* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download The replication of DNA
DNA paternity testing wikipedia , lookup
Genetic engineering wikipedia , lookup
DNA barcoding wikipedia , lookup
DNA methylation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
DNA sequencing wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Holliday junction wikipedia , lookup
Comparative genomic hybridization wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Genomic library wikipedia , lookup
DNA profiling wikipedia , lookup
Cancer epigenetics wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Microevolution wikipedia , lookup
Point mutation wikipedia , lookup
SNP genotyping wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
DNA nanotechnology wikipedia , lookup
Eukaryotic DNA replication wikipedia , lookup
DNA vaccination wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Primary transcript wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Microsatellite wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Non-coding DNA wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Epigenomics wikipedia , lookup
Molecular cloning wikipedia , lookup
History of genetic engineering wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
DNA replication wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
DNA supercoil wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
DNA polymerase wikipedia , lookup
Helitron (biology) wikipedia , lookup
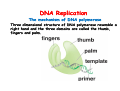
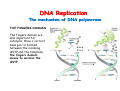
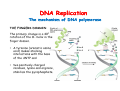
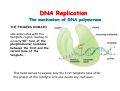
The replication of DNA Kornberg Meselson and Stahl Cairns Okazaki 1957 1958 1963 1968 DNA Replication The driving force for DNA synthesis. The addition of a nucleotide to a growing polynucleotide chain by a phosphodiester bonds release one molecule of pyrophosphate. The free energy release from this reaction is rather small. Additional free energy is provided by the rapid hydrolysis of the pyrophosphate into two phosphate groups by an enzyme know as pyrophosphatase. DNA synthesis reaction is essentially irreversible. DNA Replication The mechanism of DNA polymerase • DNA polymerase uses a single active site to catalyze the addition of any of the four dNTPs. • DNA polymerase monitors the ability of the incoming nucleotide to form an A:T or G:C base pair; incorrect base pairing leads to drammatically lower rates of nucleotide addition. DNA Replication The mechanism of DNA polymerase . • DNA polymerases show an impressive ability to distinguish between rNTPs and dNTPs. This discrimination is mediated by the steric exclusion of rNTPs from the active site; the nucleotide binding pocket is to small to allow the presence of a 2’-OH on the incoming nucleotide. This space is occupied by two “discriminator” amino acids that make van der Waals contact with the sugar ring. DNA Replication The mechanism of DNA polymerase Three dimensional structure of DNA polymerase resemble a right hand and the three domains are called the thumb, fingers and palm. DNA Replication The mechanism of DNA polymerase THE PALM DOMAIN The palm domain contain the primary elements of the catalytic site. This region binds two divalent metal ions (Mg2+ or Zn2+). DNA Replication The mechanism of DNA polymerase THE PALM DOMAIN One metal ion (A) reduce the affinity of the 3’-OH for its hydrogen. This generate a 3’O- ready for the nucleophilic attack of the α-phosphate of the incoming dNTPs. The second metal ion (B) coordinates the negative charges of the β- and γphosphate of the dNTP and stabilizes the pyrophosphate produced. DNA Replication The mechanism of DNA polymerase THE FINGERS DOMAIN The fingers domain are also important for catalysis. Once a correct base pair is formed between the incoming dNTP and the template, the fingers domain moves to enclose the dNTP. DNA Replication The mechanism of DNA polymerase THE FINGERS DOMAIN The primary change is a 40° rotation of the O- helix in the finger domain. • A tyrosine (aromatic amino acid) makes stacking interactions with the base of the dNTP and • two positively charged residues, lysine and arginine, stabilize the pyrophosphate. DNA Replication The mechanism of DNA polymerase THE FINGERS DOMAIN also associates with the template region, leading to a nearly 90° turn of the phosphodiester backbone between the first and the second base of the template. This bend serves to expose only the first template base after the primer at the catalytic site and avoids any confusion. DNA Replication The mechanism of DNA polymerase THE THUMB DOMAIN is not intimately involved in catalysis. It interacts with DNA that has been mostly recently synthesized. This serve two purpose: • It maintains the correct position of the primer and the active site • helps to maintain a strong association between the DNA polymerase and its substrate. It binds a primer-template junction. DNA Replication The mechanism of DNA polymerase THE PROOFREADING EXONUCLEASE The palm domain also monitors the base pairing of the most recently added nucleotide. If the recently added nucleotide are correctly base-paired, the palm domain makes extensive hydrogen bond contact with base pairs in the minor groove of the newly synthesized DNA. Mismatched DNA in this region interferes with the minor groove contacts and dramatically slows catalysis. DNA Replication The mechanism of DNA polymerase THE PROOFREADING EXONUCLEASE Proofreading of DNA synthesis is mediated by exonucleases present in the palm domain of the DNA polymerase. These exonucleases are capable of degrading DNA