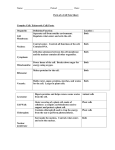
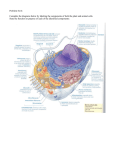
* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Protein translocation pathways across the inner and outer
Model lipid bilayer wikipedia , lookup
Chloroplast DNA wikipedia , lookup
Theories of general anaesthetic action wikipedia , lookup
Protein phosphorylation wikipedia , lookup
Magnesium transporter wikipedia , lookup
G protein–coupled receptor wikipedia , lookup
Protein moonlighting wikipedia , lookup
Cell membrane wikipedia , lookup
SNARE (protein) wikipedia , lookup
Intrinsically disordered proteins wikipedia , lookup
Signal transduction wikipedia , lookup
Protein–protein interaction wikipedia , lookup
Endomembrane system wikipedia , lookup
List of types of proteins wikipedia , lookup
Proteolysis wikipedia , lookup
Indian Journal of Biochemistry & Biophysics Vol. 45, June 2008, pp 149-156 Minireview Protein translocation pathways across the inner and outer mitochondrial membranes Jaspreet Kaur and Gautam Kaul* Department of Animal Biochemistry, National Dairy Research Institute, Karnal, Haryana 132001, India Received 29 June 2007; revised 02 May 2008 Mitochondria import different proteins that are encoded by nuclear genes and synthesized in the cytosol. Separate translocases in inner and outer mitochondrial membranes like TOM, TIM23, TOB/SAM, TIM22 complex facilitate recognition, import and intramitochondrial sorting of preproteins. Various cytosolic factors as Hsp70 and auxillary factors assist in targeting these preproteins to their destinations. Also, different protein components in the matrix participate in this energetically driven translocation process in a reaction that depends upon membrane potential and matrix-ATP. This review summarizes the present knowledge on import and sorting of mitochondrial precursor proteins with glance on unresolved questions. Keywords: Mitochondria, Translocation, Membrane potential, ATP hydrolysis. Introduction Mitochondria are not only of key importance for the bioenergetics of eukaryotic cells, but have critical functions in metabolism of various biomolecules including amino acids and lipids and also play a role in apoptosis1. They are essential for cell viability, proliferation and actively participate in development and differentiation processes. Mitochondria contain about 800 proteins in yeast2 to 1500 different proteins in humans. Although there is a complete genetic system in the matrix, the innermost compartment of mitochondria, only ~1% of all mitochondrial proteins are encoded by organelle genome and synthesized in the matrix2 and rest are encoded by nuclear genes. Basic aspect of this protein sorting is the transfer of nuclearly encoded and cytoplasmically synthesized proteins both across and into the mitochondrial inner and outer membranes. Many hundreds of different mitochondrial proteins participate in these processes. In contrast, only a few protein components are encoded by mitochondrial DNA itself, synthesized on organellar ribosomes and inserted from matrix side into inner membrane3. The translocase of outer mitochondrial membrane, TOM complex represents a common entry gate for __________________________ *Author for correspondence Tel: 0184-2259133, 0184-2259115 Fax: 0184- 2250042 E-mail: [email protected] usually all the nuclear encoded mitochondrial proteins. After making their way through TOM complex, these precursor proteins are directed to atleast four different import pathways: (i) Presequence pathway: imports the matrix proteins using TIM23 complex, (ii) Carrier pathway: imports hydrophobic inner membrane proteins, (iii) TOB/SAM or SAM/TOB pathway: the mitochondrial outer membrane β-barrel proteins are inserted into membrane via TOB/SAM (topogenesis of mitochondrial outer membrane β-barrel proteins also called as sorting and assembly machinery), and (iv) recently identified specific protein import machinery for small intermembrane space proteins4, but little is known about this pathway. Since mitochondria play essential roles in eukaryotic cells, knowledge regarding the protein import and biogenesis of this organelle is equally important. In this review, presequence, carrier, TOB/SAM import pathways as well as import of preproteins in intermembrane space of mitochondria are discussed. Mitochondrial targeting signals The targeting signals or sequences for all matrixtargeted and inner membrane (IM) proteins that contain a single transmembrane anchor, as well as some intermembrane space (IMS) proteins are typically N-terminal extensions of about 15-40 amino acids, rich in basic and hydroxylated amino acids, 150 INDIAN J. BIOCHEM. BIOPHYS., VOL. 45, JUNE 2008 which can form amphipathic α-helices5. However, targeting information is lost in the mature sequences of approximately 30% of mitochondrial-targeted proteins6. Presequence