* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
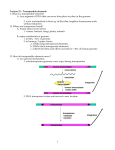
Download Ds - e-Acharya
Public health genomics wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Pathogenomics wikipedia , lookup
Gene expression profiling wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
X-inactivation wikipedia , lookup
Primary transcript wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Minimal genome wikipedia , lookup
Point mutation wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Genomic library wikipedia , lookup
Human genome wikipedia , lookup
Gene expression programming wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genome (book) wikipedia , lookup
Microevolution wikipedia , lookup
Genome editing wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Non-coding DNA wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Genome evolution wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Text Historical Background Transposable elements were discovered by Barbara McClintock through an analysis of genetic instability in maize. She was studying maize genetics and, in particular, was interested in understanding a phenomenon of genetic instability which was recognizable by the appearance of a spotted phenotype in a somatic cell tissue by using the ‘breakage–fusion–bridge’ strains to map genes. She identified two interesting loci-Dissociator (Ds) and Activator (Ac), which could transpose or change positions on the chromosome. In one case, she observed frequent chromosome break occurring at the Ds locus on chromosome-9 in Acdependent manner. Interestingly, this Ds element could move in the genome even to different chromosomes. In another case, the Ds locus seemed to regulate the expression of neighbouring genes. The change in position of the Ds element correlated with the expression of the C gene, and resulted in variegation of the kernel colour. Based on her observations, McClintock proposed that Ac and Ds were ‘controlling elements’ that regulated the expression of other genes. In the wake of the discovery of Ac/Ds, few more transposable element systems were identified in maize. In 1936, Marcus Rhoades observed frequent mutability of anthocyanin pigmentation in the aleurone of the endosperm in Mexican Black corn, giving a dotted phenotype. He determined that the unstable phenotype of the a1 gene that was responsible for the aleurone pigmentation was controlled by the Dotted (Dt) gene. The years following the discovery of the Ac/Ds elements were marked by genetic characterization of new transposons in maize. However, there was an uncertainty among the biologists whether transposons caused alteration in phenotype of other genes because they disrupt other genes by insertion or they cause alteration in phenotype by their ‘controlling’ influence. When the limitations of maize genetics to resolve this puzzle was clearly felt, research in bacterial genetics was breaking new grounds. In 1961, Jacob and Monod proposed the operon model for regulation of expression of the b-galactosidase gene in E. coli. Here the LacI gene regulates the expression of the Lac operon. Based on this observation, McClintock suggested a parallel between regulation of gene expression in bacteria and the effect of the ‘controlling elements’ in maize. In the following decade, more transposable elements, including IS and Tn elements in bacteria; Ty elements in yeast; and copia-like elements, Foldback elements and P elements in Drosophila were identified, mainly due to their ability to cause mutations, rearrange the genome, and regulate or modify the expression of other genes. The power of genetics in these systems, their simplicity and small size of genome, coupled with the advantage of emerging molecular techniques enabled the characterization of many transposable elements at the genetic and molecular level. Introduction Transposable elements are repetitive genomic sequences that are able to move or transpose from one chromosomal location to another. These mobile elements have been variously called jumping genes, mobile elements, insertion sequences and transposons. The formal name for this family of mobile genes is transposable element, and their movement is called transposition. Transposition has also been called illegimate recombination because it requires no homology between sequences nor is it site-specific. Consequently, it is relatively inefficient. The term transposon was coined by Hedges and Jacob in 1974 for a DNA segment which could move from one DNA molecule to other and usually disrupts genetic function resulting in the phenotypic variation. The transposable elements often replicate themselves. The replicative capacity of transposable elements is essence of their evolutionary success and has important implications for the dynamics between elements and their hosts. The transposable elements are present in both prokaryotic and eukaryotic chromosomes. These are even more prevalent in eukaryotic chromosomes than in bacterial chromosomes. For instance, 25% to 40% of mammalian chromosomes consist of transposable elements that have accumulated in the course of evolution. In addition, half of the spontaneous mutations seen in Drosophila are attributed to the movement and insertion of transposable elements. The contribution of the transposable elements to genome architecture and to the advent of genome novelty is likely to be dependent, at least in part, on the transposition mechanism, diversity, number and rate of turnover of