* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Genome Biology and
Molecular Inversion Probe wikipedia , lookup
Adeno-associated virus wikipedia , lookup
Point mutation wikipedia , lookup
Genomic imprinting wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Gene desert wikipedia , lookup
Gene expression profiling wikipedia , lookup
Epigenomics wikipedia , lookup
Human genetic variation wikipedia , lookup
Genetic engineering wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Segmental Duplication on the Human Y Chromosome wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Copy-number variation wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Microevolution wikipedia , lookup
Oncogenomics wikipedia , lookup
Genome (book) wikipedia , lookup
Designer baby wikipedia , lookup
Public health genomics wikipedia , lookup
Transposable element wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
DNA sequencing wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Microsatellite wikipedia , lookup
History of genetic engineering wikipedia , lookup
Non-coding DNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Minimal genome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Helitron (biology) wikipedia , lookup
Human genome wikipedia , lookup
Pathogenomics wikipedia , lookup
Genome editing wikipedia , lookup
Genomic library wikipedia , lookup
Metagenomics wikipedia , lookup
Human Genome Project wikipedia , lookup
Whole genome sequencing wikipedia , lookup
Genome Biology and
Biotechnology
The genomics revolution
Prof. M. Zabeau
Department of Plant Systems Biology
Flanders Interuniversity Institute for Biotechnology (VIB)
University of Gent
International course 2005
The Human Genome Project
1990
1995
2000
2005
Human Genome Project
Technological innovations
High throughput automation
Large scale genome sequencing
<1Mb/year
1000-fold
>1000 Mb/year
20.000 Mb/year
Technological Innovations
1.
High throughput fingerprinting of BAC clones
–
Construction of physical maps
–
Starting DNA for large scale sequencing
1
2 Mb
Technological Innovations
1. High throughput fingerprinting of BAC clones
–
Construction of physical maps
2. Improvements of the dideoxy sequencing technique
–
Fluorescent labeling and improved sequencing enzymes
3. Improved sequencing strategies
–
Shotgun sequencing, improved shotgun libraries
4. Software for automated interpretation of
fluorograms
–
Assigns 'assembly-quality scores' to each base in the assembled
sequence
–
Assembly of high quality sequence contigs
Shotgun DNA Sequencing Strategy
BAC clone
High throughput automation
¤ Automated DNA sequence gel readers
–
First generation: slab gel-based DNA sequencers
•
•
•
–
32 – 96 samples per run
Manual loading
Difficulties in lane tracking causing considerable losses in data
Second generation: capillary DNA sequencers
•
•
Automated loading, allowing unattended operation and perfect lane
tracking
20 * 96 samples/day = ~2 million bases of raw sequence/day
¤ Automation of sample preparation and handling
–
Liquid handling robots made the up scaling feasible
–
Eliminated most of the “human error”
Sequencing Complex Genomes : the Challenge
¤ Difficulties arise because of repeated sequences
– Small amounts of repeated sequence pose little problem for
shotgun sequencing
• Bacterial genomes (about 1.5% repeat)
– Mammalian genomes are filled (> 50%) with repeated sequences
• Interspersed repeats derived from transposable elements
• Large duplicated segments with high sequence identity (98–99.9%),
– Repeated sequences complicate the correct assembly of shotgun
sequence reads
¤ Two strategies for sequencing complex genomes
– Hierarchical shotgun sequencing strategy ('map-based', 'BACbased' or 'clone-by-clone‘ strategy)
– Whole genome shotgun (WGS) sequencing strategy
Hierarchical Shotgun Sequencing Strategy
Reprinted from: International Human Genome Sequencing Consortium Nature 409, 860 (2001)
Whole Genome Shotgun Sequencing
¤ Different insert sizes of cloned DNA
– 2 kb in multi copy vectors
– 10 kb in fosmid vectors
– 100 - 200 kb in BACs
Reprinted from: Venter et. al., Science 280: 1540 (1998)
Whole-genome shotgun sequence assembly
STS Sequence tagged Sites
Reprinted from: Venter et. al., Science, 291, 1304 (2001)
Comparison of the two strategies
¤ The hierarchical shotgun sequencing strategy is
– Slower and has a higher upfront cost
• create a detailed physical map of clones
• Sequencing of 10.000s of individual BAC clones involves more
handling steps
– Is indispensable for the production of a finished sequence
¤ The whole-genome shotgun approach is
– Faster and more cost effective
• Fully exploits the potential of a streamlined robotics-based
operation
– But, cannot deliver more than a (high quality) draft sequence
Draft Sequences versus Finished Sequences
¤ Draft genome sequences
– High quality draft sequence high (8 to 10-fold) coverage
• Yields sequence contigs that cover 95% - 98% of the sequence
– Draft sequence is by definition incomplete
• 10.000 – 100.000 gaps
• Incorrectly assembled sequences – duplicated segments
¤ Finished genome sequences
– Close gaps and resolve ambiguities in draft sequences
• Correct order and orientation of sequence contigs
• Resolution of duplicated regions: collapsed in the draft sequence
• Standard error rate: < 1 error per 10,000 bases
Sequencing Complex Genomes
¤ Projects currently underway use
