* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Nature Rev.Genet. 8
DNA vaccination wikipedia , lookup
Gene expression programming wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Epitranscriptome wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Ridge (biology) wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
Non-coding DNA wikipedia , lookup
Minimal genome wikipedia , lookup
Genome (book) wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Point mutation wikipedia , lookup
Oncogenomics wikipedia , lookup
Histone acetyltransferase wikipedia , lookup
Epigenetic clock wikipedia , lookup
Microevolution wikipedia , lookup
Gene expression profiling wikipedia , lookup
Transgenerational epigenetic inheritance wikipedia , lookup
Designer baby wikipedia , lookup
History of genetic engineering wikipedia , lookup
Neocentromere wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Primary transcript wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Behavioral epigenetics wikipedia , lookup
Epigenetics of depression wikipedia , lookup
DNA methylation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenetics wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Cancer epigenetics wikipedia , lookup
X-inactivation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Epigenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
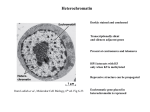
Heterochromatin Darkly stained and condensed Transcriptionally silent and silences adjacent genes Present at centromeres and telomeres HP1 interacts with H3 only when K9 is methylated Repressive structure can be propagated from Lodish et al., Molecular Cell Biology, 6th ed. Fig 6-33 Euchromatic gene placed in heterochromatin is repressed Histone Modifications Associated with Heterochromatin and Euchromatin from Lodish et al., Molecular Cell Biology, 6th ed. Fig 6-33 Initiation of Heterochromatin Assembly from Grewal and Gia, Nature Rev.Genet. 8, 35 (2007) Transcription factors and RNAi machinery bind to specific sequences or repetitive elements to recruit histone modifying enzymes Modified histones recruit HP1 HP1 recruits histone modifying enzymes to facilitate heterochromatin spread Boundary elements prevent further heterochromatin spread Mechanism of Heterochromatin Spreading HP1 binds to H3K9me3 HP1 recruits SUV39H1 methylase SUV39H1 methylates H3K9 on neighboring nucleosomes from Bannister et al., Nature 410, 120 (2001) Heterochromatin spreading is restricted by boundary elements Propagation of Heterochromatin Passage of the replication fork releases parental modified nucleosomes Nucleosome binding sites are created by recruitment of CAF1 by PCNA CAF1-bound HP1 recruits Suv39h, Dnmt1, and HDAC Methylated histones provide new HP1 binding sites Structural RNA associates from Maison and Almounzi, Nature Rev.Mol.Cell Biol. 5, 296 (2004) Heterochromatin Functions DNA or H3 methylation recruits adaptors such as HP1 Adaptors recruit effectors that are involved in chromosome segregation, gene silencing, transcriptional activation, and histone modification from Grewal and Gia, Nature Rev.Genet. 8, 35 (2007) Role of RNAi in Heterochromatin Formation in S. pombe dsRNA is transcribed from centromeric repeats or synthetic hairpin RNAs dsRNA is processed to siRNA siRNA promotes H3 K9 methylation by Clr4 Methylated H3 K9 recruits Swi6 to form silenced chromatin Transcription of the top strand of centromeric repeats is repressed Rdp1 activity ensures continuous dsRNA synthesis from Schramke and Allshire, Science 301, 1069 (2003) Recruitment of Clr4 by Swi6 chromatin leads to spread of heterochromatin Formation of Telomeric Heterochromatin RAP1 binds to C1-3A repeats Recruits Sir proteins Overexpression of Sir3 causes spread of telomeric heterochromatin Silencing decreases exponentially with distance from Grunstein, Cell 93, 325 (1998) Mechanism of Silencing at Telomeres Sir2 deacetylates histones Sir3,4 binds deacetylated histones and recruits additional Sir2 from Lodish et al., Molecular Cell Biology, 6th ed. Fig 7-35 Insulators Prevent the Progression of Condensed Chromatin Insulators protect genes from inappropriate signals Insulators block the action of distal enhancers from West et al, Genes Dev. 16, 271 (2002) Insulators prevent the spreading of heterochromatin gypsy Retrotransposon Contains an Insulator gypsy protects a transgene from position effects su(Hw) is necessary for enhancer blocking activity gypsy contains a su(Hw) binding site su(Hw) blocks the process that brings enhancer and promoter together Formation of insulator bodies at the nuclear periphery to divide the chromosome into looped domains Multiple su(Hw) binding sites can inhibit enhancer blocking activity Models for Heterochromatin Barrier Formation Stable block interrupts propagation of heterochromatin Active barrier recruits a complex containing chromatin remodeling activity from Donze and Kamakaka, BioEssays 24, 344 (2002) Epigenetics Heritable changes in gene function that cannot be explained by changes in gene sequences DNA methylation Histone variants and modifications Nucleosome positioning Epigenetic Modifications During Development Epigenetically imposed restrictions to plasticity are erased in the germ line Early mammalian development is characterized by progressive restriction of cellular plasticity accompanied by acquisition of epigenetic modifications Epigenetic modifications impose a cellular memory that accompanies and enables stable differentiation Epigenetic Modifications Within an Arabidopsis Chromosome Heterochromatin correlates with epigenetic marks from Zhang, Science 320, 489 (2008) DNA Methylation Methylation at CpG residues Sites of methylation Inactive X Imprinted loci Transposon-derived sequences CpG islands and CpG island shores Methylation patterns are reproduced at each round of cell division Methylated CpG Islands Inhibit Transcription from Portela and Esteller, Nature Biotechnol. 