomologous enjoining. When a double-strand break is identified, if it cannot use homologous recombination, the cell will just cut back the ends of the DNA to make them blunt, or even, and then just jam the two ends together. You have deleted DNA base pairs, in this method, which, for a protein, could be quite disastrous. Losing even a single base will change the reading frame, which can then change every subsequent amino acid. And while that's better than having a broken chromosome, nonhomologous enjoining is a last resort. [00:06:28.52] A preferred method which we'll talk about is homologous recombination. And what's important to remember is that you have two copies of every chromosome. You have one chromosome from your father and one chromosome from your mother. [00:06:40.51] And what the cell will do is use the other chromosome to guide the repair. It will cut away the affected region. It will use, as a template, the other chromosome. And it will synthesize new DNA. So it will change your genetic material, replacing, for instance, your copy of your dad's gene with your mother's version. [00:06:59.69] So now you will have two copies of the maternal version. But hopefully it won't incorporate a deleterious mutation. And this is a far preferable alternative to simply jamming the two ends back together. [00:07:14.93] This is the molecular mechanism for homologous recombination. In this example, we have a double-strand break. First, the cell uses an exonuclease to chew back from the breaks in the five-prime direction. The purpose is to create a segment of the DNA which is singlestranded and then available for base pairing. [00:07:34.93] Miraculously, the single-stranded session aligns with the other parental chromosome and now is part of a double helix there. This is called a D-loop. And it can go on synthesizing in the normal five prime to three prime direction, where the bulge of the other strand of the paternal chromosome is being successively displaced, until it has traveled some distance. [00:07:56.38] Now it has enough material to reach the existing maternal chromosome. And then the ligase can seal the nick. At the same time, the existing part of the strand can now be used as a template for the synthesis of the other strand. And again, the ligase will seal the nick here. So overall, this is pretty amazing. [00:08:17.61] One of the ways that double-strand breaks occur is through ionizing radiation, like x-rays or cosmic rays. But a lot of us don't get exposed to x-rays that often. But a common way that double-strand breaks form is when a replication fork encounters a nick in the backbone. [00:08:35.32] If you have a leading-strand synthesis and you encounter a nick, that's it. You're done. There's no further extending, because you no longer have a complete strand that is in this proximity as a template. The cell will begin homologous recombination by using the exonuclease activity in the five-prime direction and having a strand that can base pair to the other chromosome and start the repair process. [00:09:01.63] We will encounter homologous recombination later in the course, when we're studying meiosis. As you already know, you inherit one chromosome from your father and one chromosome from your mother. But the chromosomes that you will pass on to your offspring are not purely either your paternal or your maternal chromosome, but a combination chromosome that is unique to you. [00:09:20.04] You might be wondering, how is it that, in some cases, homologous recombination can exchange genetic material, versus sometimes simply repairing a damaged segment? So we'll spend just a few minutes dissecting the mechanism. [00:09:35.91] The key to this is to focus on this particular intermediate called a Holliday junction. That's a rather famous term, and the difference is how these crossover intermediates get resolved. If we cleave the chiasma or cross over this way, you will have an exchange of genetic material. If you cleave this intermediate that way, you will have a repaired chromosome. [00:09:59.73] It's impossible to fully visualize here, but let me try to make it a little clearer. On the left is the form that was shown in the previous slide. On the right is the same molecule, but it's unwound in this direction. And now we have an open form. And I think you can see here, if you cleave it this way, that in one case you'll have repaired material, or in this case, if you cleave it here, you'll have translocation, with some of the paternal and maternal material mixed. [00:10:28.37] And this is made even clearer here, on this slide. Cut it vertically, and you have double-strand break repair. Cut it horizontally, and you have crossing over, as in meiosis. OK. This is a bit challenging to visualize, but work through it, and I think you'll be able to see the molecular difference. [00:10:44.76] So that concludes the video on three different types of DNA repair-- base excision repair, nucleotide excision repair, and homologous recombination. Thanks for listening.