– Model organisms where a finished genome sequence is
indispensable use a combination of the two approaches
• Human, Mouse, Drosophila, zebrafish
– Whole genome shotgun to generate high quality drafts
• Comparative genome analysis
– Hierarchical strategy for genomes with repetitive DNA is
clustered in centromeres or telomeres
• Plant genomes
– Alternative strategies
• Methyl filtration or Cot enriched libraries are used for
particular (large) plant genomes
Genome sequencing: progress to date
¤ Extraordinary progress in sequencing technologies
development in the past 15 years has resulted in
– Completion of the human genome project ahead of schedule
(2004)
– Over 30 eukaryotic genome sequences (including 6 vertebrate
genomes)
– Over 200 bacterial and archean genome sequences
¤ The completion of the human genome marks the “end
of the beginning”
– Many more genomes are to follow
– awaits the daunting task of unraveling its secrets
Genome Sequencing Milestones
1995
6
8
7
9
2000
H. influenza
1
2
3
4
2005
Human chrom 20
S. cerevisae
S. pombe
yeasts
Fugu
Tetrahodon
Mouse
Rat
Anopheles
Chicken
Neurospora
alga
Ciona
silkworm
C. elegans
Human chrom 21 & 22
Drosophila melanogaster
Arabidopsis thaliana
Human working draft
Human finished
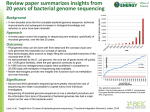
The global sequencing output to date
Equivalent of
15 human genomes
Feb 2004
GenBank website: http://www.ncbi.nlm.nih.gov/Genbank/GenbankOverview.html
Annotation of Genome Sequences
¤ The challenge of identifying genes in genomic
sequences varies greatly among organisms
– Gene identification is almost trivial in bacteria and yeasts
• Genes are readily recognized by ab initio analysis as ORFs
coding for >100 amino acids (no introns)
– Smaller ORFs and overlapping genes are missed
– Gene identification is relatively straightforward in small genomes,
such as worm, plant and Drosophila
• Coding sequences comprise a large proportion of the genome
(~50%)
• Introns are relatively small
– Gene identification is very difficult in large complex genomes
(mammalian)
• Coding sequences comprise only a few per cent of the genome
• Exons are small and introns are very large
Gene Prediction Methods
¤ Three basic approaches
– Direct evidence of transcription: ESTs or full length cDNAs
• Limited to the more frequently expressed genes – misses rarely
expressed genes
– Indirect evidence based on sequence similarity to previously
identified genes and proteins
• Correctly identifies genes, but these may be pseudogenes
• Limited to known genes – misses unknown genes
– Ab initio prediction of groups of exons on the basis of hidden
Markov models (HMMs) that
• Combine statistical information about splice sites, coding bias and
exon and intron lengths (for example, Genscan, Genie and FGENES)
Genome annotation: state-of-the-art
¤ Genome annotation is an ongoing effort
– In all published model genomes the gene counts and gene models
are constantly being revised
• The gene numbers do not change drastically (10% range)
• Gene models are often subject to considerable change
– Improvements will result from
• The availability of many more complete genome sequences
• Comparative genome analysis between related species
• Larger databases of confirmed gene and protein sequences
¤ The challenge ahead is the identification of
regulatory sequences
– Comparing multiple genomes related species
• Yeast and the mammalian genome projects
Principal Types of Microarrays
¤ Oligonucleotide arrays
– Produced by in situ synthesis, of
short 25-70 mer oligonucleotides
onto glass slides
¤ Spotted arrays
– Produced by robotic deposition of
nucleic acids (PCR products,
plasmids or oligonucleotides) onto
a glass slide
Reprinted from: Lockhart and Winzeler, Nature 405, 827 (2000)
Photolithographic microarrays
Reprinted from: Lipshutz et. al., Nature Genet. 21, 20 (1999)
Spotted Microarrays
¤ Technology developed in the early 90’s
– Deposit micro droplets (nanoliter volumes) onto chemically treated
glass surfaces
• Multi-pin tools transfer liquid from micro titer plates on glass surface
• Chemical coating is necessary for binding nucleic acids
DNA spotting
Prehybridization Blocking
Silanized Slides
Transcribe RNA to
labeled cDNA
Washing
Hybridization
Future Perspectives
¤ Technology developments will continue to drive the
genomics field
– Large scale genome sequencing improvements
• Higher throughput and accuracy– more genomes
• Lower the cost of genome sequencing
– Microarray technology improvements
• Higher probe densities – higher resolution data sets
• Enable novel applications – functional genomics
– Revolutionary new technologies are now being pioneered
• 1000€ (human) genome programmes
Recommended reading
¤ Genome sequencing
– The sequencing of the human genome
• International Human Genome Sequencing Consortium Nature 409,
860 (2001)
¤ Microarrays
– Photolithographic oligonucleotide arrays
• Lipshutz et. al., Nature Genet. 21, 20 (1999)
Further reading
¤ Large scale sequencing technologies
– Whole-genome shotgun sequencing
• Venter et. al., Science 280: 1540 (1998) ,
• Venter et. al., Science, 291, 1304 (2001)
– High throughput fingerprint analysis of large-insert clones
• Marra et al., Genome Res 7: 1072–1084 (1997)
¤ Microarray technologies
– Mask driven photolithographic oligonucleotide arrays
• Lockhart and Winzeler, Nature 405, 827 (2000)
– Maskless photolithography oligonucleotide arrays
– Nuwaysir et al., Genome Res. 12, 1749 (2002)