28, 1057 (2010) More than half of human promoters contain CpG islands Promoters are usually unmethylated Methylated DNA recruits methyl-CpG-binding domain proteins which recruit histone modifying and chromatin-remodelling complexes Unmethylated CpG islands recruit Cfp1 which associates with a histone methyltransferase creating H3K4me3 Methylated CpG Islands Inhibit Transcription from Portela and Esteller, Nature Biotechnol. 28, 1057 (2010) More than half of human promoters contain CpG islands Promoters are usually unmethylated Methylated DNA recruits methyl-CpG-binding domain proteins which recruit histone modifying and chromatin-remodelling complexes Unmethylated CpG islands recruit Cfp1 which associates with a histone methyltransferase creating H3K4me3 Methylation of Repetitive Sequences Stabilize Chromosomes from Portela and Esteller, Nature Biotechnol. 28, 1057 (2010) Unmethylated repetitive sequences cause reactivation of endoparasitic sequences RNA-dependent DNA Methylation in Plants Methylation occurs in transposons and repetitive elements PolIV transcribes ssRNA which is converted to dsRNA by RDR2 siRNA is produced by DCL3 and loaded onto AGO4 from Law and Jacobsen, Nature Rev.Genet. 11, 204 (2010) PolV produces IGN transcripts and recruits AGO4 siRNA-IGN duplex is formed and recruits DRM2 De Novo DNA Methylation in Mammals DNMT3L interacts with unmethylated H3K4 DNMT3A is recruited and activated and forms a tetrameric complex Active sites are separated by 8-10 bp and methylates opposite DNA strands from Law and Jacobsen, Nature Rev.Genet. 11, 204 (2010) Tetramer oligomerizes and results in 10 bp pattern of methylation on the same strand Establishment of DNA Methylation Pattern Most CpGs are unmethylated before implantation RNA pol II recruits H3K4 methyltransferase DNMT3L only binds unmethylated H3K4 and recruits DNA methyltransferases from Cedar and Bergman, Nature Rev.Genet. 10, 295 (2009) Propagation of DNA Methylation State Newly synthesized methylated DNA is hemimethylated NP95 binds hemimethylated DNA DNMT1 is a maintenance methyltransferase and binds PCNA NP95 links DNMT1 to hemimethylated DNA from Richly et al., BioEssays 32, 669 (2010) Mechanisms for Repression Mediated by MBD Proteins from Wade, BioEssays 23, 1131 (2001) MeCP2 Regulates Gene Expression in Response to Neural Activity Rett Syndrome is linked to mutations in MECP2 on the X chromosome MeCP2 binds CpG residues and silences target genes such as BDNF and corticotropin-releasing hormone from Bienvenu and Chelly, Nature Rev.Genet. 7, 415 (2006) from Miller, Science 314, 1356 (2006) Neural activity triggers MeCP2 phosphorylation and target gene activation Hippocampal neurons grow dendrites with fewer branches when MeCP2 is blocked Establishment of Cell Identity in Drosophila Embryos Segment identity is established by sequential spatially-localized expression of specific genes Regulatory genes are expressed transiently Transcriptional memory is maintained throughout development from Lodish et al., Molecular Cell Biology, 5th ed. Fig 15-24 Misexpression of Homeotic Genes Lead to Morphological Abominations from Lodish et al., Molecular Cell Biology, 5th ed. Fig 15-25 Polycomb and Trithorax Complexes Prevents changes in cell identity by preserving transcription patterns Chromatin is altered in a heritable manner Polycomb-group Proteins Maintains a silenced state Prevents chromatin remodelling Trithorax-group Proteins Maintains an active state Counteracts the action of PcG proteins Memory system composed of PcG and trxG complexes is linked to the histone code Model for PcG Formation and Function PcG complexes are recruited to PREs PRC2 complex methylates H3 K9 and K27 H3K27me3 recruits Polycomb and PRC1 complex H3K27me3 is segregated to both daughter DNAs to maintain repression from Lund and van Lohuizen, Curr.Opin.Cell Biol. 16, 239 (2004) Propagation of H3K27 Methylation EED2 binds H3K27me3 EED2 binding stimulates PRC2 activity EZH2 methylates H3K27 from Richly et al., BioEssays 32, 669 (2010) Demethylation of H3K27me3 Promotes Gene Activation PRC2 is recruited to H3K27me3 to mediate gene repression UTX and JMJD3 are recruited to Hox promoters and reverse repression Change in cell fate is mediated by H3K27 demethylation and H3K4 methylation, whose activities are present in the same complex from Rivenbark and Strahl, Science 318, 403 (2007) Trithorax Complex Mechanism of Action TrxC methylates H3K4 and recruits HAT and remodeling complexes Acetylated H3K9 prevents methylation, and prevents HP1 binding Somatic Cell Reprogramming Pleuripotency genes in somatic cells have methylated CpG islands Epigenetic marks must be reset to generate induced pleuripotent stem (iPS) cells from Cedar and Bergman, Nature Rev.Genet. 