* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Clinical and Molecular Aspects of Diseases of Mitochondrial DNA
SNP genotyping wikipedia , lookup
DNA profiling wikipedia , lookup
Frameshift mutation wikipedia , lookup
Genetic engineering wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Genomic library wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Human genome wikipedia , lookup
Designer baby wikipedia , lookup
DNA polymerase wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
List of haplogroups of historic people wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
DNA barcoding wikipedia , lookup
Epigenomics wikipedia , lookup
Molecular cloning wikipedia , lookup
DNA vaccination wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Microevolution wikipedia , lookup
Primary transcript wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
DNA supercoil wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Non-coding DNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Helitron (biology) wikipedia , lookup
Point mutation wikipedia , lookup
Oncogenomics wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Mitochondrial Eve wikipedia , lookup
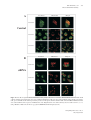
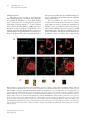
Special Edition 354 Clinical and Molecular Aspects of Diseases of Mitochondrial DNA Instability Chih-Chieh Mao, MD, PhD; Ian J. Holt1, PhD Mitochondria within human cells contain vast numbers of mitochondrial DNA (mtDNA), which are small, circular, and double-stranded. The proper functions of mtDNA depend totally on specific proteins that are encoded by the nucleus and then imported into mitochondria. Thus instability of mtDNA can stem from the mtDNA itself, or secondarily from abnormalities in nuclear DNA. In this review, we will first introduce mtDNA, including its characteristics, replication, transcription, translation, and the proteins involved in its metabolism, in particular DNA polymerase γ (POLG), DNA helicase Twinkle (Twinkle), and mitochondrial transcription factor A (TFAM). Secondly, we will stress the importance of mitochondrial nucleoid structures in the protection and facilitation of mtDNA metabolism, and report on the few known protein components of nucleoid, especially Twinkle, TFAM, and the recently discovered ATAD3. Based on this information, mtDNA instabilities will be categorized in accordance with their molecular etiologies, those that are caused by primary defects of mtDNA, and those by secondary effects from abnormalities in nuclear DNA. The former includes large defects or point mutations of mtDNA. The latter involves the nuclear genes of POLG1, Twinkle, ANT1, TK2, dGK, and TP. With the comprehensive categorization in this review, links are provided between the molecular and clinical aspects of mitochondrial DNA diseases. This report should help medical staff understand the complexity of these diseases and encourage them in further investigations. (Chang Gung Med J 2009;32:354-69) Key words: mitochondrial DNA, mitochondrial nucleoid, mitochondrial diseases 1. General features of mitochondria M itochondria are subcellular organelles within the cytoplasm of nearly all eukaryotic cells (a rare exception is the erythrocyte), which usually constitute ~25% of the volume of cytoplasm. They are not static entities. Instead they move along the cytoskeletal network,(1,2) undergoing frequent fusion and fission.(3-5) When cells divide, mitochondria are shared almost equally among the daughter cells. Mitochondria have an outer membrane and an inner membrane separated by a cavity, the intermembrane space.(6) The outer membrane is highly permeable, containing many pores for small molecules to pass through. In contrast, the inner mitochondrial membrane (IMM) is impermeable to most small molecules and ions, a property required for maintenance of an electrochemical gradient.(7) The IMM folds into lamellar structures called cristae,(6) which increase its surface area and mass. It is estimated that three quarters of the mass of the IMM is protein, and many of these proteins are components of the oxidative phosphorylation system (OXPHOS). The interior com- From the Department of Anesthesiology, Chang Gung Memorial Hospital, Taipei, Chang Gung University College of Medicine, Taoyuan, Taiwan; 1Medical Research Council Dunn Human Nutrition Unit, Cambridge, U.K. Received: Jun. 17, 2008; Accepted: Dec. 26, 2008 Correspondence to: Dr. Chih-Chieh Mao, Department of Anesthesiology, Chang Gung Memorial Hospital. 5, Fusing St., Gueishan Township, Taoyuan County 333, Taiwan (R.O.C.) Tel.: 886-3-3281200 ext. 2324; Fax: 886-3-3281200 ext. 2787; E-mail: [email protected] 355 Chih-Chieh Mao, et al Mitochondrial DNA instability partment of mitochondria is composed of a gel-like substance, the matrix, which contains enzymes of the tricarboxylic acid (TCA) cycle, and β-oxidation. These oxidation-reduction reactions and energy transfer processes make mitochondria the powerhouses of eukaryotic cells. The above chemical reactions (TCA cycle, βoxidation, and OXPHOS) involve a large number of proteins in the mitochondria. It is estimated that the mitochondrion contains approximately 1000 proteins, of which only 3-32, depending on the organism, are encoded by mtDNA.(8) The remainder are encoded by nuclear DNA, translated in the cytosol and imported into mitochondria via translocases of the outer membrane (TOM complex) and inner membrane (TIM22, TIM23 complexes). 2. Mitochondrial DNA Mitochondria, like chloroplasts, have their own DNA, mitochondrial DNA (mtDNA). However, mtDNA is itself dependent on nuclear genes for its replication, expression, and maintenance (see sections 3, 4 & 5 of this report). In mammals, mtDNA are covalently closed circular molecules of doublestranded DNA interspersed with a handful of ribonucleotides on each strand.(9) In a single cell the copy number of mtDNA is typically over 1,000. This huge number of mtDNA molecules is organized into discrete foci called mitochondrial nucleoids (mtnucleoids; see sections 5 and 6), which are believed to be tethered to the IMM. Mt-nucleoids comprise multiple copies of mtDNA and an unknown number of proteins. Nuclear DNA encodes all the proteins involved in mt-nucleoid metabolism. Most of the little that is known about mammalian mt-nucleoids has come from studies of human diseases caused by mtDNA perturbation (section 10). There is little variation in the size of mtDNA in animals; the range is approximately 16-19 kb. The genes are tightly packed; there are no introns, many genes are contiguous, and some even overlap. Mammalian mtDNA differs from nuclear DNA in several ways. It is small, circular, has a high copy number per cell, and is inherited through the maternal line. The two strands of mtDNA are separable on alkaline caesium chloride gradients, owing to a bias in the nucleotide composition. The so-called heavy strand (H-strand) is rich in guanines; while the light strand (L-strand) is rich in cytosines. The human Chang Gung Med J Vol. 32 No. 4 July-August 2009 mitochondrial genome is 16,569 base pairs (Fig. 1). Despite its small size, mtDNA encodes 22 transfer RNAs, 2 ribosomal RNAs and 13 polypeptides which are required for OXPHOS. Although the number 1000 is widely quoted as the mtDNA complement of a typical cell, there is in fact considerable variation in the mtDNA copy number among different cell types and tissues. The key determinant of the copy number is energy demand; thus, cardiac muscle has a considerably higher mtDNA copy number than skin cells. A high mtDNA copy number appears to protect against defects in mtDNA because OXPHOS can still be maintained without a significant cellular phenotype, until or unless a large proportion of mtDNA molecules is defective. Animal oocytes contain an extraordinary number of mtDNA molecules, about 100,000 in frogs and humans. In contrast to oocytes, spermatozoa contain few mitochondria, even less than a typical epithelial cell, although there is no doubt that their function is required for sperm motility. Mitochondria of sperm are located in the mid-piece, which is actively degraded upon entry into the egg.