* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Yeast whole-genome analysis of conserved regulatory motifs
Epigenetics wikipedia , lookup
History of genetic engineering wikipedia , lookup
RNA interference wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Human genome wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epitranscriptome wikipedia , lookup
Pathogenomics wikipedia , lookup
Public health genomics wikipedia , lookup
Epigenetics in stem-cell differentiation wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Gene desert wikipedia , lookup
Microevolution wikipedia , lookup
Genome (book) wikipedia , lookup
Non-coding DNA wikipedia , lookup
Histone acetyltransferase wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Genomic imprinting wikipedia , lookup
Minimal genome wikipedia , lookup
Designer baby wikipedia , lookup
RNA silencing wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Primary transcript wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Gene expression programming wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Genome evolution wikipedia , lookup
Non-coding RNA wikipedia , lookup
Ridge (biology) wikipedia , lookup
Epigenomics wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Gene expression profiling wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Epigenetics of human development wikipedia , lookup
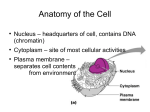
Fly ModENCODE data integration update Manolis Kellis, MIT Broad Institute of MIT and Harvard MIT Computer Science & Artificial Intelligence Laboratory modENCODE integration goals • Annotate all functional elements – Enhancers, promoters, insulators, silencers – Protein-coding genes, RNA genes, alternative splice forms • Understand their dynamics – Tissue- and stage-specific activity of each type of element • Mechanisms – Relative roles of histones, chromatin, specific/general TFs – Sequence specificity, regulatory motifs and grammars • Community involvement will be key – Seeking both computational and experimental partners – Large-scale: Complementary datasets / computation – Small-scale: Directed follow-up studies / genes, pathways • Drosophila 2009 modENCODE workshop discussion Each dataset is supported by all others Data Integration efforts Nucleosomes Henikoff Already presented Underway Transcripts Chromatin Celniker Karpen White Mac Alpine TFs/Chromatin Replication Lai Small RNAs • Each type of element requires multiple data types – – – – – – – Protein genes RNA genes Promoters Enhancers Transcripts Heterochromatin Initiation sites modENCODE is not alone • Community data types Nucleosomes – Boundaries Henikoff Transcripts Chromatin Karpen DNAse HS 12flies (+8 flies) Celniker Dam mapping Mac Alpine Boundaries – Small RNAs etc TFs/Chromatin Lai Small RNAs – evolutionary properties (correlations with conserved/nonconserved properties) – Dam mapping White Replication – DNAse HS sites, low buoyant density (protein binding) • Techniques and functional genomics – Gene Disruption projects – RNAi collection – Recombineering – Computational analyses Comparative resources for Drosophila genomes New Species D. ficusphila D. biarmipes D. elegans D. kikkawai D. eugracilis D. takahashii D. rhopaloa D. bipectinata Dist 0.80 0.70 0.72 0.89 0.76 0.65 0.66 0.99 done priority1 priority2 • Identify functional elements by their evolutionary signatures: complement experimental studies Evolutionary signatures for diverse functions Protein-coding genes - Codon Substitution Frequencies - Reading Frame Conservation RNA structures - Compensatory changes - Silent G-U substitutions microRNAs - Shape of conservation profile - Structural features: loops, pairs - Relationship with 3’UTR motifs Stark et