* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download doc Genetics 03-22
Gene therapy of the human retina wikipedia , lookup
RNA interference wikipedia , lookup
Non-coding RNA wikipedia , lookup
Cancer epigenetics wikipedia , lookup
RNA silencing wikipedia , lookup
X-inactivation wikipedia , lookup
Metagenomics wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Gene therapy wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Oncogenomics wikipedia , lookup
Gene nomenclature wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Primary transcript wikipedia , lookup
Gene desert wikipedia , lookup
Ridge (biology) wikipedia , lookup
Public health genomics wikipedia , lookup
Genomic imprinting wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Human genome wikipedia , lookup
Point mutation wikipedia , lookup
Pathogenomics wikipedia , lookup
Genomic library wikipedia , lookup
Genetic engineering wikipedia , lookup
Gene expression programming wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Non-coding DNA wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Minimal genome wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome (book) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Transposable element wikipedia , lookup
Gene expression profiling wikipedia , lookup
Genome editing wikipedia , lookup
Designer baby wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
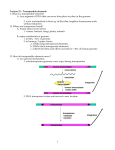
GENETICS – 03-22-2010 Slide: Hybrid dysgenesis is the result of DNA Transposons (P elements) P elements of Drosophila Are also nonautonomous versions with internal deletions. There are defective p elements that can transpose on their own but can with a complete P element. P repressor and transposase are a result from alternative splicing of the same gene- splicing of P element is tissue specific – Spliced different in somatic cells and germ line cells – Entron 3 is spliced out and a longer proteine of 87KD is made Both the repressor and the transposase are the products of the same gene. P element is one of these class 2 type DNA transposon – incredibly valuable for molecular bio – allowed developmental genetics for fruit flies. It can be used as a tool for mutagenesis – because they produce a tag of a known DNA sequence and therefore easy to identity the gene that’s causing mutation because this known DNA sequence is sitting in the middle of it. Approach called Transposon tagging. Harnessing of Eukaryotic Transposable elements P elements can also be used for introducing Genes back into Drosophila – used carrier to insert your gene of interest into the germ line of dysgenic flies – progeny will express that gene. Originally you needed to inject two plasmids into the germ cells of the dysgenic flies. Progeny will reflect the expression of that Gene. Tricks now to use different genetic vectors to express the gene in any kind of cell/tissue that you’d like. Transposable elements & host genomes: Why do organisms have these? Maybe they are simply selfish DNA capable of self-replication and that’s a necessary and sufficient explanation. But the transposable elements largely explain a long-standing paradox called the C-value paradox – the Genome size doesn’t really correlate with the complexity of the organism or with the number of genes. – Table of various organisms and genome size (protein-coding genes) and the number of predicted genes. Number of genes and size hard to argue it correlates well with complexity of organism What does correlate is that the larger genomes full of transposable elements. Transposons & the human genome: So for us – roughly 45 percent of genome composed by transposable elements – 20-40 thousand of LINe elements SINEs are very short but 20-30 thousand in our genome. Roughly 300 0000 DNA transposons – How can so many Transposons be maintained? What do all these things do and how do they avoid replicating totally out of control/killing their host – but if the host dies then the transposable elements die too. Most transposons are dead – dead because either they have a mutation in their transposase genes and they also have mutations in their flanking repeats – they can’t hop anymore – A lot of transposons are inactive –capable of mobility but kept in one place by repressors. Those transposons can be activated under certain conditions – could be advantageous for the organism because it could induce rapid mutation. They are found in between genes and introns. They are inconspicuous – they insert one into another – so if a transposon goes into another – not a great effect on a gene. There also seem to be safe havens – areas of the chromosome in which transposons really don’t move around – usually heterchromatic (highly condensed) regions – Host is able to inactivate to inactivate – often found in heterochromatin – kept off by methylation, etc. Transposable elements and genome structure: Useful – structural role around centromeres? Other host mechanisms related to those used to suppress virus replication. Transposable elements can be harnessed by their hosts – they can drive evolution of the genome – also play structural roles. The other thing that transposable elements can do is since you have homologous sequences – they can allow recombination events and that can make new combinations of genes or elements of genes. GENETIC DISSECTION 1 –Reverse Genetics & Functional Dissection: Functional dissection of genomes Gene expression analysis – reporter genes/in situ- trying to work from phenotype back to molecular function. Reverse Genetics & gene knockouts (e.g RNA) – starting with molecular info about a gene and you’re trying to work back to functional info. Reverse Genetics: What genes are encoded in a sequence? Bioinformatics & genome annotation – identifying where in the sequence particular genes are – Another tool is to look whether or not they are conserved between organisms – comparative genomics valuable for getting info. What roles do those genes play: when? Where? What kind of protein? How are they regulated? When? – one can find out when/where using molecular techniques – given that you know its sequence you can design hybrodization probes to find RNA expressed by that gene - look for population of RNA in a developmental time or a certain tissue. Where? – Expression analysis (transcript + protein) immuno blotting experiments – What does the Gene do? – one can use sequence to knock out the function of the Gene – so that mutational analysis can follow. How? – one can do biochemical experiments on the protein that it produces – (identity of protein, sub-cellular localization, modifications, interactors). FUNCTIONAL DISSECTION – GENE EXPRESION STUDIES – mRNA In situ hybridization – can be used to tell you where on a chromosome a particular gene is located – Perhaps now a more important use for situ hybridization is to do the same thing on a tissue section or a whole organism if small – and actually see the cells that are producing the messenger RNA of your favorite gene. Use RNA probe complementary to mRNA (“anti-mRNA) Label probe with radioactivity or with tag for colourmetric enzyme dissection. Reporter Genes: Promoter of genes includes place where RNA will bind and promoter also includes regulatory regions that drive tissue specific expression of the gene. Reporter protein: some protein not normally expressed – something that’s really easy to detect in the lab –