10, 295 (2009) Repressive histone methylation marks must be removed, followed by removal of DNA methylation which activates the gene Epigenetics and Heart Failure Brg1, a SWI/SNF component, is activated by cardiac stress Brg1 suppresses expression of a CKI to promote myocyte proliferation Brg1 promotes reprogramming to an embryonic state of transcription Brg1 forms a complex with HDAC and PARP and triggers a shift from a-myosin heavy chain expression to b-myosin heavy chain expression from Hang et al., Nature 466, 62 (2010) Epigenetic Modifications May Drive Cognitive Decline Chromatin remodeling in the hippocampus is necessary for stabilizing long term memories Aged mice have lower H4K12 acetylation HDAC inhibitor restores H4K12 acetylation and improved memory function from Sweatt, Science 328, 701 (2010) SIRT1 Deacetylase and Alzheimer’s Disease SIRT1 deacetylates RARb SIRT1 lof causes Alzheimer’s-like phenotype SIRT1 lof results in decreased a-secretase transcription and increased Ab production from Wolfe and Selkoe, Cell 142, 194 (2010) SIRT1 lof causes decreased Notch pathway activity and decreased neuronal repair Prion Epigenetics Prions template conformational conversion of other molecules of the same protein Prions are formed through an oligomeric nucleus, and the elongating polymer is severed by protein remodeling factors Prions are disseminated to daughter cells during cell division from Halfmann and Lindquist, Science 330, 629 (2010) Stress Accelerates Prion Appearance Abrupt changes have consequences for protein folding Prion-free cells are adapted to environment 1, but poorly adapted to environment 2 Prion formation and disappearance provide fitness advantages in different environments from Halfmann and Lindquist, Science 330, 629 (2010) Prions connect environmental conditions to acquisition and inheritance of new traits Co-suppression Increase in gene copy number results in decreased expression Dependent on PcG genes PcG complexes interact in trans from Pirrotta, Cell 93, 333 (1998) Imprinting Expression of only one allele of a locus Only 80 genes in mammals are imprinted Most imprinted genes are involved in growth control Imprinted genes involves allele specific methylation and is resistant to genome-wide demethylation Clusters of imprinted genes contain noncoding RNAs that are involved control allele-specific expression Imprinted Expression of the H19 and Igf2 Genes Maternal – H19 expression Paternal – Igf2 expression from Reik and Murrell, Nature 405, 408 (2000) Imprinting is Regulated by a Methylation-sensitive Boundary ICR is methylated in the male germ line ICR is protected from methylation in the female germ line by CTCF CTCF binding to the ICR in females prevents activation of Igf2 by downstream enhancer from Reik and Murrell, Nature 405, 408 (2000) In males, the downstream enhancer activates Igf2 and H19 expression is repressed by DNA methylation Imprinting of the PWS-AS Locus from Ferguson-Smith and Surani, Science 293, 1086 (2001) The AS-ICR is required for methylation and inactivation of the PWS-ICR in females to repress nearby genes The AS-ICR is nonfunctional in males allowing the PWS-ICR to activate nearby genes The PWS-ICR promotes expression of an antisense Ube3a transcript in males Dosage Compensation Mechanisms Genomes compensate for different numbers of sex chromosomes by adjusting gene expression levels from Straub and Becker, Nature Rev.Genet. 8, 47 (2007) X Inactivation in Mammals X inactivation is initiated from the Xic Xist and Tsix partially overlap and are transcribed in opposite directions from the Xic from Lodish et al., Molecular Cell Biology, 6th ed. Fig 22-7 Model for the Initiation of X Inactivation The Xic in female cells colocalize prior to X inactivation Low expression of Tsix from Xi leads to Xist transcription Xist RNA coats the Xi in cis The chromatin structure of Xi becomes condensed from Lodish et al., Molecular Cell Biology, 6th ed. Fig 22-7 Stepwise Progression of X Inactivation in Differentiating ES Cells One X chromosome is converted to facultative heterochromatin Xist transcription off the inactive X initiates chromatin modification events X inactivation is maintained epigenetically from Brockdorff, Trends Genet. 18, 352 (2002) Calico Cats One of the genes controlling fur color is on the X chromosome B – orange b - black Female mammals are genetic mosaics Random X inactivation early in embryonic development leads to patchworks of skin cells expressing each allele The Dosage Compensation Complex in Drosophila SXL in females prevents MSL2 translation MSL2 in males stabilizes roX, MSL1, and MSL3 DCC binds to high affinity sites on X chromosome from Gilfillan et al., FEBS Lett. 567, 8 (2004) DCC spreads to nearby sites on active chromatin H4K16 acetylation impedes formation of condensed chromatin structure DCC is Localized to the X Chromosome DCC localization is determined by staining of polytene chromosomes with anti-MSL1 DCC associates almost exclusively with transcribed regions from Straub and Becker, Nature Rev.Genet. 8, 47 (2007)