starting from a 3’ DNA end. When an incorrect nucleotide is incorporated, the rate of DNA synthesis is reduced because of the incorrect positioning of the 3’-OH DNA Replication The mechanism of DNA polymerase THE PROOFREADING EXONUCLEASE In the presence of a mismatch 3’-end, the last 3-4 nucleotide became single stranded. The exonuclease active site has a ten fold higher affinity for single stranded 3’ ends respect to the active site. Once bound, the mismatched nucleotide is removed and a properly base paired is reformed in the active site. Proofreading exonuclease activity reduces the error rate from 10-7 to 10-8 errors per base pair. DNA Replication The mechanism of DNA polymerase PROCESSIVITY The degree of processivity of DNA polymerases is defined as the average number of nucleotides added each time the enzyme binds a primer-template junction. • Each DNA polymerase has a characteristic processivity that can range from only a few nucleotide to more than 50.000 bases added per binding event. • Once bound addition of nucleotides is very fast. The fastest DNA polymerases are capable of adding as many as 1000 nucleotides per second to a primer strand. • Increased processivity is facilitated by the ability of DNA polymerases to slide along DNA template. Each time a nucleotide is added the DNA partially release from the polymerase (H-bond with the minor groove are broken, but the electrostatic interaction with the thumb are maintained). The DNA rapidly rebinds to the polymerase in a position that is shifted by 1 bp. DNA Replication Sliding DNA clamps Sliding DNA clamps are proteins composed of multiple identical subunits that encircle the DNA double helix and also bind tightly to DNA polymerases at replication forks. In absence of the sliding clamp, DNA polymerase dissociates from the template DNA on average once every 20-100 bp synthesized. DNA Replication Sliding DNA clamps In the presence of the sliding clamp, the DNA polymerase still disengages its active site from the 3’-OH frequently, but the association with the sliding clamps prevents polymerase from diffusing away from DNA. Thus, sliding clamp increase the processivity of the DNA polymerase. These proteins are present from bacteria to human. DNA Replication Sliding DNA clamps loaders Sliding clamp loaders are proteins that catalyze the opening and placement of sliding camp on DNA. These enzyme couple ATP binding and hydrolysis to the placement of sliding clamp around primer template junction, every time that this junction is present in the cell. The clamp loaders also remove the slide clamp from DNA once all of the enzymes that interact with them have completed their function. DNA Replication • In general, DNA is replicated by: – uncoiling of the helix – strand separation by breaking of the hydrogen bonds between the complementary strands – synthesis of two new strands by complementary base pairing Replication begins at a specific site in the DNA called the origin of replication (ori) DNA Replication Semidiscontinuous DNA replication Problem DNA polymerases can only add nucleotides from 5′→3′ but, the two strands of the double helix are antiparallel. Solution The major form of replication that occurs in nuclear DNA (eukaryotes), some viruses (e.g. the papovavirus SV40), and bacteria is called semidiscontinuous DNA replication.. replication DNA Replication Semidiscontinuous DNA replication • Two strands in the double helix separate at an origin of replication, exposing bases to form a replication bubble that contains two replication fork in opposite direction. • One strand is synthesized continuously from start to finish and the other strand is synthesized in short, discontinuous fragments. DNA Replication Semidiscontinuous DNA replication • Once primed, continuous replication is possible on the 3′→ 5′ template strand (leading strand). Leading strand synthesis occurs in the same direction as movement of the replication fork. DNA Replication Semidiscontinuous DNA replication • Discontinuous replication occurs on the 5′→3′ template strand (lagging strand). DNA is copied in short segments called Okazaki fragments moving in the opposite direction to the replication fork. The lagging strand requires the repetition of primer synthesis, elongation, primer removal with gap filling and joining of Okazaki fragments. DNA Replication Semidiscontinuous DNA replication • Despite this extra steps synthesis of both strands occurs concurrently • Nucleotides are added to the leading and lagging strands at the same time and rate by two DNA polymerases, one for each strand. • Fundamental features of DNA replication are conserved from E. coli to humans. DNA Replication in Bacteria • DNA replication is bidirectional from the origin of replication • DNA replication occurs in both directions from the origin of replication in the circular DNA found in most bacteria. DNA Replication in Bacteria Initiation of replication • An origin of replication is a site on chromosomal DNA where a bidirectional replication fork initiates or fires. • Most bacteria have a single, well-defined origin (e.g. oriC in E. coli) • Usually A-T rich. DNA Replication in Bacteria Initiation of replication • In E. coli the initiator protein DnaA binds to oriC and recruits a complex of two protein; the DNA helicase (DnaB) and helicase loader (DnaC). Three AA-T rich repeated elements that are the site of initial DNA unwinding. Five DnaA DnaA-binding sites The combination of all the protein that function at the replication fork is referred to as the REPLISOME DNA Replication in Bacteria DNA polymerases in bacteria E. coli has