al, Nature 2007; Clark et al, Nature 2007 Regulatory motifs - Mutations preserve consensus - Increased Branch Length Score - Genome-wide conservation Frequency Fraction Functional annotation of Novel Transcripts using evo. sigs -20 0 20 40 60 CSF Score (best 30 aa window) CSF = Heuristic metric for codon substitution frequency -20 0 20 40 60 CSF Score (best 30 aa window) 73 Putative protein coding 57 Putative non-coding Mike Lin, Jane Landolin, Sue Celniker Discover motifs associated with binding 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 Consensus CTAATTAAA TTKCAATTAA WATTRATTK AAATTTATGCK GCAATAAA DTAATTTRYNR TGATTAAT YMATTAAAA AAACNNGTT RATTKAATT GCACGTGT AACASCTG AATTRMATTA TATGCWAAT TAATTATG CATNAATCA TTACATAA RTAAATCAA AATKNMATTT ATGTCAAHT ATAAAYAAA YYAATCAAA WTTTTATG TTTYMATTA TGTMAATA TAAYGAG AAAKTGA AAANNAAA RTAAWTTAT TTATTTAYR MCS 65.6 57.3 54.9 54.4 51 46.7 45.7 43.1 41.2 40 39.5 38.8 38.2 37.8 37.5 36.9 36.9 36.3 36 35.6 35.5 33.9 33.8 33.6 33.2 33.1 32.9 32.9 32.9 32.9 Matches to known engrailed (en) reversed-polarity (repo) araucan (ara) paired (prd) ventral veins lacking (vvl) Ultrabithorax (Ubx) apterous (ap) abdominal A (abd-A) fushi tarazu (ftz) broad-Z3 (br-Z3) Antennapedia (Antp) Abdominal B (Abd-B) extradenticle (exd) gooseberry-neuro (gsb-n) Deformed (Dfd) Tissue specific target expression Promoters 25.4 5.8 11.7 4.5 13.2 16 7.1 7 20.1 3.9 17.9 10.7 19.5 5.8 14.1 1.8 5.4 3.2 3.6 2.4 57.2 5.3 6.3 6.7 8.9 4.7 7.6 449.7 11 30.7 Enhancers 2 4.2 2.6 16.5 0.3 3.3 1.7 2.2 4.3 0.7 1.2 2 5.4 1.7 2.8 0 4.6 -0.5 0.6 6 1.7 1.6 2.7 0.3 0.8 0.8 Ability to discover full dictionary of regulatory motifs de novo Stark et al, Nature, 2007 Initial regulatory network for an animal genome • ChIP-grade quality – Similar functional enrichment – High sens. High spec. • Systems-level – – – – 81% of Transc. Factors 86% of microRNAs 8k + 2k targets 46k connections • Lessons learned – Pre- and post- are correlated (hihi/lolo) – Regulators are heavily targeted, feedback loop Sushmita Roy Kheradpour et al, Genome Research, 2007 Temporal latencies in regulatory networks • TF-specific latencies, coherent with TF function • Latencies associated with network motifs • Extensions to tissue-specific networks Rogerio Candeias Incorporating ENCODE functional datasets Pouya Kheradpour, Jason Ernst, Chris Bristow, Rachel Sealfon modENCODE and gene regulation Goal: Understand the DNA elements responsible for gene regulation: • The regulators: TFs, GFs, miRNAs, their specificities • The regions: enhancers, promoters, insulators • The targets: individual regulatory motif instances • The grammars: combinations predictive of tissue-specific activity Building blocks of gene regulation Our tools: Comparative genomics & large-scale experimental datasets. • Evolutionary signatures for promoter/enhancer/3’UTR motif annotation • Chromatin signatures for integrating histone modification datasets • TFs, GFs, motifs, instances associated with tissue-specific activity • Infer regulatory networks, their temporal and spatial dynamics Integrate diverse datasets Sequence motifs predictive of insulators • Understand specificity of each factor • How predictable are these of binding • Motif combinations and grammars GAF, check Su(Hw), check CTCF, check BEAF-32, variant CP190, novel Pouya Kheradpour Motifs specific to each insulator Mod(mdg4), novel SPP, 40bp window Narrow Peak Interval Rank x104 Performance (higher is better) Fraction overlapping CTCF motif instances Motif instances correlate with ChIP peaks Recovery of CTCF inst. at 90% confid. Peak size • CTCF motif instances correlate strongly with narrow peak calls from multiple peak callers, even at 40bp window • Correlation extends down rank link (to