(10) Immediately following fertilization, mtDNA replication is quiescent, and is only activated at the 2-cell stage in mouse embryos.(11) The degradation of sperm mitochondria and the preponderance of oocyte mtDNAs ensure strict maternal inheritance of mtDNA. Human mtDNA is compact and economical, managing to compress 37 genes into fewer than 20 kilobases of DNA. There is, however, one substantial non-coding region (NCR), of ~900 bp in mammals (Fig. 1). Within the NCR, many molecules of mtDNA contain a triple-stranded region, or displacement loop (D-loop). (12-14) The third strand, or 7S DNA, is synthesised via a transcription event initiating at the light strand promoter (LSP), and then a transition to DNA synthesis at the origin of heavy strand replication (OH). In humans, the size of 7S DNA is about 650 bp, spanning nucleotide numbers 16,111 to nucleotide 191. The purpose of the D-loop is a matter of conjecture. 3. Mitochondrial DNA replication The mitochondrial DNA copy number is related to energy demand: tissues with the highest respiratory demand, such as muscle, liver, brain, and pancreatic islets, have the greatest number of copies of mtDNA. Therefore replication of mtDNA not only is Chih-Chieh Mao, et al Mitochondrial DNA instability HSP2 OH HSP F 12S rRNA 356 T NCR V Cyt b P LSP 16S rRNA E ND6 ND5 MELAS 3243G LUUR LHON 14484C LCUN ND1 LHON 3460A SAGY H 16,569 bp I LHON 11778A Q ND4 NARP T9176G/C M ND4L NARP 8993G/C QL ND2 N ND3 G COX3 MERFF 8344G A W R C Y ATPase 6 SUCN COX1 K ATPase 8 D COX2 5 kb deletion Complex I genes Complex III genes Transfer RNA genes Complex IV genes ATPase genes Ribosomal RNA genes Fig. 1 Schematic representation of 16,569 base-pair circular human mitochondrial DNA (mtDNA) showing genes encoding subunits of complex I, III, IV, and ATPase; ribosomal RNAs, and transfer RNAs according to the single letter code of the amino acids. The heavy strand promoter (HSP), light strand promoter (LSP), origin of heavy strand replication (OH), and a second heavy strand promoter (HSP2) are marked by bent arrows. The origin of lagging strand synthesis (OL) in a short spacing region is also marked by a bent arrow. Common pathogenic mutations (NARP, LHON, MERFF, MELAS) associated with mitochondrial disease are indicated by arrows. A large scale partial deletion (nucleotide position 8469 to 13447), seen in one third of patients with PEO/KSS/PS, is marked as a 5 kb deletion. Abbreviations used: NARP, neurogenic muscle weakness, ataxia, and retinitis pigmentosa syndrome; LHON, Leber’s hereditary optic neuropathy syndrome; MERFF, myoclonus epilepsy with ragged-red fibres syndrome; MELAS, mitochondrial encephalopathy, lactic acidosis, and stroke-like episodes syndrome; PEO/KSS/PS, progressive external ophthalmoplegia/Kearns-Sayre syndrome/Pearson’s syndrome. fundamental to cell biogenesis, but should also be regulated by the metabolic status of a given tissue at different stages of development or diseases. 3.1. Mechanisms of mtDNA replication There are two competing models of mtDNA replication in mammals, strand-asynchronous repli- cation, and coupled leading and lagging strand replication. In the strand-asynchronous model, the 2 strands of mtDNA initiate DNA synthesis in a temporally asynchronous way from 2 spatially separate points on the mtDNA, termed heavy (OH) and light (OL) strand origins. H strand replication is primed from a processed transcript initiated at the LSP with- Chang Gung Med J Vol. 32 No. 4 July-August 2009 357 Chih-Chieh Mao, et al Mitochondrial DNA instability in the NCR. The transcript is terminated at another region within the NCR known as the conserved sequences blocks (CSBs) and then processed, thereby creating a primer for DNA synthesis; the RNADNA transition point defines the origin of heavy strand replication (OH, typically nucleotide 191 of human mtDNA). H-strand DNA synthesis proceeds until it has traversed two-thirds of the genome, whereupon it exposes a short spacer region which is theoretically capable of forming a stem loop structure; separation of the two strands allows initiation of DNA synthesis of the second strand to commence.(15) This mechanism predicts extensive single-stranded regions in replicating molecules. Recently the model has been revised and alternative light strand ‘origins’ have been proposed.(16) Over the past few years an alternative view of mtDNA replication has been elaborated. Mitochondrial DNA molecules separated by twodimensional agarose gel electrophoresis have the appearance and properties of duplex replication intermediates. A fraction of these have all the hallmarks of products of conventional coupled leading and lagging strand synthesis.(17) However the majority are sensitive to RNase H(18) and this has led recently to the proposal that RNA is incorporated throughout the lagging strand at high frequency during mtDNA replication, the RITOLS mechanism.(19) ty, and initiation and elongation of DNA replication.(25) Mitochondria also contain a DNA helicase called Twinkle, which is related to the primase-helicase of T7 phage. (26) The helicase domain is conserved; however, the primase domain of T7 primasehelicase has no obvious similarity to Twinkle. Recombinant Twinkle has a DNA helicase activity with 5’ to 3’ directionality in vitro, and DNA unwinding by Twinkle is specifically stimulated by mtSSB.(27) In vitro, Twinkle, POLG, and mtSSB form processive replication machinery, which can use double-stranded DNA as a template to synthesize singlestranded DNA molecules of about 16 kb.(28) Mitochondrial DNA is supercoiled in vivo and its replication, as in other DNA systems, requires topoisomerases to relax and wind up DNA molecules. Mitochondrial topoisomerase 1 (TOP1MT) is homologous to nuclear topoisomerase 1, except that it has an N-terminal mitochondrial targeting signal in place of a nuclear localisation sequence.(29) TOP1MT releases the tension generated in the circular mitochondrial genome during replication and transcription. 3.2. Proteins involved in mtDNA replication Genetic information is contained in both strands of mammalian mtDNA (Fig. 1). Transcription initiates at two promoters in the NCR, one for each strand, hence heavy strand promoter (HSP), and light strand promoter (LSP). These give rise to polycistronic transcripts. H-strand transcription starts at nucleotide position 561 located within HSP, whereas L-strand transcription starts at nucleotide position 407 within LSP. The initiation of transcription is facilitated by mitochondrial transcription factor A (TFAM),(30) and mitochondrial transcription factor B (TFBM).(31) RNA synthesis in mitochondria is mediated by MTRPOL, in which the conserved C-domain specifies promoter selectivity and polymerase activity.(32) A second HSP is located immediately outside the NCR within the tRNAPhe gene. A ‘transcription termination factor’ (mTERF) simultaneously binds to this site and another position immediately downstream of the two rRNA genes, thereby forming a DNA loop, and then generating truncated H-strand The machinery of mtDNA replication in mammals involves a host of proteins. DNA polymerase γ, POLG, is believed to be the replicative polymerase for mtDNA. It forms a heterotrimer(20) composed of one molecule of POLG1, which is the catalytic subunit, and two molecules of POLG2, which are accessory subunits required for synthesizing long stretches of DNA in vitro.(21) The catalytic subunit, POLG1, has one conserved domain for polymerase activity located towards the C-terminus; at the other end is a domain for 3’ 5’ exonuclease ‘proofreading’ activity.(22) Mitochondrial single-stranded DNA-binding protein (mtSSB) has an N-terminal domain similar in sequence to prokaryotic SSB.