transposable elements in the genome at any given time. The transposable elements transpose by a mechanism that involves DNA synthesis followed by random integration at a new target site in the genome. All transposable elements encode for transposase, the special enzyme activity that helps in the insertion of transposons at a new site and most of them contain inverted repeats at their ends. The major structural difference between bacterial transposable elements and their eukaryotic counterparts is the mechanism of transposition. Only eukaryotic genomes contain a special type of transposable elements, called retroposons, which use reverse transcriptase to transpose through an RNA intermediate. Retroposons are more common than transposons. They are either retroviral or nonviral. Viral retroposons encode for the enzymes reverse transcriptase and integrase and are flanked by long terminal repeats (LTRs) in the same way as retroviruses. Nonviral retroposons are the most frequently encountered transposons in mammals. The typical and most abundant nonviral retroposons are the short interspersed elements (SINEs) and the long interspersed elements (LINEs), which are usually repeated many times in the mammalian genome. Both SINEs and LINEs lack LTRs and are thought to transpose through special retrotransposition mechanism that involves transcription of one strand of the retroposon in to RNA. This RNA undergoes conformation change (looping) and provides a primer for the synthesis of single-stranded cDNA. The cDNA later serves as template for the synthesis of a double-stranded DNA that is inserted in the genome by an unknown mechanism. Characteristics of Transposable Elements 1. The transposable elements were found to be DNA sequences that code for enzymes which bring about the insertion of an identical copy in to a new DNA site. 2. Transposition events involve both recombination and replication processes which frequently generate two daughter copies of the original transposable elements. One copy remains at the parent site while the other appears at the target site. 3. The insertion of transposable elements invariably disrupts the integrity of their target genes. For example, if an IS (insertion sequence) becomes inserted in to an operon, it interrupts the coding sequence and inactivates the expression of the target gene in to which it inserts as well as any gene downstream in that same operon. This is because an IS contains transcription and/or translation termination signals that block the expression of other genes downstream in an operon. This ‘one-way’ mutational effect or polarity is referred to as polar mutation. 4. Since transposable elements carry signals for the initiation for the initiation for RNA synthesis, they sometimes activate previously dormant genes. 5. A transposable element is not a replicon, thus, it cannot replicate apart from the host chromosome the way that plasmids and phage can. 6. The frequency of transposition is low for most transposons, in the range of one transposition per 104 to 107 cells per generation in prokaryotes. 7. Transposons can grow or diminish by accumulating or losing DNA sequences that are in addition to those necessary for transposition. 8. There exists no homology between the transposon and the target site for its insertion. Many transposons can insert at virtually any position in the host chromosome or on to a plasmid. Some transposons seem to be more likely to insert at certain positions (hot-spots), but rarely at base-specific target sites. Classification of Transposable Elements in Eukaryotes Eukaryotic transposable elements are ubiquitous and widespread mobile genetic entities. These elements often make up a substantial fraction of the host genomes in which they reside. For example, approximately 1/2 of the human genome was recently shown to consist of transposable element sequences. Transposable elements are classified based upon their mechanism of transposition as well as by comparison of their genomic structures and sequences Class 1 In Class 1 transposons, the transposon is transcribed into RNA, which is reverse transcribed by the transposable element encoded reverse transcriptase. The cDNA is inserted at different loci by the transposase. Class 1 elements are also called retrotransposons, and include both autonomous and non-autonomous elements. Retrotransposons are further classified as LTR elements and non-LTR elements, based on their transposition mechanism and structure. LTRs are named after the ‘Long Terminal Repeats’ in direct orientation at either end of the transposon. The LTRs bear structural similarity to retroviruses. The autonomous elements encode at least two genes: the gag gene encodes a capsid-like protein and the pol gene encodes a polyprotein with various activities, including protease, reverse transcriptase, RNaseH and integrase. LTR elements are further classified into copia-like and gypsy-like elements. The non-LTR retrotransposons lack the long terminal repeats; instead they terminate by simple sequence repeats. They are divided into two: long interspersed nuclear elements (LINEs) and short interspersed nuclear elements (SINEs). Their coding regions include a gag-like protein, an endonuclease and reverse transcriptase. Class 2 The Class 2nd transposons transpose by direct cut-and-paste mechanism without any RNA intermediate. The transposase recognizes the terminal inverted repeat sequences (TIRs) on either ends of the transposable element, cuts it out and inserts at a new site. The