* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Genetics and Genomics of Core Short Tandem Repeat Loci
Medical genetics wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Molecular cloning wikipedia , lookup
Genetic engineering wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Molecular Inversion Probe wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Polymorphism (biology) wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Genomic library wikipedia , lookup
Designer baby wikipedia , lookup
DNA paternity testing wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Genomic imprinting wikipedia , lookup
Skewed X-inactivation wikipedia , lookup
X-inactivation wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
Neocentromere wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
Epigenomics wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Genome (book) wikipedia , lookup
DNA profiling wikipedia , lookup
DNA supercoil wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Helitron (biology) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Hardy–Weinberg principle wikipedia , lookup
Human genetic variation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Point mutation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Quantitative trait locus wikipedia , lookup
SNP genotyping wikipedia , lookup
Human leukocyte antigen wikipedia , lookup
Genetic drift wikipedia , lookup
Population genetics wikipedia , lookup
Microsatellite wikipedia , lookup
Haplogroup G-P303 wikipedia , lookup
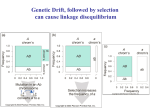
J Forensic Sci, March 2006, Vol. 51, No. 2 doi:10.1111/j.1556-4029.2006.00046.x Available online at: www.blackwell-synergy.com Genetics and Genomics of Core Short Tandem Repeat Loci Used in Human Identity Testing John M. Butler,Ph.D 2012.10.08 Sung Hwa Young . 13 genetic markers form the core of the FBI Laboratory’s Combined DNA Index System (CODIS) were selected in November 1997. Short tandem repeat (STR) loci dominate the genetic information that has been collected to date on human beings. In the U.S. and U.K. alone, more than 5 million profiles now exist in criminal justice DNA databases. The U.S.(13 core loci ): CSF1PO, FGA, TH01, TPOX, VWA, D3S1358, D5S818, D7S820, D8S1179, D13S317, D16S539, D18S51, and D21S11 http://cincinnati.com/niecincy/archive/2011/10/24/ The U.K. and Europe (10 core loci) additional markers : D2S1338 and D19S433 eight overlapping loci : FGA, TH01, VWA, D3S1358, D8S1179, D16S539, D18S51, and D21S11. This review article describes 1. Commonly used STR markers in terms of their population genetic variation and genomic locations. 2. Potential linkage of STR loci to genetic disease-causing genes 3. Desirable characteristics for additional STR loci 4. Commonly used Y-chromosome STR loci STR loci - ease of use in the form of commercial STR kits - human identity testing both in forensic casework and paternity testing - Missing persons investigations and mass disaster victim identification involve the same STR markers and kits - Observed allele ranges for each locus with with PCR product sizes and dye labels for the various STR kits STR allele sizes - measured relative to an internal size standard during electrophoresis - depending on the DNA strand that is dye labeled - may have a different apparent measured size than the actual DNA sequence PCR product size (bp) AmpFlSTR Identifiler™ kit (Applied Biosystems) 6-FAM (Blue) D8S1179 VIC (Green) D3S1358 NED (Yellow) PET (Red) LIZ (Orange) D21S11 TH01 D19S433 AMEL CSF1PO D13S317 D16S539 D2S1338 VWA D5S818 D7S820 TPOX D18S51 FGA GS500 LIZ size standard 1. genomic information and population genetic variation (1) Genomic information - the core loci are located on separate chromosomes - expected to segregate independently of one another during meiosis - use of the product rule in estimating random match probabilities with DNA profiles generated from multiple STR loci Chromosome 12 telomer e p (short arm) Band 3 12 p3 Band 5 12 q5 centromer e q (long arm) telom ere Figure 2.4, J.M. Butler (2005) Forensic DNA Typing, 2nd Edition © 2005 Elsevier Academic Press - the exceptions : CSF1PO and D5S818 (chromosome 5)- separated by approximately 26.3 Mb Penta D and D21(chromosome 21)- separated by approximately 24.4 Mb hundreds of population studies involving D5S818 and CSF1PO conducted on unrelated individuals have failed to show any signs of significant linkage between these two loci. (2) Population Variation 1) Allele Range and Variants - STR typing : size comparisons with standardized allelic ladders that possess the most common alleles new alleles can be discovered that occur outside the range defined by the commercially available allelic ladder ‘‘off-ladder’’ alleles can be variants with more or less of the core repeat unit than present in the common alleles found in the commercially available allelic ladder. these variant alleles may contain partial repeats or insertions/deletions in the flanking region close to the repeat 28.1 Figure 6.6, J.M. Butler (2005) Forensic DNA Typing, 