(23) Like its counterparts elsewhere, it binds single-stranded DNA cooperatively as tetramers. (24) In flies and frogs, mtSSB enhances the functions of POLG in various assays of primer-template binding, 3’ 5’ exonuclease activi- Chang Gung Med J Vol. 32 No. 4 July-August 2009 4. Transcription, RNA processing, post-transcriptional modification, and translation 4.1. Transcription of human mtDNA and processing of primary transcripts Chih-Chieh Mao, et al Mitochondrial DNA instability transcripts containing the rRNAs with little else.(33) In the absence of mTERF, full-length H-strand transcripts are produced, which specify 10 mRNAs and 14 tRNAs. Since tRNA genes abut mRNAs in the polycistronic transcripts, cleavage of tRNAs by RNase P and RNase Z yields mature mRNAs. (34) Thus there is no requirement for a complex spliceosome in mitochondria. 4.2. Post-transcriptional modifications Post-transcriptional modifications of several bases in excised tRNAs are important for folding, aminoacylation, and codon recognition. Maturation of tRNAs requires addition of CCA to the 3’ end for the attachment of amino acids. The 3’ ends of mRNA undergo polyadenylation, which is believed to stabilize transcripts. 4.3. Translation Mitochondrial ribosomes are broadly prokaryotic-like. They are sensitive to aminoglycosides and chloramphenicol, and resistant to cyclohexamide(35) Mammalian mitochondrial ribosomes contain relatively little RNA, but have a high protein content of 29 and 48 polypeptides in the small and large ribosomal subunits, respectively. The result is a low sedimentation coefficient of 55S, compared with the 80S of nuclear ribosomes and 70S of bacterial ribosomes. Mitochondrial mRNAs have neither upstream untranslated regions to facilitate ribosomal binding, nor a cap structure at the 5’-end to direct ribosomes to the initiation codon. Instead, the small subunit of mitochondrial ribosomes binds mRNA in a sequence-independent way, and moves to the initiation codon. There the small subunit is bound by mitochondrial translation initiation factor 2 (mtIF2) and N-formylmethionyl-tRNA, facilitating recruitment of the large ribosomal subunit. Elongation of translation is carried out by mitochondrial elongation factors (mtEFs) that are homologous to EF-Tu, EFTs, and EF-G of E. coli. Likewise, termination of translation is believed to involve the recognition of stop codons by mitochondrial release factors (mtRFs) homologous to those in E. coli. 5. Nucleoids In vivo DNA is rarely, if ever, naked. Rather, it adopts an ordered condensed structure in association with a number of proteins. In bacteria, such nucleic 358 acid-protein complexes are termed nucleoids. (36) Packaging of DNA with protein offers a means of protection from extraneous damage, facilitating orchestrated copying and segregation of DNA molecules. Both are processes essential to the faithful transmission of genetic material. Mitochondrial DNA is no exception; it also forms nucleoprotein complexes, which have become the subject of intense study in recent years. (37) Mitochondrial nucleoprotein complexes, or mitochondrial nucleoids (mt-nucleoids), are believed to contain multi-copy mitochondrial genomes based on microscopic studies in which the number of mtDNA foci in a cell, indicated by a fluorescent dye specific to DNA, was significantly less than the mtDNA copy number. In humans, mt-nucleoids are about 0.068 µm in diameter as judged from immunocytochemical staining, and the mitochondrial genome per nucleoid is estimated to be between 2 and 10 copies.(38,39) In S cerevisiae, mt-nucleoids are larger than humans. In yeast growing aerobically, mt-nucleoids are 0.2 to 0.4 µm in diameter, and the mitochondrial genome per nucleoid is estimated to be 1 to 2 copies. Under anaerobic conditions, yeast mt-nucleoids grow to 0.6 to 0.9 µm and each contains the equivalent of ~20 copies of the mitochondrial genome.(37) Most progress in understanding mitochondrial nucleoids has been made in yeast. The concept of mitochondrial nucleoids is based on in vivo and ex vivo observations that yeast mtDNA forms stable complexes which are the units of segregation within the mitochondrial network. Yeast studies have led the field, largely because yeast genetics allows the rapid creation and screening of mutants. Caron et al.(40) first identified a histone-like protein from yeast mitochondria. This protein is encoded by the ABF2 gene. It has two high-mobility-group (HMG) domains capable of binding DNA and is required for maintenance of mtDNA.(41) The bacterial DNA packaging protein HU can restore mtDNA stability to abf2 mutant cells when targeted to mitochondria.(42) Thus Abf2 is likely to function, at least in part, as a DNApackaging protein. However, it is not clearly understood how Abf2 is involved in the maintenance of the mitochondrial genome because Yhm2, which is an inner membrane protein with in vitro DNA binding activity, and aconitase (Aco1), which is required for TCA cycle, can both complement abf2 mutant cells.(43,44) Chang Gung Med J Vol. 32 No. 4 July-August 2009 359 Chih-Chieh Mao, et al Mitochondrial DNA instability In organello formaldehyde cross-linking experiments identified a number of candidate mitochondrial nucleoid proteins of yeast;(45) subsequently several of these have been shown to associate closely with mtDNA and to contribute to mtDNA stability.(44,46,47) These include Abf2, Aco1, Rim1, Mgm101, Hsp60, and Ilv5. Rim1 is a single-stranded binding protein and by analogy with other systems is presumed to be involved in yeast mtDNA replication.(48) Mgm101 binds DNA and is required for the repair of damaged mtDNA. (49) Further investigation revealed that in vivo, Mgm101 colocalized with actively replicating mtDNA.(50) Even in the absence of mtDNA, Mgm101 forms a self-replicating complex with an outer membrane protein Mmm1, which itself is required for mtDNA stability.(51) Hsp60, a well known mitochondrial chaperone and analogue of E. coli GroEL, was found in yeast mitochondrial nucleoprotein preparations; when tested in vitro it was shown to bind to mtDNA ‘ori-like sequences’ preferentially. (45) Moreover, mtDNA is unstable in a hsp60 mutant and further studies showed that this instability was caused by a defect in mtDNA transmission to daughter cells,(47) suggesting the nucleoid dynamics underlying mtDNA transmission are regulated by the interaction between Hsp60 and mtDNA ori sequences. Ilv5 participates in branched-chain amino acid biosynthesis and plays a direct role in the parsing of mtDNA into nucleoids.(46) Ilv5 is essentially two proteins glued together, as the two functions are quite separable.(52) These findings provide convincing evidence of highly organized mitochondrial nucleoids whose maintenance and transmission is presumably dependent on mechanisms similar to bacteria. However, yeast mitochondria have few obvious direct homologues of bacterial DNA maintenance and segregation proteins.(36,37) Few protein components of yeast mitochondrial nucleoids have sequence-conserved homologues in mammals. In addition, targeted mutation of mammalian cells is cumbersome and laborious. Therefore less is known about mammalian mitochondrial nucleoids; they contain Twinkle, a DNA helicase, which is localized with mtDNA when overexpressed in cultured cells,(26) and TFAM,(53) which is believed to be the major packaging protein of mammalian mtDNA.(54) TFAM is a homologue of yeast Abf2, based on the conserved high-mobility-group (HMG) domains and its complementary effect in Chang Gung Med J Vol. 32 No. 4 July-August 2009 yeast cells devoid of Abf2. (55) Both Twinkle and TFAM are essential for mtDNA maintenance.(56,57) A fraction of POLG and mtSSB colocalize with Twinkle and mtDNA, as would be expected. (58) Additional proteins co-purify with frog oocyte mtDNA, (59) although the role of these, if any, in mtDNA metabolism is unknown. Studies of yeast mitochondrial nucleoid dynamics have been aided by live-staining with 4’, 6diamidino-2-phenylindole (DAPI). DAPI is a fluorescent dye that can pass through intact membranes to bind mtDNA and nuclear DNA in yeast. Unfortunately DAPI stains human mtDNA weakly. Recently PicoGreen, a fluorescent dye that binds specifically to double strand DNA in solution, has been used to stain mtDNA in living mammalian cells with good results.