transposase makes a staggered cut at the target site producing sticky ends, cuts out the transposon and ligates it into the target site. A DNA polymerase fills in the resulting gaps from the sticky ends and DNA ligase closes the sugar-phosphate backbone. This results in target site duplication and the insertion sites of DNA transposons may be identified by short direct repeats followed by inverted repeats. The duplications at the target site can result in gene duplication, which plays an important role in evolution. DNA transposons are grouped into several super families and families based on sequence similarity, length of the TIRs, and length of the TSD. hAT, CACTA, Mutator, PIF/Harbinger, Tc1/mariner are well represented class 2 transposable element groups in eukaryotes. Transposable elements in eukaryotes can be classified in to four types on the basis of their structure. The basis of their classification includes the size and nature of the terminal repeats or inverted repeats. The four types of transposons are: (i) With long terminal direct repeats (e.g. Copia in Drosophila, Ty in Yeast and IAP in mice); (ii) with long terminal inverted repeats (e.g. FB, TE in Drosophila); (iii) with short terminal inverted repeats (e.g. P and I in Drosophila; Ac/Ds in maize; TamI in snapdragon and TcI in Coenorhabditis) and (iv) without terminal repeats (e.g. Alu in mammals). Transposable Elements in Drosophila melanogastere The presence of transposable elements in D. melanogaster was first inferred from observations analogous to those that identified the first insertion sequence in E.coli. These sequences include the copia retroposon, the foldback (FB) family and the P elements. (a) Copia-like elements Its name reflects the presence of a large number of closely related sequences that code for abundant mRNAs, as the name derives from the Latin copia means abundance. The copia-like elements of Drosophila constitute atleast seven families, ranging in size from 5-8.5 kb. Members of each family appear at 100-1000 positions in the Drosophila genome. Each member carries a long, direct, terminal repeat and a short, imperfect inverted repeat. The number of copies of the copia-like element depends on the strain of fly. The members of the family are widely dispersed. The locations of the copia elements show a different (although overlapping) spectrum in each strain of D. melanogaster. These differences have developed over evolutionary periods. The copia element is ~5000 bp long, with identical direct terminal repeats of 276 bp. Each of the direct repeats itself ends in related inverted repeats. A direct repeat of 5 bp of targer DNA is generated at the site of insertion. The divergence between individual members of the copia family is slight, at <5%; variations often contain small deletions. All of these features are common to the other copia-like families, although their individual members display greater divergence. Copia-like elements cause a duplication of a characteristic number of base pairs of Drosophila DNA on insertion. Certain classic Drosophila mutations result from the insertion of copia-like and other elements, e.g. the white apricot (wa) mutation for eye colour is caused by the insertion of an element from the copia family in to the white locus. (b) P elements The P element in Drosophila is one of the best examples of exploiting the properties of transposable elements in eukaryotes. This element is 2907 bp long and features a 31 bp inverted repeat at each end. The element encodes a transposase. Although, the transposase is required for transposition, it can be supplied by a second element. Therefore, P element with internal deletions can be mobilized and then remain fixed in the new position in the absence of the second element, thus the P element can serve as a convenient marker. P elements do not utilize an RNA intermediate during transposition and can insert at many different positions in the Drosophila chromosome. The transposition of a P element is controlled by repressors encoded by the element. When P elements are mobilized, they produce a syndrome of traits known collectively as hybrid dysgenesis. These traits include temperaturedependent sterility, elevated rates of mutation, chromosome rearrangement and recombination. The syndrome is usually seen only in the progeny of males with autonomous P elements and females that lack P elements. These two kinds of strains are called ‘P’ and ‘M’ because they contribute paternally and maternally respectively to hybrid dysgenesis. The reciprocal cross P (female) × M (male) yields hybrids in which the dysgenic traits are much reduced, due to the maternal component of P element regulation by cytotype. The dysgenic traits can be explained largely by genomic changes due to P element transposition and excision in developing germ cells. The sterility is due to loss of germ cells early in development. It is more pronounced in females, where there are fewer germ cells to spare than in males, and at temperatures above 250C. The mutations come about through several mechanisms, but are primarily P insertions in to genes and imprecise excision of P element near genes. Chromosome rearrangements usually result from breakage at the sites of two or more P element insertions, followed by rejoining of the chromosome segments in a different order. P-induced recombination occurs preferentially in the genetic intervals containing mobile P elements, and usually within 2 kb of the insertion site. (b) Foldback (FB) elements