of these precursor proteins interacts directly with the receptors of translocase of outer membrane (Tom20 and Tom22); one half of this amphipathic helix having a hydrophobic surface, is recognized by the binding groove within Tom207, whereas the other half, which is positively-charged, is recognized by the Tom22 of TOM complex. The movement of preproteins through cytosol is associated with cytosolic factors namely chaperones and cytosolic Hsp70, an eukaryotic homologue of bacterial DNaK is one of the chaperones considered as an important factor in intracellular protein traffic. In addition, several other cytosolic factors may also be considered as targeting factors as they bind preproteins in a specific manner and deliver them to the receptor components of TOM complex. Amongst these, the best-characterized ones are mitochondrial import stimulation factor (MSF) and aryl-hydrocarbon receptor interacting protein which interacts with preproteins and Tom207. MSF has shown two activities: (i) it recognizes and forms stable complexes with mature parts of mitochondrial precursors and also facilitates depolymerizaton, and (ii) promotes unfolding of preproteins3. Aggregated preadrenodoxin (involved in electron transfer reaction) synthesized in Escherichia coli8 becomes import competent in presence of MSF. Both Hsp70 and MSF mediate translocation of preproteins from cytosol to TOM complex and involvement of each depends on the nature of preprotein. TOM or TOM40 complex The holo-TOM complex has a size of ~450 kDa and contains seven different proteins viz., the receptors (Tom20 and Tom70) and five additional proteins (Tom40, Tom22, Tom5, Tom6 and Tom7) make up the TOM-core complex which acts as a general insertion pore (GIP ~400 kDa)9-12 (Fig. 1). Tom20 and Tom70 of TOM complex project into cytosol and both have an N-terminal membrane anchor and hydrophilic C-terminal domain of ~17 kDa and ~65 kDa, respectively13. Also, Tom22 extends an N-terminal domain of ~85 amino acid residues into cytosol and has a single transmembrane segment and a smaller C-terminal domain (~45 residues) facing the intermembrane space. Tom40 protein appears to be deeply embedded in the outer membrane of mitochondria, although it does not have many obvious contiguous hydrophobic sequence segments. In recent years, most of the homologous components of TOM complex in mammalian system have also been identified which include: Tom2014, Tom2215, Tom7016, Tom4017 and Tom718. The yeast GIP complex resolves as a ~400 kDa species on native-PAGE, while the mammalian complex is a ~380 kDa species containing at least Tom40, Tom22 and Tom718. TOM complex plays an important role during translocation of preproteins to subsequent translocases in the import pathway19. Role of different proteins of TOM complex Tom20 and Tom70 act as receptors for the precursor proteins20. Tom22 has various roles including receptor functions21, helps in passage of precursor proteins from the receptors to pore22, and binds precursors at trans site23. Tom40 forms poreforming component of the complex and is essential for cell viability. Different mutations generated by altering 10 conserved regions of Neurospora crassa Tom40 protein revealed that all these affect the ability of altered Tom40 to be assembled into TOM complex in isolated mitochondria and such mitochondria have defects in importing preproteins to different subcomparments19. Although the receptors of TOM complex (Tom20 and Tom22) preferentially recognize presequences, Tom70 interacts mainly with hydrophobic precursor proteins carrying internal targeting signals (carrier pathway preproteins). Tom6 plays a major role in the stability of TOM complex25 and is required to promote assembly of Tom40 and Tom22, but not for their interaction during translocation10. Absence of Tom7 results in more stable associations between Tom40, Tom20 and Tom22, suggesting that it plays a role opposite to that of Tom6 with respect to TOM stability10. Tom7 is also involved in the trans site binding and passage of precursors to TIM23 complex25. Mutants of N. crassa Tom proteins lacking both Tom6 and Tom7 have an extremely labile TOM complex and exhibit altered growth phenotype20. Tom5 in Saccharomyces cerevisiae maintains the structure of TOM complex rather than acting in preprotein transfer26, whereas single mutants lacking N. crassa Tom5 display no apparent TOM complex abnormalities and has minor role in maintaining TOM complex stability20. Proteins of mitochondrial inner membrane contain transmembrane α-helices, like proteins present in most other membranes of eukaryotic cell. However, KAUR & KAUL: PROTEIN IMPORT INTO MITOCHONDRIA 151 Fig. 1—A model for protein import into mitochondria [Preproteins are synthesized in the cytosol and are directed to the mitochondrial surface by either N-terminal or internal targeting