(60) 6. Recent progress in mammalian mitochondrial nucleoids A new procedure using the bacterial nucleoid protein HU to capture mitochondrial nucleoprotein from rat liver identified ATAD3, a member of the AAA+ family of ATPases.(61) Repressing expression of the homologue of ATAD3 in C. elegans (F54B3.3) caused embryonic lethality, larval arrest, slow growth, and maternal and progeny sterility, indicating it is an important mitochondrial protein involved in cell viability (http://www.wormbase.org/db/gene/gene?name=F54B3.3). Other members of the AAA+ family(62) include the bacterial nucleoid protein FtsK,(63-65) eukaryotic initiation protein Cdc6, and components of the origin recognition complex (Orcs 1, 4 and 5) required for the initiation of DNA replication.(66) ATAD3 was studied in further detail using RNA interference in human 143B cells.(61) Dramatic reductions in the PicoGreen signal of mt-nucleoids were revealed in ATAD3 siRNA-treated cells, suggesting that it plays a role in mtDNA metabolism (Fig. 2). In ATAD3 gene-silenced cells, DNA analysis by neutral/neutral two-dimensional agarose gel electrophoresis indicated that ATAD3 binds within the NCR of human mtDNA and contributes to the maintenance or formation of mtDNA multimers. (61,67) Hence, a coherent hypothesis is beginning to emerge. ATAD3 binds specifically to a region of mtDNA in the vicinity of the NCR, possibly the D-loop itself, and this brings together or stabilizes multiple Chih-Chieh Mao, et al Mitochondrial DNA instability PicoGreen A Mitotracker 360 Merged Zoom x 4 Control Zoom x 2 Zoom x 1 PicoGreen B Mitotracker Merged Zoom x 4 siRNA Zoom x 2 Zoom x 1 Fig. 2 The PicoGreen signal of mitochondrial nucleoids is greatly reduced after 2 rounds of ATAD3-siRNA. Panel A, human 143B cells mock transfected with nuclease free water (Ambion, Huntingdon, U.K.) were live-stained with PicoGreen (Invitrogen, Paisley, U.K.; green) and Mitotracker (Invitrogen, Paisley, U.K; red in the panel). Panel B, human 143B cells with siRNA ATAD3 treatment were stained with PicoGreen (green) and Mitotracker (red). Magnifications were with a 60x objective lens with zoom x 4, 2, or 1, using a Radiance 2000 confocal microscopy system (BIORAD, Hemel Hempstead, U.K.). Chang Gung Med J Vol. 32 No. 4 July-August 2009 361 Chih-Chieh Mao, et al Mitochondrial DNA instability mtDNA molecules. The obvious next step was to investigate the DNA binding properties of ATAD3. Two fragments of recombinant ATAD3 for in vitro DNA binding assays showed that one (residues 44-247) preferentially binds to D-loop structures,(61) even in solutions where double-stranded DNA is 1,000 times more abundant. Coupled with the 2D-AGE data, it is strongly suggested that ATAD3 binds to clusters of mitochondrial D-loops through the N-terminal onethird of the protein. The three-stranded stretches of D-loop that frequently turn up in mtDNA might provide a scaffold for a protein that helps mitochondria organize their DNA. The N-terminal one-third of the protein (residues 44-247) had also been used to raise an antibody which was used to examine the distribution of ATAD3 relative to mtDNA. Confocal microscopy indicated that ATAD3 is localized exclusively within mitochondria (Fig. 3A). However, ATAD3 is not distributed uniformly throughout mitochondria but is restricted to discrete foci (Fig. 3A), a characteristic A B 1 2 3 4 5 C Fig. 3 ATAD3 co-localized with mitochondria and mtDNA. Panel A, localization of mitochondria and ATAD3. Human 143B cells were live-stained with Mitotracker (red), then fixed, permeabilized, and immuno-labelled with a rabbit primary antibody against human ATAD3 (a kind gift from MRC Dunn Human Nutrition Unit, Cambridge, U.K.), and then with a goat anti-rabbit IgG secondary antibody pre-labelled with Oregon Green® 488 (Invitrogen, Paisley, U.K.; green). Panel B, localization of ATAD3 and DNA. Human 143B cells were fixed, permeabilized, and immuno-labelled for ATAD3 (as in panel A; green). In addition the cells were labelled with a mouse monoclonal anti-DNA IgM primary antibody (PROGEN Biotechnik, Heidelberg, Germany), and a goat antimouse secondary antibody Cy3 (Amersham Bioscience, Little Chalfont, U.K.; red). Panel C, enlargement of selected regions from the merged image of panel B. Cells were examined by confocal microscopy using an Olympus FluoView 1000 confocal microscopy system (Olympus, KeyMed House, U.K.), with magnifications from a 60x objective lens and zoom x 4 in panel A, and a 60x objective lens with zoom x 6 in panel B, in conjunction with the computer program Photoshop Elements (Adobe Systems Inc., Uxbridge, U.K.) in panel C. Chang Gung Med J Vol. 32 No. 4 July-August 2009 Chih-Chieh Mao, et al Mitochondrial DNA instability of mt-nucleoid.(26,61,67) Confocal microscopy further revealed frequent co-localization (yellow in Fig. 3C1, 3C-2, 3C-3) of ATAD3 (green in Fig. 3B, 3C) and mitochondrial nucleoids (red in Fig. 3B, 3C). Nevertheless a fraction of ATAD3 is in the form of ‘filamentous’ strings, flanked by mitochondrial nucleoids (Fig. 3C-4, 3C-5). Therefore, a model of mitochondrial nucleoid division has been proposed.(67) This model presumes that ATAD3 binding to mitochondrial D-loops enhances protein-protein interactions, thereby creating a filament, which drives apart the mitochondrial nucleoid. This also implies that mitochondrial nucleoids lacking ATAD3 are incapable of undergoing nucleoid division. Substantiating this model will require extensive investigation of cell biology and further in vitro analysis of ATAD3. 362 mtDNA deletions are invariably found in skeletal muscle but are often absent from blood. The clinical outcome is highly variable; the onset can range from birth to late adulthood (> 60 years). The reasons for this are not fully understood, but it relates in part to the mutant load in various tissues. The mildest clinical manifestation is progressive external ophthalmoplegia (PEO), which is characterised by droopy eyelids, and restricted eye movement, and may be accompanied by exercise intolerance. Much more severe is Kearns-Sayre syndrome (KSS), which involves cerebellar or cardiac dysfunction. Pearson’s syndrome (PS) is a fatal infantile disorder which is characterised by anemia and impaired synthesis of digestive enzymes in the pancreas. One third of PEO/KSS/PS patients have the so-called common deletion that removes 4977 bp of mtDNA (nucleotide position 8469 to 13447).(68,69) 7. Human mitochondrial diseases Mitochondrial diseases are a diverse group of human diseases characterised by defects in any aspect of mitochondrial function. Often for simplicity, defects relating to energy production, especially OXPHOS, are considered separately. Since the components of OXPHOS are encoded by mtDNA and nuclear DNA, and mtDNA is itself dependent on nuclear genes for its maintenance and expression, mitochondrial diseases of OXPHOS fall into three categories, mtDNA mutations as the primary cause, nuclear DNA defects affecting OXPHOS but unrelated to mtDNA, and nuclear DNA defects as a cause of mtDNA perturbation. 8. Mitochondrial DNA as the primary cause Mitochondrial DNA mutations can themselves be divided into two broad categories, those that affect translation, and those affecting a specific protein-encoding gene. The 22 tRNA and 2 rRNA genes are all required for mitochondrial translation; hence, deleterious mutations in any one will affect the production of all 13 mitochondrially-encoded components of OXPHOS. 