at least 5 DNA polymerases that are distinguished by their enzymatic properties, subunits composition and abundance. • DNA Pol III is the primary enzyme involved in the replication of the chromosome. One subunits called Klenow fragment has 5’-3’ polymerase activity; the other has a proofreading exonuclease activity 3’-5’ • DNA Pol I is specialized for the removal of the RNA primer and also has a proofreading exonuclease. • The remaining three DNA Pol are specialized for DNA repair and lack proofreading activities. DNA Replication in Bacteria DNA polymerases in bacteria In E. coli, the coordinate action of these polymerases is facilitated by physically linking them together in a large multiprotein complex called the DNA Pol III holoenzyme that confers very high processivity. The DNA pol III holoenzyme includes two copies of the core DNA Pol III and one copy of the five protein γ complex (the E. coli sliding camp loader). The γ complex includes two copies of the τ protein DNA Replication in Bacteria Initiation of replication • To begin DNA replication, unwinding enzymes called DNA helicases cause the two parent DNA strands to unwind and separate from one another at the origin of replication to form two "Y"-shaped replication forks. • These replication forks are the actual site of DNA copying DNA Replication in Bacteria DNA HELICASES DNA helicases, catalyze the separation of the two strand of the double helix breaking only the H-bonds that hold the two strands together. DNA helicases are hexameric proteins that assume the shape of a ring. DNA helicases use the energy coming from the ATP hydrolysis to encircle one of the two single strands at the replication fork. DNA Replication in Bacteria DNA HELICASES Each subunit has a hairpin protein loop that binds a phosphate of the DNA backbone and its two adjacent deoxyribose components. The coordinated movement of these protein hairpins can pull the ssDNA through the central pore of the helicase. DNA Replication in Bacteria DNA HELICASES There are specialized mechanisms that open the DNA helicase ring and place it around the DNA before forming the ring (initiation of replication). Each DNA helicases moves along ssDNA in a defined direction (POLARITY).This direction is define according to the strand of DNA bound. In case of a DNA helicases that functions on the lagging strand the polarity is 5’-> 3’; On the leading strand 3’-> 5’ . DNA Replication in Bacteria ssDNAssDNA -binding protein (SSB) • Helix destabilizing proteins bind to the single-stranded regions so the two strands do not rejoin Active E. coli SSB is composed of four identical 19 kDa subunits. DNA Replication in Bacteria ssDNAssDNA -binding protein (SSB) Single stranded binding proteins prevent premature annealing, to protect the single-stranded DNA from being digested by nucleases, and to remove secondary structure from the DNA to allow other enzymes to function effectively upon it. DNA Replication in Bacteria PRIMASE DNA primase is a type of RNA polymerase which creates a RNA primer Primase catalyzes the synthesis of a short RNA segment complementary to a ssDNA template. DNA polymerases can not initiate the synthesis of a DNA strand without an initial RNA or DNA primer (for temporary DNA elongation). DNA Replication in Bacteria PRIMASE In bacteria, primase binds to the DNA helicase forming a complex called the primosome. Primase is activated by DNA helicase where it then synthesizes a short RNA primer approximately 11 ±1 nucleotides long, to which new nucleotides can be added by DNA polymerase. DNA Replication in Bacteria • The τ proteins interact with DNA helicase and also bind to both DNA polymerase. • One DNA Pol III core replicates the leading strand and the other the lagging strand. • SSB coats the ssDNA regions. DNA Replication in Bacteria • Periodically DNA primase will associate with the DNA helicase and synthetize a new primer on the lagging strand template. • Lagging strand Pol complete the previous Okazaki fragment. DNA Replication in Bacteria When the lagging strand DNA Pol completes an Okazaki fragment, it is released from the sliding clamp and the DNA. DNA Replication in Bacteria The recently primed lagging strand is then a target of the clamp loader, which assembles a new sliding clamp at the primertemplate junction DNA Replication in Bacteria DNA Pol binds the sliding clamp and begin to synthesize a new Okazaki fragment This process continues around the circular genome until the replication forks meet each other at the terminus to generate two daughter molecules DNA Replication in Bacteria 1. Unwinding The enzyme helicase separates and unwinds the DNA double helix. The separated strands are held apart by single-strand binding proteins, forming a replication fork DNA Replication in Bacteria 2. Priming the Leading Strand Starting from a short RNA primer, DNA polymerase III assembles a new complementary strand on each parental (template) strand. The two strands are replicated in opposite directions, always 5' -> 3'. DNA Replication in Bacteria 3. Building the Leading Strand The strand that is polymerized toward the replication fork is called the leading strand. DNA polymerase III replicates the leading strand as a continuous strand moving into (5' -> 3') the replication fork. The RNA primer is removed and replaced with DNA by DNA polymerase I. DNA Replication in Bacteria 4. Priming and Building the Lagging Strand The lagging strand is synthesized in short pieces called Okazaki fragments, each with an RNAprimer. These fragments are joined together by DNA ligase, after DNA polymerase I has removed the RNA primers.