2nd Edition © 2005 Elsevier Academic Press - Triallelic patterns have been observed for many of the core STR loci and recorded on the NIST STRBase Web site can occur as an imbalance in amounts between the three alleles (type 1) or equal amounts of all three alleles (type 2) Ex) TPOX, which occurs closest to the tip of a chromosome, has the highest number of observed triallelic patterns Thus, it is possible that this section of chromosome 2 is more likely to be duplicated in some individuals for telomere maintenance to keep the end of the chromosome intact 2) Characterizing a Variant Allele That Occurs Between Two Loci Locus 1 with only an allele ‘‘a’’ Locus 2 only has an allele ‘‘c’’ with an allele ‘‘b’’ occurring between the two loci the possible genotypes : locus1 (a,b) and locus2 (c,c) or locus1(a,a) and locus2 (b,c) Ex) if a green-colored peak occurs between D16S539 and D2S1338 in the Identifiler kit and only a single allele is observed in each of the D16 and D2 normal allele ranges, then the interlocus allele more likely belongs to D2S1338 because D2 has a higher heterozygosity. 3) Null Alleles with Commercial STR Kits Sequence variation does occur in the flanking regions surrounding STR loci Some PCR primers have been noted to be impacted by a primer binding site mutation, which can lead to allele dropout. Heterozygous alleles are well balanced 6 8 Imbalance in allele peak heights 6 8 No mutation Mutation in middle of primer binding site * 8 * Figure 6.9, J.M. Butler (2005) Forensic DNA Typing, 2nd Edition © 2005 Elsevier Academic Press Allele 6 amplicon has “dropped out” Mutation at 3’end of primer binding site (allele dropout) D5S818 (74), D16S539 (75), and D18S51 (76) alleles 4) Mutation Rates with comparisons between relatives in parentage testing and kinship analysis, mass disaster victim Identification : mutational events can play a significant role. SE33, FGA, D18S51 : the loci with the highest mutation rate the most polymorphic and possess the highest number of alleles (3) Population Studies - As of early 2005, this list contains 365 population studies based on 183 literature references. - OmniPop (Brian Burritt) permits calculation of a user inputted profile’s frequency using allele frequencies from 166 published population surveys. - Large data sets typically identify a greater number of rare alleles as more individuals in a population are included in the analysis. 2. Potential Linkage to Disease Genes (1) The X-chromosome STR locus HumARA : a CAG repeat located in a coding region located in a gene coding region (i.e., exon) trinucleotide repeats, which can be prone to expansions that cause genetic defects (2) to be useful in tracking various genetic diseases through loss of heterozygosity or allelic imbalance. ex) - D8S1179 was used to localize a gene connected to Meckel–Gruber syndrome (monogenic cause of neural tube defects), elevated risk for cardiovascular disease - TH01 (the first intron of the tyrosine hydroxylase gene) : schizophrenic and bipolar disorders - Individuals possessing TH01 allele 7 : less nicotine dependence - D21S11(a useful test) : trisomy-21(Down’s syndrome), three alleles in any polymorphic marker found on chromosome 21 - D18S51 : trisomy-18 (Edwards’ syndrome) - cancer : loss of heterozygosity or extreme allelic imbalance 3. Additional STRs Beyond the Current Core Loci (1) Less polymorphic loci have lower mutation rates, which can make them more useful in some parentage testing situations (2) Two or three moderately polymorphic STR loci on separate chromosomes would be more powerful when the product rule was applied and would easily fit into the same PCR product space ex) a higher molecular weight FGA allele ( 40 repeat units or 160 bp) (3) DNA types can be recovered more effectively from degraded DNA samples when the PCR . products are smaller future loci should contain a more compact allele range and be able to be amplified as small PCR products 4. Y-Chromosome STR Loci (1) Found only in males, specific to the male portion of a male–female DNA mixture (sexual assault cases) (2) passed from father to son without changes (3) Alleles observed with Y-STR markers are concatenated to form a haplotype (4) Y-STR results from individual loci cannot be combined with the product rule, because the core Y-STR loci are all on the nonrecombining portion of the Y chromosome Figures for John M. Butler’s Advanced Topics in Forensic DNA Typing: Methodology (2012) 5. Conclusions (1) STR markers : important tools for human identity testing continue to be widely used for many years : their high degree of variability, ease of use in multiplex amplification formats and implementation in National DNA Databases (2) A uniform set of core STR loci : the capability for national and international sharing of criminal DNA profiles. (3) Robust commercial STR kits : reliable amplification of these core loci from small amounts of starting DNA template. (4) Resulting STR profiles : high powers of discrimination to be achieved among both related and unrelated individuals.