8.1. Large-scale mtDNA deletions The occurrences of a variety of mtDNA deletions (∆-mtDNA, ~1 to 8 kb in size) are mostly sporadic, always heteroplasmic, and always result in the loss of one or more tRNA genes, therefore affecting translation of all 13 mitochondrial genes. Such 8.2. Point mutations affecting tRNA or rRNA The most common pathological mtDNA mutations are located in tRNA genes; somewhat unexpectedly they produce distinct clinical syndromes. MELAS syndrome (mitochondrial encephalopathy, lactic acidosis, and stroke-like episodes) usually presents before the age of 40, after normal early development. The majority of MELAS patients have an A to G transition at nucleotide position 3243 (A3243G) within the tRNALeuUUR gene,(70) which intriguingly is within the mTERF binding site (section 4.1). Another point mutation, A8344G, within the tRNALys gene, is associated with MERRF syndrome (myoclonus epilepsy and ragged-red fibres). (71,72) Ragged-red fibres (RRF) are a key indicator of mitochondrial dysfunction in skeletal muscle; high concentrations of mitochondria stain reddish-purple with the Gomori Trichrome stain, hence ragged red fibres mark areas of mitochondrial proliferation. In situ hybridisation studies indicate that the mitochondria in ragged red regions contain high levels of mutant mtDNA. (73) Hence, proliferation appears to be an attempt to boost mitochondrial output, although this is ultimately futile in the case of mutant mtDNA, as RRF are almost invariably cytochrome c oxidase negative. Both the MELAS and MERFF mutant tRNAs are non-functional, as they lack the usual post-transcriptional modification of uridine at the first position of the anticodon.(74) The best-known rRNA mutation occurs at Chang Gung Med J Vol. 32 No. 4 July-August 2009 363 Chih-Chieh Mao, et al Mitochondrial DNA instability nucleotide position 1555 in the 12S rRNA.(75) This A1555G mutation is associated with aminoglycoside antibiotic-induced deafness, and maternally inherited non-syndromic deafness. It has been demonstrated that the new G-C pair, created by the mutation, facilitates the binding of aminoglycoside antibiotics to 12S rRNA,(76) which explains the molecular pathophysiology of aminoglycoside antibiotic-induced deafness. 8.3. Point mutations affecting protein-coding genes Point mutations T8993G/C and T9176G/C in human mtDNA convert a conserved leucine to an arginine or proline, near the C-terminus of subunit 6 of ATP synthase.(77) Disease severity is directly related to the percentage of mutant mtDNA in the blood. When the percentage of T8993G mtDNA is less than 70%, patients are asymptomatic or oligosymptomatic and when it exceeds the threshold of 70%, NARP (neurogenic muscle weakness, ataxia, and retinitis pigmentosa) develops. Point mutations of mitochondrial genes encoding subunits of complex I are associated with LHON (Leber ’s hereditary optic neuropathy) syndrome, a form of adult onset blindness. The pathogenic mutations are G11778A in ND4, G3460A in ND1, and T14884C in ND6. Primary mtDNA mutations can be sporadic, particularly in the case of large deletions. Where mutations are transmitted, they are invariably through the maternal line. Genetic counselling in the case of mtDNA mutations is a primitive art as penetrance and clinical phenotypes are so variable. However in the case of affected fathers, (almost) complete reassurance can be offered; they will not transmit the disease to their offspring. 9. Nuclear DNA defects affecting OXPHOS rather than mtDNA Most of the 1000 or more mitochondrial proteins encoded in nuclear DNA do not directly affect mtDNA stability, but relate to mitochondrial metabolism and OXPHOS.(78,79) However, since OXPHOS is formed through the complementation of nuclear and mitochondrial DNA, mutations in nuclear genes encoding either structural components of OXPHOS,(80,81) or factors involved in the assembly of OXPHOS,(82) can all lead to clinical phenotypes similar to mtDNA mutations. Chang Gung Med J Vol. 32 No. 4 July-August 2009 10.Nuclear DNA defects as a cause of mtDNA perturbations Nuclear genes encode all the mtDNA binding proteins. Some are essential for replication and others are presumed to play roles in maintenance and segregation. As mitochondria are the major site of free radical production, protection from oxidative stress is another key function of the mtDNA binding proteins. Mutations in these nuclear genes can therefore produce mutations in mtDNA, but unlike primary mtDNA mutations, these are heterogeneous (multiple deletions and/or point mutations). Predictably, the clinical phenotypes associated with nuclear gene defects of proteins involved in mtDNA metabolism are often similar to those linked to specific mtDNA mutations, but the transmission follows Mendelian rules rather than a maternal pattern of inheritance. 10.1. POLG1, Twinkle, and Adenine Nucleotide Translocator 1 (ANT1) Autosomal dominant progressive external ophthalmoplegia (adPEO) is characterised by multiple large deletions of mtDNA in not only the ocular and skeletal muscles, but also the brain and heart.(83-85) adPEO is genetically heterogeneous with disease loci mapped to chromosome 15q25, 10q24, and 4q34-35. Chromosome 15-linked adPEO is caused by mutations in the POLG1 gene.(86) In fact, mutations of POLG1 are the most frequent cause of adPEO, accounting for approximately half of family cohorts. (86,87) Amino acid position 955 of POLG1, which is located within the catalytic polymerase domain, or C-domain, is the most frequent mutation site. (87) Mutations within the 3’ 5’ exonuclease domain, or N-domain, are associated with a recessively inherited form of PEO. Mutations of POLG1 are also associated with Parkinsonism.(88) Chromosome 10-linked adPEO is caused by mutations in Twinkle, a DNA helicase, which colocalizes with mtDNA(26) and is a component of mitochondrial nucleoids.(57) Mutations in Twinkle include various missense mutations and an in-frame duplication of 13 amino acids (dup352-364).(26,89) A recent study showed that transgenic mice expressing mutant Twinkle with in-frame dup352-364 develop multiple mtDNA deletions and a late-onset mitochondrial disease.(90) Chromosome 4-linked adPEO is caused by Chih-Chieh Mao, et al Mitochondrial DNA instability mutations in adenine nucleotide translocator 1 (ANT1).(91) The familial mutation substitutes a proline for a highly conserved alanine at position 114 in the ANT1 protein. ANT1 is a heart- and muscle-specific form of the ADP/ATP transporter, which is located in the mitochondrial inner membrane and is responsible for transporting ADP in and ATP out of mitochondria. It is likely that the mutation disturbs mitochondrial nucleotide pools, although it is not clear why this should cause multiple mtDNA deletions rather than depletion (as in the next section). 10.2. Thymidine Kinase 2 (TK2) and Deoxyguanosine Kinase (dGK) Mutations in the genes encoding thymidine kinase 2 (TK2) (92) and deoxyguanosine kinase (dGK)(93) are associated with mtDNA depletion syndrome (MDS), a severe infantile disease inherited in an autosomal recessive pattern. Patients develop tissue-specific depletion of mtDNA, with its level in the liver dropping to less than 10% of normal.(94,95) In mitochondria, both TK2 and dGK are required for the synthesis of deoxyribonucleoside triphosphates (dNTPs), the building blocks of DNA. In differentiated neurons and muscle cells, nuclear DNA replication is suspended and the cytosolic dNTP pool is lowered accordingly. There is no known de novo synthesis of dNTP in mitochondria, and so deoxyribonucleosides have to be recycled to replenish the mitochondrial dNTP pool. The salvage pathway for pyrimidines initially generates dTMP and dCMP by the action of TK2, whereas dGK generates dAMP and dGMP for purine biosynthesis. There is evidence showing that dNTPs need to be in balance for optimal DNA synthesis;(96) an excess or deficiency in any one dNTP can result in error-prone or complete cessation of DNA synthesis.