* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download 1. The I gene determines the synthesis of a repressor molecule
Gene desert wikipedia , lookup
Transcription factor wikipedia , lookup
Saethre–Chotzen syndrome wikipedia , lookup
DNA supercoil wikipedia , lookup
Behavioral epigenetics wikipedia , lookup
Biology and consumer behaviour wikipedia , lookup
Minimal genome wikipedia , lookup
Dominance (genetics) wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Ridge (biology) wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Frameshift mutation wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Quantitative trait locus wikipedia , lookup
Transgenerational epigenetic inheritance wikipedia , lookup
Epigenetics wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenomics wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Oncogenomics wikipedia , lookup
Genome evolution wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Gene expression profiling wikipedia , lookup
X-inactivation wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genome (book) wikipedia , lookup
History of genetic engineering wikipedia , lookup
Primary transcript wikipedia , lookup
Designer baby wikipedia , lookup
Helitron (biology) wikipedia , lookup
Genomic imprinting wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Point mutation wikipedia , lookup
Microevolution wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
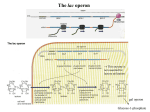
1. The I gene determines the synthesis of a repressor molecule, which blocks expression of the lac operon and which is inactivated by the inducer. The presence of the repressor I+ will be dominant to the absence of a repressor I–. Is mutants are unresponsive to an inducer. For this reason, the gene product cannot be stopped from interacting with the operator and blocking the lac operon. Therefore, Is is dominant to I+. 2. Oc mutants are changes in the DNA sequence of the operator that impair the binding of the lac repressor. Therefore, the lac operon associated with the Oc operator cannot be turned off. Because an operator controls only the genes on the same DNA strand, it is cis (on the same strand) and dominant (cannot be turned off). 3. a. You are told that a, b, and c represent lacI, lacO, and lacZ, but you do not know which is which. Both a– and c– have constitutive phenotypes (lines 1 and 2) and therefore must represent mutations in either the operator (lacO) or the repressor (lac I). b– (line 3) shows no ß-gal activity and by elimination must represent the lacZ gene. Mutations in the operator will be cis-dominant and will cause constitutive expression of the lacZ gene only if it’s on the same chromosome. Line 6 has c– on the same chromosome as b+ but the phenotype is still inducible (owing to c+ in trans). Line 7 has a– on the same chromosome as b+ and is constitutive even though the other chromosome is a+ . Therefore a is lacO, c is lacI, and b is lacZ. b. 4. Another way of labeling mutants of the operator is to denote that they lead to a constitutive phenotype; lacO– (or a–) can also be written as lacOc. There are also mutations of the repressor that fail to bind inducer (allolactose) as opposed to fail to bind DNA. These two classes have quite different phenotypes and are distinguished by lacIs (fails to bind allolactose and leads to a dominant uninducible phenotype in the presence of a wild-type operator) and lacI– (fails to bind DNA and is recessive). It is possible that line 3, line 4, and line 7 have lacIs mutations (because dominance cannot be ascertained in a cell that is also lacOc) but the other c– alleles must be lacI–. ß-Galactosidase Part a b c d e f g No lactose + + – – + + – Lactose + + – – + + + Permease No lactose – – – – + – – Lactose + – – – + – + Chapter Ten 171 a. The Oc mutation leads to the constitutive synthesis of ß-galactosidase because it is cis to a lacZ+ gene, but the permease is inducible because the lacY+ gene is cis to a wild-type operator. b. The lacP– mutation prevents transcription so only the genes cis to lacP+ will be transcribed. These genes are also cis to Oc so the lacZ+ gene is transcribed constitutively. c. The lacIs is a trans-dominant mutation and prevents transcription from either operon. d. Same as part c. e. There is no functional repressor made (and one operator is mutant as well). f. Same as part b. g. Both operators are wild type and the one functional copy of lacI will direct the synthesis of enough repressor to control both operons. 9. The term epigenetic inheritance is used to describe heritable alterations in which the DNA sequence itself is not changed. Paramutation and parental imprinting are two such examples. 11. Imprinted genes are functionally hemizygous. Maternally imprinted genes are inactive when inherited from the mother, and paternally imprinted genes are inactive when inherited from the father. A mutation in one of these genes is dominant when an offspring inherits a mutant allele from one parent and a “normal” but inactivated allele from the other parent. 13. The inheritance of chromatin structure is thought to be responsible for the inheritance of epigenetic information. This is due to the inheritance of the histone code and may also include inheritance of DNA methylation patterns. 14. Many DNA-protein interactions are shared by prokaryotes and eukaryotes, but the mechanisms by which proteins bound to DNA at great distances from the start of transcription affect that transcription is unique to eukaryotes. Also, mechanisms of gene regulation based on chromatin structure are distinctly eukaryotic. 29. a. D through J — the primary transcript will include all exons and introns. b. E, G, I — all introns will be removed. c. A, C, L — the promoter and enhancer regions will bind various transcription factors that may interact with RNA polymerase.