signals. These precursor proteins after passing through TOM complex are directed to different import pathways: the presequence pathway which involves the TOM complex and TIM23 complex, the carrier pathway requiring TOM complex and TIM22 complex. Translocation of mitochondrial outer membrane complex proteins, β-barrel proteins requires SAM/TOB complex, in addition to TOM complex, whereas proteins with simple topology require only TOM complex. OM, outer membrane; IM, inner membrane. Modified and produced from ref. 5] outer membrane of mitochondria contains proteins anchored in membrane by multiple β-strands9. Recently, it has been revealed that TOM complex is not sufficient to integrate β-barrel proteins into outer membrane and also not able to assemble its own precursor proteins into functional complexes. This led to the discovery of topogenesis of mitochondrial outer membrame β-barrel proteins (TOB) complex, also called the sorting and assembly machinery (SAM) complex (TOB/SAM complex). Mitochondria lacking protein Sam37 (formerly named Tom37) are strongly impaired in Tom40 assembly, the channel protein of outer mitochondrial membrane, whereas other known import pathways, presequence and carrier pathways are not affected27. Since mitochondrial targeting signals are diverse in their nature, it is puzzling that they converge at the same import machinery for outer membrane translocation and subsequent sorting to their appropriate sub-mitochondrial compartments5 by different import routes. TOB/SAM complex The mitochondrial TOB/SAM complex is derived from the bacterial Omp85 complex, but to date no detailed structural analysis of a mitochondrial β-barrel protein has been acheived28. The founding subunit of SAM complex is Sam37/Mas37/Tom3727 (Fig. 1). For membrane integration of mitochondrial β-barrel proteins, Sam37 is required which is a subunit of SAMcore complex (~200 kDa). As compared to the integration of β-barrel proteins, simple outer membrane proteins with α-helical transmembrane segment e.g. Tom20 can be inserted into outer mitochondrial membrane with help of TOM complex alone27. The two additional subunits of SAMcore complex are Sam50/Tob55 and Sam35/Tob38. Sam50 assembles as an integral outer membrane protein with a predicted β-barrel domain29, whereas Sam35 and Sam37 behave as peripheral membrane proteins that are exposed on mitochondrial surface28,30-32. The central subunit of SAM complex is represented by Sam50, which in its purified form forms ring-shaped particles with a large channel31. N-terminal domain of Sam50/Tob55 is exposed to the intermembrane space and represents interaction site for β-barrel precursors before insertion in SAM/TOB complex2,33. Sam35 and Sam37 function codependently in the biogenesis of β-barrel proteins. Sam35 is required for SAM complex to bind outer membrane substrate 152 INDIAN J. BIOCHEM. BIOPHYS., VOL. 45, JUNE 2008 proteins and its destabilization inhibits substrate binding by Sam50, whereas Sam37 seems to assist the release of substrates from SAM comlex28. This SAMcore complex can further associate with a fourth subunit, the morphology component Mdm10 to form SAMholo complex. SAMcore complex is required for biogenesis of all β-barrel proteins of outer membrane ,whereas Mdm10 and SAMholo complex play a selective role in β-barrel biogenesis as they promote assembly of Tom40, but not porin. Tom7, a conserved subunit of TOM complex functions antagonistically to Mdm10 in the biogenesis of Tom40 and porin, suggesting that its role is not limited to TOM complex. Tom7 also functions in the mitochondrial protein biogenesis by a new mechanism, segregation of a sorting component, leading to a differentiation of β-barrel assembly34. In yeast, assembly of Tom40 and porin in outer membrane requires Tom13 in addition to SAMcore complex29. TIM complex or TIM23 complex/translocase TIM23 complex can be structurally and functionally sub-divided into two parts — membrane integrated translocation channel and import motor, which is located at the matrix face of channel35 (Fig. 1). Although membrane integrated part of this translocase is sufficient for insertion of some preproteins into the inner membrane, but complete translocation of preproteins into matrix requires a combination of both subunits. Without motor section, preproteins cannot translocate into matrix. The components of membrane integrated part of TIM23 complex include Tim17, Tim21, Tim23 and Tim50, while import motor of translocase comprises five proteins: Tim14, Tim16, Tim44, Mge1 and mtHsp70 (mitochondrial heat-shock protein 70)35. An integral 23 kDa protein of yeast inner membrane, N-terminal half of Tim23 is hydrophilic in nature, whereas Cterminal half is hydrophobic. Tim17, also an integral membrane protein shares sequence similarity with the hydrophobic portion of Tim233. Tim23 also