(97,98) 10.3. Thymidine Phosphorylase (TP) Mutations in the gene encoding thymidine phosphorylase (TP) (99) are associated with depletion and/or multiple deletions of mtDNA in the muscles of patients with MNGIE (Mitochondrial neurogastrointestinal encephalomyopathy), an autosomal recessive disease. Intriguingly TP is not located in mitochondria, but in the cytosol where it is involved in the salvage of thymidine by catalysing the conversion of thymidine to phosphoribose and thymine. Mutations of TP lead to a decrease in enzyme activi- 364 ty to 5% of that in controls,(99) which results in accumulation of thymidine and an imbalance of dNTP pools.(100) 10.4. TFAM TFAM is a component of mitochondrial nucleoids.(54,55) Although TFAM has no known genetic relationship with any mitochondrial disease, in mice TFAM knockout causes severe mtDNA depletion and death in the fetus.(56) Tissue-specific disruption of TFAM causes dilated cardiomyopathy, anaesthesia-related conduction block in the heart,(101) and impaired insulin secretion in pancreatic-β-cells.(102) 11. The importance of dissecting the mitochondrial nucleoid in the pathogenesis of mitochondrial DNA instability Since there are more than 1000 copies of mtDNA in each cell, the percentage of mutant mtDNA needs to exceed a threshold to cause cell dysfunction. The threshold varies according to energy requirements, with the brain, heart, and skeletal muscle being the most susceptible to mitochondrial dysfunction. Crossing the threshold may result from random genetic drift(103) or biased segregation.(104,105) From studies on cultured cells it is known that different nuclear backgrounds confer selective advantages to particular mitochondrial genotypes. (105,106) Identifying and characterizing all the nucleus-encoded proteins that are functionally associated with mitochondrial nucleoids will help elucidate the production or biased segregation of mutant mtDNA. These studies are relevant to patients as the same mechanisms are likely to be key determinants of mutant load in particular individuals and tissues. 12. Summary This review reports the basic facts about mitochondria and mtDNA, and introduces nucleus-encoded proteins that are involved in the metabolism of mtDNA, with special emphasis on POLG, Twinkle, and TFAM. Integrated knowledge about the protein components of the genetic units of mtDNA, namely mitochondrial nucleoids, is also included. Among them Twinkle, TFAM, and the recently identified ATAD3 are detailed. Based on the molecular causes of mtDNA instability, mitochondria DNA diseases are grouped into two categories, mtDNA as the primary cause, and nuclear DNA defects as a cause of Chang Gung Med J Vol. 32 No. 4 July-August 2009 365 Chih-Chieh Mao, et al Mitochondrial DNA instability mtDNA perturbations. However, key questions remain to be answered, in particular the production and biased segregation of mtDNA instability in specific tissues. These fields may be greatly advanced by further identification and investigation of the individual components of mitochondrial nucleoids. Acknowledgements We acknowledge Chang Gung Memorial Hospital (Taoyuan, Taiwan), especially Professors Ping-Wing Lui, Min-Wen Yang, and Angie C-Y Ho of the Department of Anesthesiology, for their encouragement and support. Chih-Chieh Mao is supported by CMRP 1331 from the Chang Gung Memorial Hospital. Ian J. Holt is supported by the UK Medical Research Council and the European Union FP6 Mitocombat Integrated Programme. REFERENCES 1. Morris RL, Hollenbeck PJ. Axonal transport of mitochondria along microtubules and F-actin in living vertebrate neurons. J Cell Biol 1995;131:1315-26. 2. Nangaku M, Sato-Yoshitake R, Okada Y, Noda Y, Takemura R, Yamazaki H, Hirokawa N. KIF1B, a novel microtubule plus end-directed monomeric motor protein for transport of mitochondria. Cell 1994;79:1209-20. 3. Okamoto K, Shaw JM. Mitochondrial morphology and dynamics in yeast and multicellular eukaryotes. Annu Rev Genet 2005;39:503-36. 4. Santel A, Fuller MT. Control of mitochondrial morphology by a human mitofusin. J Cell Sci 2001;114:867-74. 5. Smirnova E, Griparic L, Shurland DL, van der Bliek AM. Dynamin-related protein Drp1 is required for mitochondrial division in mammalian cells. Mol Biol Cell 2001;12:2245-56. 6. Frey TG, Mannella CA. The internal structure of mitochondria. Trends Biochem Sci 2000;25:319-24. 7. DiMauro S, Schon EA. Mitochondrial respiratory-chain diseases. N Engl J Med 2003;348:2656-68. 8. Gray MW, Burger G, Lang BF. Mitochondrial evolution. Science 1999;283:1476-81. 9. Grossman LI, Watson R, Vinograd J. The presence of ribonucleotides in mature closed-circular mitochondrial DNA. Proc Natl Acad Sci USA 1973;70:3339-43. 10. Sutovsky P, Moreno RD, Ramalho-Santos J, Dominko T, Simerly C, Schatten G. Ubiquitin tag for sperm mitochondria. Nature 1999;402:371-2. 11. Sutovsky P, Moreno RD, Ramalho-Santos J, Dominko T, Simerly C, Schatten G. Ubiquitinated sperm mitochondria, selective proteolysis, and the regulation of mitochondrial inheritance in mammalian embryos. Biol Reprod 2000;63:582-90. Chang Gung Med J Vol. 32 No. 4 July-August 2009 12. Arnberg A, van Bruggen EF, Borst P. The presence of DNA molecules with a displacement loop in standard mitochondrial DNA preparations. Biochim Biophys Acta 1971;246:353-7. 13. Kasamatsu H, Robberson DL, Vinograd J. A novel closed-circular mitochondrial DNA with properties of a replicating intermediate. Proc Natl Acad Sci USA 1971;68:2252-7. 14. Walberg MW, Clayton DA. Sequence and properties of the human KB cell and mouse L cell D-loop regions of mitochondrial DNA. Nucleic Acids Res 1981;9:5411-21. 15. Clayton DA. Replication of animal mitochondrial DNA. Cell 1982;28:693-705. 16. Brown TA, Cecconi C, Tkachuk AN, Bustamante C, Clayton DA. Replication of mitochondrial DNA occurs by strand displacement with alternative light-strand origins, not via a strand-coupled mechanism. Genes Dev 2005;19:2466-76. 17. Holt IJ, Lorimer HE, Jacobs HT. Coupled leading- and lagging-strand synthesis of mammalian mitochondrial DNA. Cell 2000;100:515-24. 18. Yang MY, Bowmaker M, Reyes A, Vergani L, Angeli P, Gringeri E, Jacobs HT, Holt IJ. Biased incorporation of ribonucleotides on the mitochondrial L-strand accounts for apparent strand-asymmetric DNA replication. Cell 2002;111:495-505. 19. Yasukawa T, Reyes A, Cluett TJ, Yang MY, Bowmaker M, Jacobs HT, Holt IJ. Replication of vertebrate mitochondrial DNA entails transient ribonucleotide incorporation throughout the lagging strand. EMBO J 2006;25:5358-71. 20. Yakubovskaya E, Chen Z, Carrodeguas JA, Kisker C, Bogenhagen DF. Functional human mitochondrial DNA polymerase gamma forms a heterotrimer. J Biol Chem 2006;281:374-82. 21. Carrodeguas JA, Kobayashi R, Lim SE, Copeland WC, Bogenhagen DF. The accessory subunit of Xenopus laevis mitochondrial DNA polymerase gamma increases processivity of the catalytic subunit of human DNA polymerase gamma and is related to class II aminoacyl-tRNA synthetases. Mol Cell Biol 1999;19:4039-46. 22. Ye F, Carrodeguas JA, Bogenhagen DF. The gamma subfamily of DNA polymerases: cloning of a developmentally regulated cDNA encoding Xenopus laevis mitochondrial DNA polymerase gamma. Nucleic Acids Res 1996;24:1481-8. 23. Tiranti V, Rocchi M, DiDonato S, Zeviani M. Cloning of human and rat cDNAs encoding the mitochondrial singlestranded DNA-binding protein (SSB). Gene 1993;126:219-25. 24. Li K, Williams RS. Tetramerization and single-stranded DNA binding properties of native and mutated forms of murine mitochondrial single-stranded DNA-binding proteins. J Biol Chem 1997;272:8686-94. 25. Williams AJ, Kaguni LS. Stimulation of Drosophila mitochondrial DNA polymerase by single-stranded DNA- Chih-Chieh Mao, et al Mitochondrial DNA instability binding protein. J Biol Chem 1995;270:860-5. 26. Spelbrink JN, Li FY, Tiranti V, Nikali K, Yuan QP, Tariq M, Wanrooij S, Garrido N, Comi G, Morandi L, Santoro L, Toscano A, Fabrizi GM, Somer H, Croxen R, Beeson D, Poulton J, Suomalainen A, Jacobs HT, Zeviani M, Larsson C. Human mitochondrial DNA deletions associated with mutations in the gene encoding Twinkle, a phage T7 gene 4-like protein localized in mitochondria. Nat Genet 2001;28:223-31. 