all 50,000 peaks) • Implications for peak calling and for motif discovery Pouya Kheradpour, Ben Brown Motifs and tissue-specific chromatin marks • • • • Active marks The NF-κB motif is enriched in H3K4me2 regions found uniquely in GM12878 cells It is likewise enriched in the uniquely bound regions for other active marks Conversely, it is enriched in the uniquely unbound regions for the repressive mark H3K27me3 We find that NF-κB is also over expressed in GM12878, suggesting a causative explanation Pouya Kheradpour Repressive mark NF-κB motif Fold enrichment or over expression Motifs and stage-specific chromatin marks H3K27me3 • abd-A motif is enriched in new H3K27me3 regions at L2 – Coincides with a drop in the expression of abd-A – Model: sites gain H3K27me3 as abd-A binding lost Fold enrichment • Additional intriguing stories found, to be explored or over expression What about combinations of chromatin marks? Jason Ernst A hidden Markov model for chromatin state Transcription Start Site Enhancer Transcribed Region DNA Observed Histone Modifications Most likely Hidden State 1 2 3 4 Highly Likely Modifications in State 0.8 0.8 .8 1: 4: 0.7 2: 0.9 3: 0.9 5: 0.9 0.8 6: 5 5 5 5 5 Even though modification was not observed can still infer correct state based on neighboring locations that this state is likely of the same type as its neighboring states 6 6 6 20 distinct chromatin states, combinations of marks • Combinations of chromatin marks – More informative than individual marks (A&B ≠ A&C) – Small number of states (20 instead of all 2 million=221) – Allow study of co-occurrence patterns, independence… Each chromatin state associated w/ distinct function Tentative annotations • Reveals active/repressed promoters & enhancers • Distinct enrichments for 5’UTR/3’UTR/transcripts • Distinct chromatin properties of exons / introns Transcriptional unit enrichment Transcription start site (TSS) enrichment Transcription termination site (TTS) enrichment Transcriptional unit enrichment Chromatin signatures as context for TF analysis • TF role in establishing chromatin states • Chromatin role in modulating TF function Specific enrichment for DV and AP factors Functions of 20 distinct chromatin states in fly Chromatin marks DV enhancers AP enhancers General TFs Insulators Replication Motifs The grand challenge ahead Binding sites of every developmental regulator Dorsal-Ventral Annotations & images for all expression patterns Sequence motifs for every regulator CTCF, check GAF, check Su(Hw), check Anterior-Posterior Expression domain primitives reveal underlying logic BEAF-32, variant CP190, novel Mod(mdg4), novel Understand regulatory logic specifying development Summary of our lab’s experience in (mod)ENCODE • Protein-coding genes (Mike Lin) – Hubbard: Predict new genes, evaluate novel genes – Celniker: Distinguish coding/non-coding transcripts • Chromatin domains (Jason Ernst) – Karpen: Chromatin states in Drosophila – Bernstein: Chromatin states in Human • Motif and grammar discovery (Pouya Kheradpour) – White: Motifs associated with insulator proteins – Bernstein: Tissue-specific chromatin states – White: Expression and Binding Time-course • Tissue-specific gene expression (Chris Bristow) – Celniker: Embryo expression domains – All: Predictive models of gene expression Acknowledgements Pouya Kheradpour Alex Stark Mike Lin Jason Ernst Chris Bristow Funding ENCODE, modENCODE, NHGRI, NSF, Sloan Foundation TFs/Insul. 12+8-flies Chromatin Prot.Genes Kevin White, Bing Ren, Nicolas Negre, Par Shah, Jim Posakony Andy Clark, Mike Eisen, Bill Gelbart, Doug Smith, Peter Cherbas Gary Karpen, Aki Minoda, Nicole Riddle, Peter Park + Kharchenko BDGP: Sue Celniker, Jane Landolin, FlyBase: Bill Gelbart