Foldback elements are transposable elements found in many eukaryotic genomes. They are thought to contribute significantly to genome plasticity. These elements range from a few hundred to few thousand base pairs and carry long inverted repeats at the termini. The repeats may be adjoining or may be separated, but can always fold back to form stem and loop structures. In D. melanogaster, foldback elements have been shown to be involved in the transposition of large chromosomal regions and in the genetic instability of some alleles of the white gene. They may cause mutations by insertion or by their effect on gene expression due to their presence near a control region. Transposable Elements in Maize The transposable elements in maize were discovered by Marcus Rhoades and Barbara McClintock. These genetic elements in maize were found to be responsible for turning the expression of genes ‘on’ or ‘off’ hence the name ‘controlling elements’. These controlling elements when located on maize chromosomes represented specific sites for breakage and reunion of chromosomes leading to gross changes in chromosome structure. Some of the most popular controlling elements found in maize are activatordissociation (Ac-Ds) system and suppressormutator (spm) system. The number, type and location of these elements are characteristic of each maize strain. These elements can be classified in to two classes- (i) an autonomous element, which have the ability to excise and transpose, rendering sites of their insertion mutable, and (ii) non-autonomous elements, which do not transpose and are unstable and can be derived from autonomous elements of the same family. They become unstable only when an autonomous member of the same family is present elsewhere in the genome. When complemented by an autonomous member of the same family, the non-autonomous member may display properties of the autonomous element including its ability to transpose. (a) Activator-Dissociation (Ac-Ds) system Ac-Ds system is the most popular system and was discovered in 1950 by Barbara McClintock, when she found the presence of a genetic factor Ds (dissociation) caused high tendency towards chromosome breakage at the location, where it appears. These breaks could be located either cytologically at pachytene or genetically by uncovering of recessive alleles in a heterozygote. This property of Ds was dependent on another unlinked genetic factor Ac (Activator). Thus, in presence of Ac, Ds may undergo transposition or may cause chromosome breaks and an increase in the number of Ac element in the genome decreases the frequency of transposition or chromosome breaks by Ds. The functionally autonomous element, Ac, consists of 4563 nucleotide-pairs bounded by an 8 nucleotide-pair direct repeat. This direct repeat is created at the time that the element inserts in to a site on a chromosome. Other repeat sequences are found within the element itself. The most conspicuous of these are at the ends, where an 11 nucleotide-pair sequence at one end is repeated in the opposite orientation at the other end. These inverted terminal repeats are thought to play an important role in transposition. The Ds element shows considerable structural heterogeneity. One class of Ds element is derived from Ac element by deletion of internal sequences. Another class possesses the characteristic inverted terminal repeat sequences of Ac, as well as some of the sub-terminal sequences, but the remainder of the DNA is different. These unusual members of the Ac/Ds family are called aberrant Ds elements. A third class of Ds element is characterized by a peculiar piggybacking arrangement. One Ds element is inserted into another, but in an inverted orientation. It has been shown that these double Ds elements are responsible for chromosome breakage. The activating function of Ac elements is associated with a protein that they synthesize. Since this protein is involved in transposition, it is sometimes called the transposase of the Ac/Ds family. Deletion or mutations in the gene that encode this protein abolish the activating signal and explain why Ds elements, which have such lesions, cannot activate themselves. However, since this transposase is diffusible, a single Ac element can provide it to all the Ac and Ds elements in the genome. Therefore, we can say that Ac/Ds transposase is trans-acting. (b) Supressor-mutator (spm-dSpm) system The suppressor-mutator family of maize transposon was discovered by Barbara McClintock. In this family, the autonomous element is called Spm and the non-autonomous elements are called dSpm (‘d’ stands for deleted or defective). Spm elements are 8287 nucleotide-pairs long, including 13 nucleotide-pair inverted repeats. When they insert in to a chromosome, they create 3 nucleotide-pair target site duplication. The dSpm elements are smaller than the Spm elements because part of the DNA sequence has been deleted. These deletions disrupt the functioning of a gene carried by complete Spm elements and therefore prevent the synthesis of the gene’s product. Since this product is necessary for transposition, deleted dSpm elements are unable to stimulate their own movement. The Spm family was named because its elements can suppress the function of genes in to which they have transposed. This occurs when an inserted dSpm element interacts with an Spm element located elsewhere in the genome. Although, the dSpm insertion reduces the expression of the gene, it does not abolish it completely. However, when an autonomous Spm