contains an additional N-terminal domain, not present in Tim17 which has a receptor function and also responds to membrane potential. Role of different proteins of TIM23 complex Tim17 and Tim23 form the translocation channel of TIM23 complex through which precursor proteins usually with an N-terminal cleavable presequence cross the hydrophobic barrier of inner membrane in unfolded state36. Tim21 promotes coupling of the two translocator complexes (TIM23 and TOM) during translocation process. A single transmembrane domain anchors Tim21 in the mitochondrial inner membrane and it exposes its C-terminal domain into intermembrane space. The C-terminal domain of Tim21 does not to bind to any of the TIM23 components but specifically interacts with TOM complex. It has been proposed that Tim21 binds to trans site of TOM complex, thus keeping two translocases in close contact37. Tim50 facilitates protein transfer from TOM40 complex to TIM23 comlex36. Mitochondrial Hsp70 (mtHsp70) in matrix functions as import motor to drive vectorial translocation and unfolding of precursor proteins in cooperation with its partner proteins, mitochondrial associated motor and chaperone (MMC) proteins. Tim44 provides anchor for this motor (mtHsp70), so that it can bind to the polypeptide emerging out from TIM23 channel. The cochaperone Mge1 is nucleotide exchange factor for mtHsp70 and enhances its ATPase activity38. A new member of this translocator system Tam41 (translocation assembly and maintenance 41) has been identified from the yeast is a peripheral inner mitochondrial membrane protein facing matrix and not a constituent of TIM23 complex. It does not promote motor function of mtHsp70 but is instead involved in the maintenance of TIM23 complex integrity36. Roles of Tim14 and Tim16 are discussed later. Insertion of preproteins into TIM23 channel depends mainly on the presence of membrane potential across inner membrane which plays a dual role in import process. Firstly, it activates the channel protein itself and secondly it exerts an electrophoretic effect on positively-charged presequence, thereby driving presequence to matrix side7. Although, the exact mechanism of import motor is still not understood, it is clear that several rounds of ATPdependent binding to and release from the mtHsp70 lead to complete translocation of polypeptides into matrix of mitochondria. The components of TIM23 translocase are highly conserved throughout eukaryotic kingdom35. Pathways of preprotein translocation across and into the outer membrane After translocation through β-barrel protein the Tom40 pore, presequence binds to intermembrane space domain of receptor Tom223. Tom40 itself KAUR & KAUL: PROTEIN IMPORT INTO MITOCHONDRIA interacts with preproteins in transit. Preproteins which follow the presequence pathway have a cleavable presequence and are guided across outer membrane by a chain of binding sites, including the cytosolic receptors Tom20, Tom22, Tom5 and Tom40 which form translocation channel and intermembrane space domain of Tom227. Other small proteins Tom6 and Tom7 are required for assembly and stability of TOM complex (Fig. 1), but they do not interact with the precursor proteins21. Tom70 recognizes the precursors of hydrophobic carrier proteins of inner membrane such as the ADP/ATP carrier. These precursors possess multiple internal targeting sequences but lack presequence. The cytosolic chaperones, particularly Hsp70 and Hsp90 classes bind these hydrophobic precursors and prevent their aggregation in the cytosol. Heat-shock proteins then specifically interact with Tom70 and deliver the carrier precursors to this receptor. These heat-shock proteins are essential in mitochondrial biogenesis, as revealed in Caenorhabditis elegans which develops early aging or progeria-like phenotypes, if HSP6 (a nematode orthologue of mtHsp70) is reduced by RNA interference39. Several Tom70 molecules simultaneously bind to one precursor molecule which is subsequently transferred to the import pore, Tom407. The effective internal diameter of protein import channel in mitochondrial outer membrane appears to be between 20-26 Å during translocation40. β-Barrel proteins represent a distinct class of mitochondrial outer membrane proteins41, but little is known about how these newly synthesized proteins sort in eukaryotic cell, integrate into lipid bilayers and assemble into oligomeric structures33. Like other outer membrane proteins, β-barrel proteins such as porin and Tom40 first translocate across TOM complex and in a manner dependent on chaperones in intermembrane space, pass on to TOB/SAM complex for their insertion into outer membrane28,33. In the intermembrane space, small TIM complexes, essential Tim9-Tim10 complex and non-essential Tim8-Tim13 complex promote transfer of β-barrel precursors to TOB/SAM complex. These small proteins