27. Korhonen JA, Gaspari M, Falkenberg M. TWINKLE Has 5’ 3’ DNA helicase activity and is specifically stimulated by mitochondrial single-stranded DNA-binding protein. J Biol Chem 2003;278:48627-32. 28. Korhonen JA, Pham XH, Pellegrini M, Falkenberg M. Reconstitution of a minimal mtDNA replisome in vitro. EMBO J 2004;23:2423-9. 29. Zhang H, Barcelo JM, Lee B, Kohlhagen G, Zimonjic DB, Popescu NC, Pommier Y. Human mitochondrial topoisomerase I. Proc Natl Acad Sci USA 2001;98:10608-13. 30. Fisher RP, Lisowsky T, Parisi MA, Clayton DA. DNA wrapping and bending by a mitochondrial high mobility group-like transcriptional activator protein. J Biol Chem 1992;267:3358-67. 31. Falkenberg M, Gaspari M, Rantanen A, Trifunovic A, Larsson NG, Gustafsson CM. Mitochondrial transcription factors B1 and B2 activate transcription of human mtDNA. Nat Genet 2002;31:289-94. 32. Gardner LP, Mookhtiar KA, Coleman JE. Initiation, elongation, and processivity of carboxyl-terminal mutants of T7 RNA polymerase. Biochemistry 1997;36:2908-18. 33. Martin M, Cho J, Cesare AJ, Griffith JD, Attardi G. Termination factor-mediated DNA loop between termination and initiation sites drives mitochondrial rRNA synthesis. Cell 2005;123:1227-40. 34. Ojala D, Montoya J, Attardi G. tRNA punctuation model of RNA processing in human mitochondria. Nature 1981;290:470-4. 35. Borst P, Grivell LA. Mitochondrial ribosomes. FEBS Lett 1971;13:73-88. 36. Dame RT. The role of nucleoid-associated proteins in the organization and compaction of bacterial chromatin. Mol Microbiol 2005;56:858-70. 37. Chen XJ, Butow RA. The organization and inheritance of the mitochondrial genome. Nat Rev Genet 2005;6:815-25. 38. Legros F, Malka F, Frachon P, Lombes A, Rojo M. Organization and dynamics of human mitochondrial DNA. J Cell Sci 2004;117:2653-62. 39. Iborra FJ, Kimura H, Cook PR. The functional organization of mitochondrial genomes in human cells. BMC Biol 2004;2:9. 40. Caron F, Jacq C, Rouviere-Yaniv J. Characterization of a histone-like protein extracted from yeast mitochondria. Proc Natl Acad Sci USA 1979;76:4265-9. 41. Diffley JF, Stillman B. DNA binding properties of an HMG1-related protein from yeast mitochondria. J Biol 366 Chem 1992;267:3368-74. 42. Megraw TL, Chae CB. Functional complementarity between the HMG1-like yeast mitochondrial histone HM and the bacterial histone-like protein HU. J Biol Chem 1993;268:12758-63. 43. Cho JH, Ha SJ, Kao LR, Megraw TL, Chae CB. A novel DNA-binding protein bound to the mitochondrial inner membrane restores the null mutation of mitochondrial histone Abf2p in Saccharomyces cerevisiae. Mol Cell Biol 1998;18:5712-23. 44. Chen XJ, Wang X, Kaufman BA, Butow RA. Aconitase couples metabolic regulation to mitochondrial DNA maintenance. Science 2005;307:714-7. 45. Kaufman BA, Newman SM, Hallberg RL, Slaughter CA, Perlman PS, Butow RA. In organello formaldehyde crosslinking of proteins to mtDNA: identification of bifunctional proteins. Proc Natl Acad Sci USA 2000;97:7772-7. 46. MacAlpine DM, Perlman PS, Butow RA. The numbers of individual mitochondrial DNA molecules and mitochondrial DNA nucleoids in yeast are co-regulated by the general amino acid control pathway. EMBO J 2000;19:76775. 47. Kaufman BA, Kolesar JE, Perlman PS, Butow RA. A function for the mitochondrial chaperonin Hsp60 in the structure and transmission of mitochondrial DNA nucleoids in Saccharomyces cerevisiae. J Cell Biol 2003;163:457-61. 48. Van Dyck E, Foury F, Stillman B, Brill SJ. A singlestranded DNA binding protein required for mitochondrial DNA replication in S. cerevisiae is homologous to E. coli SSB. EMBO J 1992;11:3421-30. 49. Meeusen S, Tieu Q, Wong E, Weiss E, Schieltz D, Yates JR, Nunnari J. Mgm101p is a novel component of the mitochondrial nucleoid that binds DNA and is required for the repair of oxidatively damaged mitochondrial DNA. J Cell Biol 1999;145:291-304. 50. Meeusen S, Nunnari J. Evidence for a two membranespanning autonomous mitochondrial DNA replisome. J Cell Biol 2003;163:503-10. 51. Hobbs AE, Srinivasan M, McCaffery JM, Jensen RE. Mmm1p, a mitochondrial outer membrane protein, is connected to mitochondrial DNA (mtDNA) nucleoids and required for mtDNA stability. J Cell Biol 2001;152:40110. 52. Bateman JM, Perlman PS, Butow RA. Mutational bisection of the mitochondrial DNA stability and amino acid biosynthetic functions of ilv5p of budding yeast. Genetics 2002;161:1043-52. 53. Parisi MA, Clayton DA. Similarity of human mitochondrial transcription factor 1 to high mobility group proteins. Science 1991;252:965-9. 54. Alam TI, Kanki T, Muta T, Ukaji K, Abe Y, Nakayama H, Takio K, Hamasaki N, Kang D. Human mitochondrial DNA is packaged with TFAM. Nucleic Acids Res 2003;31:1640-5. Chang Gung Med J Vol. 32 No. 4 July-August 2009 367 Chih-Chieh Mao, et al Mitochondrial DNA instability 55. Parisi MA, Xu B, Clayton DA. A human mitochondrial transcriptional activator can functionally replace a yeast mitochondrial HMG-box protein both in vivo and in vitro. Mol Cell Biol 1993;13:1951-61. 56. Larsson NG, Wang J, Wilhelmsson H, Oldfors A, Rustin P, Lewandoski M, Barsh GS, Clayton DA. Mitochondrial transcription factor A is necessary for mtDNA maintenance and embryogenesis in mice. Nat Genet 1998;18:231-6. 57. Tyynismaa H, Sembongi H, Bokori-Brown M, Granycome C, Ashley N, Poulton J, Jalanko A, Spelbrink JN, Holt IJ, Suomalainen A. Twinkle helicase is essential for mtDNA maintenance and regulates mtDNA copy number. Hum Mol Genet 2004;13:3219-27. 58. Garrido N, Griparic L, Jokitalo E, Wartiovaara J, van der Bliek AM, Spelbrink JN. Composition and dynamics of human mitochondrial nucleoids. Mol Biol Cell 2003;14:1583-96. 59. Bogenhagen DF, Wang Y, Shen EL, Kobayashi R. Protein components of mitochondrial DNA nucleoids in higher eukaryotes. Mol Cell Proteomics 2003;2:1205-16. 60. Ashley N, Harris D, Poulton J. Detection of mitochondrial DNA depletion in living human cells using PicoGreen staining. Exp Cell Res 2005;303:432-46. 61. He J, Mao CC, Reyes A, Sembongi H, Di Re M, Granycome C, Clippingdale AB, Fearnley IM, Harbour M, Robinson AJ, Reichelt S, Spelbrink JN, Walker JE, Holt IJ. The AAA+ protein ATAD3 has displacement loop binding properties and is involved in mitochondrial nucleoid organization. J Cell Biol 2007;176:141-6. 62. Hanson PI, Whiteheart SW. AAA+ proteins: have engine, will work. Nat Rev Mol Cell Biol 2005;6:519-29. 63. Yu XC, Weihe EK, Margolin W. Role of the C terminus of FtsK in Escherichia coli chromosome segregation. J Bacteriol 1998;180:6424-8. 64. Begg KJ, Dewar SJ, Donachie WD. A new Escherichia coli cell division gene, ftsK. J Bacteriol 1995;177:621122. 65. Aussel L, Barre FX, Aroyo M, Stasiak A, Stasiak AZ, Sherratt D. FtsK Is a DNA motor protein that activates chromosome dimer resolution by switching the catalytic state of the XerC and XerD recombinases. Cell 2002;108:195-205. 66. Gerbi SA, Strezoska Z, Waggener JM. Initiation of DNA replication in multicellular eukaryotes. J Struct Biol 2003;140:17-30. 67. Holt IJ, He J, Mao CC, Boyd-Kirkup JD, Martinsson P, Sembongi H, Reyes A, Spelbrink JN. Mammalian mitochondrial nucleoids: organizing an independently minded genome. Mitochondrion 2007;7:311-21. 68. Schon EA, Rizzuto R, Moraes CT, Nakase H, Zeviani M, DiMauro S. A direct repeat is a hotspot for large-scale deletion of human mitochondrial DNA. Science 1989;244:346-9. 69. Mita S, Rizzuto R, Moraes CT, Shanske S, Arnaudo E, Fabrizi GM, Koga Y, DiMauro S, Schon EA. Chang Gung Med J Vol. 32 No. 4 July-August 2009 70. 71. 72. 73. 74. 75. 76. 77. 78. 79. 80. 81. 82. 