element is introduced in to the genome, the expression of the pigmentation gene is completely inhibited in most of the kernel. This indicates the ‘supressor’ action of the Spm element. In addition, this element stimulates the excision of the dSpm element in some of the cells, leading to clones in which gene function is partially restored. These clones, which are recognized by their moderate to heavy pigmentation, demonstrate the trans-acting, ‘mutator’ function of the Spm element. Transposable Elements in Yeast The yeast Saccharomyces cerevisiae carries transposable elements called Ty elements which comprise a family of dispersed repetitive DNA sequences that are found at different sites in different strains of yeast. Ty is an abbreviation for ‘transposon yeast’. Ty elements are retroposons, with a reverse transciptase activity, that transpose via an RNA intermediate. The frequency of Ty transposition is lower than that of most bacterial transposons, ~10-7 to 10-8. There is considerable divergence between individual Ty elements. Most elements fall into one of two major classes, called Ty1 and Ty917. They have the same general organization. Each element is 6.3 kb long; the last 330 bp at each end constitute direct repeats, called delta. Individual Ty elements of each type have many changes from the prototype of their class, including base pair substitutions, insertions and deletions. There are ~30 copies of Ty1 and ~6 of the Ty917 type in a typical yeast genome. In addition, there are ~100 independent delta elements called solo deltas. Not all of the Ty elements in any yeast genome are active. Most have lost the ability to transpose and are analogous to insert endogenous proviruses. These ‘dead’ elements retain the delta repeats, though, and as a result they provide targets for transposition in response to the proteins synthesized by an active element. A Ty element is altered by adding a promoter that can be activated by the addition of galactose. Activation of the promoter will increase transcription through the Ty element. Then an intron from another gene is inserted into the Ty element. Because the final product of transposition contains no intron, the intron must have been spliced out of an RNA transcript. This splicing must have taken place, where the primary transcript contains the intron but the final processed mRNA does not. This RNA is then copied by reverse transcriptase and integrated into the chromosomal DNA. The addition of galactose greatly increases the frequency of transposition of the altered Ty element. This increased frequency suggests the involvement of RNA, because galactose-stimulated transcription begins at the galactose-sensitive promoter. Because introns are excised only in the course of RNA processing, the transposed Ty DNA must have been copied from an RNA intermediate transcribed from the original Ty element and then processed by RNA splicing. The DNA copy of the spliced mRNA is then integrated into the yeast chromosome. Significance of transposable elements Transposons seem to play a role in evolution and biology by promoting rearrangement and restructuring of genes. Transposition may directly cause both deletion and inversion mutagenesis. They cause mutations by insertion into genes and affect the regulation of genes by inserting near promoters. They also provide substrates for genetic rearrangements and thus act as agents of genome evolution. Furthermore, transposable elements mediate the movement of host DNA sequences to new locations, enrich the genome with identical sequences positioned at different locations, and promote homologous recombination. Such recombination may eventually result in deletions, inversions and translocations. Transposons usually influence the expression of the proximity of their insertion sites. They are therefore extensively used as tools to create random insertion in bacteria, yeast and higher eukaryotes. They are also used in large scale genomic studies. Transposable elements have played an important role in the diversification and enrichment of mammalian transcriptomes through various mechanisms such as exonization and intronozation (the birth of new exons/introns from previously intronic/exonic sequences respectively), and insertion in to first and last exons. Transposable elements may play a crucial and essential role in evolution and could be the ‘missing link’ in our understanding of how multicellular and vertebrate organisms arose. The whole idea of transposons as purely selfish DNA is beginning to crumble. It now appears that at least some transposable elements may be essential to the organisms in which they reside. Even more interesting is the growing likelihood that transposable elements have played an essential role in the evolution of higher organisms, including humans. Transposable elements have also been used as vectors to transform the organisms genetically. In prokaryotes, the antibioticresistance marker carried by different transposons serves as a convenient marker. Transposable elements have many potential applications in genetic research including insertional mutagenesis, gene mapping, gene cloning, gene transfer within and between species, and identification of genes expressed in specific tissues at a particular time. All these genetic approaches are important in the study of molecular biology and evolution. As the number of known transposable elements increases and their properties are further documented, their utility as genetic research tools will become greater.