have chaperone-like functions which probably shield hydrophobic segments of the precursor proteins against aggregation in aqueous intermembrane space, keeping β-barrel precursors in a competent state for their insertion. Biogenesis of Tom40, pore-forming channel of outer mitochondrial membrane involves the 153 movement of precursors in unfolded state to SAMcore complex to form an assembly intermediate I (~250 kDa) which further assembles into mature TOM complex by passing through assembly intermediate II (~100 kDa)42. How this assembly intermediate II separates from SAMcore complex is under investigation. Mdm10 also associates with SAM to form a larger complex of ~350 kDa and plays a specific role in assembly of TOM complex, probably by functioning as a scaffold, where the late steps of assembly of the multi-subunits TOM complex takes place2. Transfer of preproteins from TOM to TIM 23: Translocation of preproteins across the inner mitochondrial membrane Majority of presequence carrying proteins are imported into the matrix by cooperation of presequence translocase of inner membrane, TIM23 complex and the associated motor (presequence associated motor, PAM). Protein import into matrix requires two forms of energy – an electrochemical gradient (positive outside) across inner membrane which exerts an electrophoretic force on the positively charged N-terminal targeting sequence43 and hydrolysis of ATP in matrix44. Ability of generating strong translocating driving force solely depends upon ATP-driven import motor and membrane potentialincreases the interaction of preprotein with motor41. Precursor proteins are imported with help of TOM complex and TIM23 complex into matrix (Fig. 1). This ATP-driven import motor consists of five proteins, three essential proteins–mtHsp70, peripheral inner membrane protein Tim44 and a nucleotide exchange factor of matrix termed mitochondrial GrpE (Mge1)38 and two cochaperones Pam18/Tim14 and Pam16/Tim16. Molecular mechanism of the import motor involves the following steps: (i) mtHsp70 with bound ATP interacts with Tim44 and preproteins, but with low affinity (ii) Pam18/Tim14 stimulates ATP hydrolysis which in turn leads to the ADP-bound form of mtHsp70, stabilizing its interaction with Tim44 and preproteins. (iii) Mge1, a soluble matrix protein then promotes the release of ADP-from mtHsp70. PAM, thus operates as a multi-step motor, but the exact order of events during reaction cycle of mtHsp 70 is not fully understood7. TOM and TIM23 complexes do not appear to form stable super complex in the absence of precursor proteins35. TIM23 channel is also tightly regulated, so as to maintain the permeability barrier of inner 154 INDIAN J. BIOCHEM. BIOPHYS., VOL. 45, JUNE 2008 mitochondrial membrane. In absence of precursor proteins, it remains in its inactive state, but regains its active state, when precursor proteins are present and opens up for translocation of polypeptide chains. Both Tim50 and presequence act in an antagonistic manner in this process. Whereas the hydrophilic IMS domain of Tim50 promotes oligomerization and voltagedependent closure of channel, presequences selectively override Tim50-induced closure and activate the channel, thereby crossing the channel45. This process is very complicated under in vivo conditions and there is no clear-cut answer to the question as how energy sources of mitochondria, membrane potential and matrix ATP cooperate to drive preprotein import. A new model for the mechanism of import motor called “entropic pulling” has been proposed that postulates generation of an active import-driving force based on thermodynamic principles and molecular geometries during translocation. According to this model, translocation force is generated by the excluded-volume constraint between mtHsp70 and membrane and requires efficient interaction of mtHsp70 and Tim44 for efficient translocation and unfolding of proteins46. Protein insertion machinery of the inner membrane (TIM22 complex) Substrate proteins for TIM22-mediated translocation contain multiple hydrophobic segments that can be inserted into inner membrane. Import of ADP-ATP carrier (AAC) protein of inner mitochondrial membrane has been characterized47. This 300 kDa complex consists of a pore-forming subunit Tim22 along with Tim18 and Tim54 that play unknown function and small Tim proteins Tim9, Tim10 and Tim1248. The general import of inner membrane proteins is summarized here (Fig. 1). After recognition by Tom70 and translocation by TOM pore across outer membrane as a loop, these precursor proteins are bound by Tim9Tim10 (70 kDa) complex49. Non-essential Tim proteins (Tim8-Tim13 complex) also support the transfer of selected inner membrane proteins, whereas Tim9– Tim10 complex docks precursor proteins to the TIM22 complex. These Tim proteins contain the “twin CX3C” motif