83. Recombination via flanking direct repeats is a major cause of large-scale deletions of human mitochondrial DNA. Nucleic Acids Res 1990;18:561-7. Goto Y, Nonaka I, Horai S. A mutation in the tRNA(Leu)(UUR) gene associated with the MELAS subgroup of mitochondrial encephalomyopathies. Nature 1990;348:651-3. Hammans SR, Sweeney MG, Brockington M, MorganHughes JA, Harding AE. Mitochondrial encephalopathies: molecular genetic diagnosis from blood samples. Lancet 1991;337:1311-3. Silvestri G, Ciafaloni E, Santorelli FM, Shanske S, Servidei S, Graf WD, Sumi M, DiMauro S. Clinical features associated with the A G transition at nucleotide 8344 of mtDNA (“MERRF mutation”). Neurology 1993;43:1200-6. Hammans SR, Sweeney MG, Wicks DA, Morgan-Hughes JA, Harding AE. A molecular genetic study of focal histochemical defects in mitochondrial encephalomyopathies. Brain 1992;115 (Pt 2):343-65. Yasukawa T, Suzuki T, Ohta S, Watanabe K. Wobble modification defect suppresses translational activity of tRNAs with MERRF and MELAS mutations. Mitochondrion 2002;2:129-41. Prezant TR, Agapian JV, Bohlman MC, Bu X, Oztas S, Qiu WQ, Arnos KS, Cortopassi GA, Jaber L, Rotter JI, Shohat M, Fischel-Ghodsian N. Mitochondrial ribosomal RNA mutation associated with both antibiotic-induced and non-syndromic deafness. Nat Genet 1993;4:289-94. Hutchin T, Haworth I, Higashi K, Fischel-Ghodsian N, Stoneking M, Saha N, Arnos C, Cortopassi G. A molecular basis for human hypersensitivity to aminoglycoside antibiotics. Nucleic Acids Res 1993;21:4174-9. Holt IJ, Harding AE, Petty RK, Morgan-Hughes JA. A new mitochondrial disease associated with mitochondrial DNA heteroplasmy. Am J Hum Genet 1990;46:428-33. Poyton RO, McEwen JE. Crosstalk between nuclear and mitochondrial genomes. Annu Rev Biochem 1996;65:563-607. Sue CM, Schon EA. Mitochondrial respiratory chain diseases and mutations in nuclear DNA: a promising start? Brain Pathol 2000;10:442-50. Smeitink J, van den Heuvel L. Human mitochondrial complex I in health and disease. Am J Hum Genet 1999;64:1505-10. Budde SM, van den Heuvel LP, Smeets RJ, Skladal D, Mayr JA, Boelen C, Petruzzella V, Papa S, Smeitink JA. Clinical heterogeneity in patients with mutations in the NDUFS4 gene of mitochondrial complex I. J Inherit Metab Dis 2003;26:813-5. Pecina P, Houstkova H, Hansikova H, Zeman J, Houstek J. Genetic defects of cytochrome c oxidase assembly. Physiol Res 2004;53 Suppl 1:S213-23. Suomalainen A, Majander A, Haltia M, Somer H, Lonnqvist J, Savontaus ML, Peltonen L. Multiple deletions of mitochondrial DNA in several tissues of a patient Chih-Chieh Mao, et al Mitochondrial DNA instability 84. 85. 86. 87. 88. 89. 90. 91. 92. 93. 94. with severe retarded depression and familial progressive external ophthalmoplegia. J Clin Invest 1992;90:61-6. Zeviani M, Bresolin N, Gellera C, Bordoni A, Pannacci M, Amati P, Moggio M, Servidei S, Scarlato G, DiDonato S. Nucleus-driven multiple large-scale deletions of the human mitochondrial genome: a new autosomal dominant disease. Am J Hum Genet 1990;47:904-14. Suomalainen A, Majander A, Wallin M, Setala K, Kontula K, Leinonen H, Salmi T, Paetau A, Haltia M, Valanne L, Lonnqvist J, Peltonen L, Somer H. Autosomal dominant progressive external ophthalmoplegia with multiple deletions of mtDNA: clinical, biochemical, and molecular genetic features of the 10q-linked disease. Neurology 1997;48:1244-53. Van Goethem G, Dermaut B, Lofgren A, Martin JJ, Van Broeckhoven C. Mutation of POLG is associated with progressive external ophthalmoplegia characterized by mtDNA deletions. Nat Genet 2001;28:211-2. Lamantea E, Tiranti V, Bordoni A, Toscano A, Bono F, Servidei S, Papadimitriou A, Spelbrink H, Silvestri L, Casari G, Comi GP, Zeviani M. Mutations of mitochondrial DNA polymerase gammaA are a frequent cause of autosomal dominant or recessive progressive external ophthalmoplegia. Ann Neurol 2002;52:211-9. Luoma P, Melberg A, Rinne JO, Kaukonen JA, Nupponen NN, Chalmers RM, Oldfors A, Rautakorpi I, Peltonen L, Majamaa K, Somer H, Suomalainen A. Parkinsonism, premature menopause, and mitochondrial DNA polymerase gamma mutations: clinical and molecular genetic study. Lancet 2004;364:875-82. Lewis S, Hutchison W, Thyagarajan D, Dahl HH. Clinical and molecular features of adPEO due to mutations in the Twinkle gene. J Neurol Sci 2002;201:39-44. Tyynismaa H, Mjosund KP, Wanrooij S, Lappalainen I, Ylikallio E, Jalanko A, Spelbrink JN, Paetau A, Suomalainen A. Mutant mitochondrial helicase Twinkle causes multiple mtDNA deletions and a late-onset mitochondrial disease in mice. Proc Natl Acad Sci USA 2005;102:17687-92. Kaukonen J, Juselius JK, Tiranti V, Kyttala A, Zeviani M, Comi GP, Keranen S, Peltonen L, Suomalainen A. Role of adenine nucleotide translocator 1 in mtDNA maintenance. Science 2000;289:782-5. Saada A, Shaag A, Mandel H, Nevo Y, Eriksson S, Elpeleg O. Mutant mitochondrial thymidine kinase in mitochondrial DNA depletion myopathy. Nat Genet 2001;29:342-4. Mandel H, Szargel R, Labay V, Elpeleg O, Saada A, Shalata A, Anbinder Y, Berkowitz D, Hartman C, Barak M, Eriksson S, Cohen N. The deoxyguanosine kinase gene is mutated in individuals with depleted hepatocerebral mitochondrial DNA. Nat Genet 2001;29:337-41. Morris AA, Taanman JW, Blake J, Cooper JM, Lake BD, 368 Malone M, Love S, Clayton PT, Leonard JV, Schapira AH. Liver failure associated with mitochondrial DNA depletion. J Hepatol 1998;28:556-63. 95. Vu TH, Sciacco M, Tanji K, Nichter C, Bonilla E, Chatkupt S, Maertens P, Shanske S, Mendell J, Koenigsberger MR, Sharer L, Schon EA, DiMauro S, DeVivo DC. Clinical manifestations of mitochondrial DNA depletion. Neurology 1998;50:1783-90. 96. Reichard P. Interactions between deoxyribonucleotide and DNA synthesis. Annu Rev Biochem 1988;57:349-74. 97. Kunz BA, Kohalmi SE, Kunkel TA, Mathews CK, McIntosh EM, Reidy JA. International Commission for Protection Against Environmental Mutagens and Carcinogens. Deoxyribonucleoside triphosphate levels: a critical factor in the maintenance of genetic stability. Mutat Res 1994;318:1-64. 98. Martomo SA, Mathews CK. Effects of biological DNA precursor pool asymmetry upon accuracy of DNA replication in vitro. Mutat Res 2002;499:197-211. 99. Nishino I, Spinazzola A, Hirano M. Thymidine phosphorylase gene mutations in MNGIE, a human mitochondrial disorder. Science 1999;283:689-92. 100. Spinazzola A, Marti R, Nishino I, Andreu AL, Naini A, Tadesse S, Pela I, Zammarchi E, Donati MA, Oliver JA, Hirano M. Altered thymidine metabolism due to defects of thymidine phosphorylase. J Biol Chem 2002;277:4128-33. 101. Wang J, Wilhelmsson H, Graff C, Li H, Oldfors A, Rustin P, Bruning JC, Kahn CR, Clayton DA, Barsh GS, Thoren P, Larsson NG. Dilated cardiomyopathy and atrioventricular conduction blocks induced by heart-specific inactivation of mitochondrial DNA gene expression. Nat Genet 1999;21:133-7. 102. Silva JP, Kohler M, Graff C, Oldfors A, Magnuson MA, Berggren PO, Larsson NG. Impaired insulin secretion and beta-cell loss in tissue-specific knockout mice with mitochondrial diabetes. Nat Genet 2000;26:336-40. 103. Elson JL, Samuels DC, Turnbull DM, Chinnery PF. Random intracellular drift explains the clonal expansion of mitochondrial DNA mutations with age. Am J Hum Genet 2001;68:802-6. 104. Jenuth JP, Peterson AC, Shoubridge EA. Tissue-specific selection for different mtDNA genotypes in heteroplasmic mice. Nat Genet 1997;16:93-5. 105. Dunbar DR, Moonie PA, Jacobs HT, Holt IJ. Different cellular backgrounds confer a marked advantage to either mutant or wild-type mitochondrial genomes. Proc Natl Acad Sci USA 1995;92:6562-6. 106. Holt IJ, Dunbar DR, Jacobs HT. Behaviour of a population of partially duplicated mitochondrial DNA molecules in cell culture: segregation, maintenance and recombination dependent upon nuclear background. Hum Mol Genet 1997;6:1251-60. Chang Gung Med J Vol. 32 No. 4 July-August 2009 369 DNA Ian J. Holt1 DNA DNA DNA DNA DNA DNA polymerase γ (POLG) DNA helicase Twinkle (Twinkle) transcription factor A (TFAM) Twinkle TFAM ATAD3 DNA DNA DNA DNA POLG1 Twinkle ANT1 TK2 dGK TP DNA DNA 2009;32:354-69) mitochondrial DNA ( DNA Medical Research Council Dunn Human Nutrition Unit, Cambridge, U.K. 97 12 26 333 5 Tel.: (03)3281200 2324; Fax: (03)3281200 2787; E-mail: [email protected] 1 97 6 17