that is required for formation of 70 kDa complex50. Function of this motif is not fully known, but it seems to be important for Zn2+ coordination in these small Tim proteins. It has also been suggested that cysteine present in this motif helps to form juxtapositional intramolecular disulfide bonds51. These proteins can bind zinc but insertion of intramolecular disulfide bonds is requisite for assembly of complex52. An intermembrane space protein Hot13 is also required for efficient assembly of these Tim proteins into TIM complexes. Thus, small TIM complexes have a specialized assembly and the local redox state of these complexes mediate translocation of inner membrane proteins51. Precursor is then translocated to the core of insertion machinery, channel forming protein Tim2253 which also shares homology with Tim23. Indeed, like presequence translocase, Tim22 channel is also activated by membrane potential, but no ATP-driven machinery is required in this process. Membrane potential is sufficient for both initial insertion of precursor polypeptide into translocase and for completion of transport i.e. the lateral release of protein into inner membrane7. Tanslocation of proteins to intermembrane space (IMS) Precursors destined for intermembrane space as cytochromes b2 and c1 contain a bipartite targeting sequence and N-terminal directs precursors to TIM23 complex. After cleavage by matrix processing peptidase (MPP), precursor arrests at the “stop transfer” domain in translocon54. The inner membrane protease (IMP) consisting of complex Imp1/Imp2 and facing intermembrane space mediates a second cleavage, thereby arresting cytochromeb b2 in intermembrane space55. This translocation also requires hydrolysis of matrix ATP and electrochemical membrane potential ∆ψ. For proteins lacking Nterminal and not requiring electrochemical membrane potential for their translocation thereby bypassing TIM23, their import mechanisms depends upon – energy gain of protein around the cofactor and association of a protein with high affinity binding site such as another protein56. Polynucleotide phosphorylase (PNPase), which is an exoribonuclease as well as poly(A) polymerase that regulates RNA levels is also localized in the intermembrane space and not in cytosol or matrix as considered earlier57 . Import of PNPase depends on ∆ψ and MPP cleavage of presequence. Translocation of PNPase into intermembrane requires i-AAA proteaseYme1 which has a chaperone-like activity56. This new Yme1 in protein translocation gives an idea of the diverse protein translocation pathways of intermembrane space which needs investigation. KAUR & KAUL: PROTEIN IMPORT INTO MITOCHONDRIA Conclusions and Perspective During recent years, important insights have been revealed into the machineries involved in targeting and sorting of proteins to mitochondria. However, inspite of rapid progress in our understanding of the mechanisms of protein import into mitochondria, many questions still remain to be answered. For example, general import pathways have been revealed on the basis of detailed analysis of import of various proteins, which depends upon one or two of translocators TOM, TIM23 and TIM22 complexes during transloction. But there may be more specific pathways for several mitochondrial proteins, including those synthesized inside mitochondria that remain to be elucidated. More than 30 proteins have been identified as translocator components, indicating that pathways of import and sorting are much more complex than previously envisaged. Identification and characterization of SAM/TOB pathway led to several surprising twists in the analysis of organelle biogenesis. Important areas for future studies will be: the characterization of internal targeting and sorting signals of precursor proteins, structural analysis of membrane integrated transport complexes and import machinery mechanism. 10 11 12 13 14 15 16 17 References 1 2 3 4 5 6 7 8 9 Sickmann A, Reinders J, Wagner Y, Joppich C, Zahedi R, Meyer H E, Sconfisch B, Perschil I, Chacinska A, Guiard B, Rehling P, Pfanner N & Meisinger C (2003) The proteome of Saccharomyces cerevisiae mitochondria. Proc Natl Acad Sci (USA) 100, 13207-13212 Pfanner N, Wiedemann N, Meisinger C & Lithgow T (2004) Assembling the mitochondrial outer membrane. Nat Struct Mol Biol 11, 1044-1048 Neupert W (1997) Protein import into mitochondria. Ann Rev Biochem 66, 863-917 Rissler M, Wiedemann N, Pfannschmidt S, Gabriel K, Guiard B, Pfanner N & Chacinska A (2005) The essential mitochondrial protein Erv1 cooperates with Mia40 in biogenesis of intermembrane space proteins. J Mol Biol 353, 485-492 Stojanovski D, Johnston A J, Streimann I, Hoogenraad N J & Ryan M T (2003) Import of nuclear-encoded proteins into mitochondria. Exp Physiol 88, 57-64 Diekert K, Kispal G, Guiard B & Lill R (1999) An internal targeting signal directing proteins into the mitochondrial intermembrane space. Proc Natl Acad Sci (USA) 96, 11752-11757 Wiedemann N, Frazier A E & Pfanner N (2004) The Protein Import Machinery of Mitochondria. J Biol Chem, 279, 14473-14476 Iwahashi J, Furuya S, Mihara K & Omura T (1992) Characterization of Adrenodoxin precursor expressed in Escherichia coli. J Biochem 111, 451-455 Matouschek A & Glick B S (2001) Barreling through the outer membrane. Nat Struct Biol 8, 284-286 18 19 20 21 22 23 24 155 Dekker P J T, Ryan M T, Brix J, Muller H, Honlinger A & Pfanner N (1998) Preprotein translocase of the outer mitochondrial membrane: molecular dissection and assembly of the general import pore complex. Mol Cell Biol 18, 6515-6524 Dekker P J, Muller H, Rassow J & Pfanner N (1996) Characterization of the preprotein translocase of the outer mitochondrial membrane by blue native electrophoresis. Biol Chem 377, 535-538 Kunkele K P, Heins S, Dembowski M, Nargang F E, Benz R, Thieffry M, Walz J, Lill R, Nussberger S & Neupert W (1998) The preprotein translocation channel of the outer membrane of mitochondria. Cell 93, 1009-1019 Schlossmann J, Lill R, Neupert W & Court D A (1996) Tom71, a novel homologue of the mitochondrial preprotein receptor Tom70. J Biol Chem 271, 17890-17895 Hanson B, Nuttal S & Hoogenraad N (1996) A receptor for the import of proteins into human mitochondria. Eur J Biochem 235, 750-753 Yano M, Hoogenraad N, Terada K & Mori M (2000) Identification and functional analysis of human Tom22 for protein import into mitochondria. Mol Cell Biol 20, 7205-7213 Suzuki H, Maeda M & Mihara K (2002) Characterization of rat TOM70 as a receptor of the preprotein translocase of the mitochondrial outer membrane. J Cell Sci 115, 1895-1905 Suzuki H, Okazawa Y, Komiya T, Saeki K, Mekada E, Kitada S, Ito A & Mihara K (2000) Characterization of rat TOM40, a central component of the preprotein translocase of the mitochondrial outer membrane. J Biol Chem 275, 37930-37936 Johnston A J, Hoogenraad J, Dougan D A, Truscott K N, Yano M, Mori M, Hoogenraad N J & Ryan M T (2002) Insertion and assembly of human Tom7 into the preprotein translocase complex of the outer mitochondrial membrane. J Biol Chem 277, 42197-42204 Sherman E L, Taylor R D, Go N E & Nargang F E (2006) Effect of Mutations in Tom40 on stability of the translocase of the Outer Mitochondrial Membrane (TOM) Complex, Assembly of Tom40, and Import of Mitochondrial Preproteins. J Biol Chem 281, 22554-22565 Sherman E L, Go N E & Nargang F E (2005) Functions of the Small Proteins in the TOM Complex of Neurospora crassa. Mol Biol Cell 16, 4172-4182 Mayer A, Nargang F E, Neupert W & Lill R (1995) MOM22 is a receptor for mitochondrial targeting sequences and cooperates with MOM19. EMBO J 14, 4204-4211 Kiebler M, Keil P, Schneider H, van der Klei I, Pfanner N & Neupert W (1993) The mitochondrial receptor complex: a central role of MOM22 in mediating transfer of preproteins from receptors to the general insertion pore. Cell 74, 483-492 Nakai M, Kinoshita K & Endo T (1995) Mitochondrial receptor complex protein. The intermembrane space domain of yeast Mas17 is not essential for its targeting or function. J Biol Chem 270, 30571-30575 Dietmeier K, Honlinger A, Bomer U, Dekker P J T, Eckerskorn C, Lottspeich F, Kubrich M & Pfanner N (1997) Tom5 functionally links mitochondrial preprotein receptors to the general import pore. Nature 388, 195-200 156 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 INDIAN J. BIOCHEM. BIOPHYS., VOL. 45, JUNE 2008 Esaki M, Shimizu H, Ono T, Yamamoto H, Kanamori T, Nishikawa S & Endo T (2004) Mitochondrial protein import. Requirement of presequence elements and Tom components for precursor binding of the TOM complex. J Biol Chem 279, 45701-45707 Schmitt S, Ahting U, Eichacker L, Granvogl B, Go N E, Nargang F E, Neupert W & Nussberger S (2005) Role of Tom5 in maintaining the structural stability of the TOM complex of mitochondria. J Biol Chem 280, 14499-14506 Wiedemann N, Kozjak V, Chacinska A, Schonfisch B, Rospert S, Ryan M T, Pfanner N & Meisinger C (2003) Machinery for protein sorting and assembly in the mitochondrial outer membrane. Nature 424, 565-571 Chan N C & Lithgow T (2008) The peripheral membrane subunits of the SAM complex function codependently in the mitochondrial outer membrane biogenesis. Mol Biol Cell 19, 126-136 Ishikawa D, Yamamoto H, Tamura Y, Moritoh K & Endo T (2004) Two novel proteins in the mitochondrial outer membrane mediate β-barrel protein assembly. J Cell Biol 166, 621-627 Kozjak V, Wiedemann N, Milenkovic D, Lohaus C, Meyer H E, Guiard B, Meisinger C & Pfanner N (2003) An essential role of Sam50 in the protein sorting and assembly machinery of the mitochondrial outer membrane. J Biol Chem 278, 48520-48523 Paschen S A, Waizenegger T, Stan T, Preuss M, Cyrklaff M, Hell K, Rapaport D & Neupert W (2003) Evolutionary conservation of biogenesis of β-barrel membrane proteins. Nature 426, 862-866 Milenkovic D, Kozjak V, Wiedemann N, Lohaus C, Meyer, H E, Guiard B, Pfanner N & Meisinger C (2004) Sam35 of the mitochondrial protein sorting and assembly machinery is a peripheral outer membrane protein essential for cell viability. J Biol Chem 279, 22781-22785 Habib S J, Waizenegger T, Niewienda A, Paschen S A, Neupert W & Rapaport D (2007) The N-terminal domain of Tob55 has a receptor-like function in the biogenesis of mitochondrial β-barrel proteins. J Cell Biol 176, 77-88 Meisinger C, Wiedemann N, Rissler M, Strub A, Milenkovic D, Schonfisch B, Muller H, Kozjac V & Pfanner N (2006) Mitochondrial protein sorting differentiation of β-barrel assembly by Iom7-mediated segregation of Mdm10. J Biol Chem 281, 22819-22826 Mokranjac D & Neupert W (2005) Protein import into mitochondria. Biochem Soc Trans 33, 1019-1023 Tamura Y, Harada Y, Yamano K, Watanabe K, Ishikawa D, Ohshima C, Nishikawa S-i, Yamamoto H & Endo T (2006) Identification of Tam41 maintaining integrity of the TIM23 protein translocator complex in mitochondria. J Cell Biol 174, 631-637 Mokranjac D, Popov-Celeketic D, Hell K & Neupert W (2005) Role of Tim21 in Mitochondrial Translocation Contact Sites. J Biol Chem 280, 23437-23440 Liu Q, D’Silva P, Walter W, Marszalek J & Craig E A (2003) Regulated cycling of mitochondrial Hsp70 at the protein import channel. Science 300, 139-141 Kimura K, Tanaka N, Nakamura N, Takano S & Ohkuma S (2007) Knockdown of mitochondrial heat shock protein 70 promotes Progeria-like phenotypes in Caenorhabditis elegans. J Biol Chem 282, 5910-5918 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 Schwartz M P & Matouschek A (1999) The dimensions of the protein import channels in the outer and inner mitochondrial membranes. Proc Natl Acad Sci (USA) 96, 13086-13090 Krayl M, Lim J H, Martin F, Guiard B and Voos W (2007) A cooperative action of the ATP-Dependent Import Motor Complex and the inner membrane potential drives mitochondrial preprotein import. Mol Cell Biol 27, 411-425 Taylor R, McHale B & Nargang F E (2003) Characterization of Neurospora crassa Tom40-deficient mutants and effect of specific mutations on Tom40 assembly. J Biol Chem 278, 765-775 Martin J, Mahlke K & Pfanner N (1991) Role of energized inner membrane in mitochondrial protein import. Delta psi drives the movement of presequences. J Biol Chem 266, 18051-18057 Schatz G (1993) The protein import machinery of mitochondria. Protein Sci 2, 141-146 Meinecke M, Wagner R, Kovermann P, Guiard B, Mick D U, Hutu D P, Voos W, Truscott K N, Chacinska A, Pfanner N & Rehling P (2006) Tim50 Maintains the Permeability Barrier of the Mitochondrial Inner Membrane. Science 312, 1523-1526 Rios P D L, Ben-zvi A, Slutsky O, Azem A & Goloubinoff P (2006) Hsp70 chaperones accelerate protein translocation and the unfolding of stable protein aggregates by entropic pulling. Proc Natl Acad Sci (USA) 103, 6166-6171 Endo T, Yamamoto H & Esaki M (2003) Functional cooperation and separation of translocators in protein import into mitochondria, the double-membrane bounded organelles. J Cell Sci 116, 3259-3267 Pfanner N & Giessler A (2001) Versatility of the mitochondrial protein import machinery. Nat Rev Mol Cell Biol 2, 339-349 Vial S, Lu H, Allen S, Savory P, Thornton D, Sheehan J & Tokatlidis K (2002) Assembly of Tim9 and Tim10 into a functional chaperone. J Biol Chem 277, 36100-36108 Koehler C M (2004) The small Tim proteins and the twin Cx3C motif. Trends Biochem Sci 29, 1-4 Curran S P, Leuenberger D, Leverich E P, Hwang D K, Beverly K N & Koehler C M (2004) The role of Hot13p and redox chemistry in the mitochondrial TIM22 import pathway. J Biol Chem 279, 43744-43751 Lu H, Allen S, Wardleworth L, Savory P & Tokatlidis K (2004) Functional TIM10 chaperone assembly is redox-regulated in vivo. J Biol Chem 279, 18952-18958 Sirrenberg C, Bauer M F, Guiard B, Neupert W & Brunner M (1996) Import of carrier proteins into the mitochondrial inner membrane mediated by Tim22. Nature 384, 582-585 Glick B S, Brandt A, Cunningham K, Muller S, Hallberg R L & Schatz G (1992) Cytochromes c1 and b2 are sorted to the intermembrane space of yeast mitochondria by a stop-transfer mechanism. Cell 69, 809-822 Gakh O, Cavadini P, Isaya G (2002) Mitochondrial processing peptidases. Biochim Biophys Acta 1592, 63-77 Rainey R N, Glavin J D, Chen H-W, French S W, Teitell M A & Koehler C M (2006) A new function in translocation for the mitochondrial i-AAA protease Yme1: import of Polynucleotide Phosphorylase into the intermembrane space. Mol Cell Biol 26, 8488-8497 Nagaike T, Suzuki T, Katoh T & Ueda T (2005) Human mitochondrial mRNAs are stabilized with polyadenylation regulated by mitochondria-specific poly(A) polymerase and polynucleotide phosphorylase. J Biol Chem 280, 19721-19727