* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download Nanoscale Architecture of Endoplasmic Reticulum Export Sites and
Survey
Document related concepts
Cell nucleus wikipedia , lookup
Chemical synapse wikipedia , lookup
Tissue engineering wikipedia , lookup
Cytoplasmic streaming wikipedia , lookup
Model lipid bilayer wikipedia , lookup
SNARE (protein) wikipedia , lookup
Cell growth wikipedia , lookup
Cell culture wikipedia , lookup
Cellular differentiation wikipedia , lookup
Cell encapsulation wikipedia , lookup
Signal transduction wikipedia , lookup
Extracellular matrix wikipedia , lookup
Organ-on-a-chip wikipedia , lookup
Cell membrane wikipedia , lookup
Cytokinesis wikipedia , lookup
Transcript
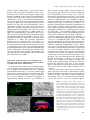
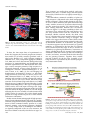
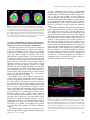
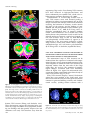
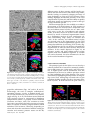
Update on Electron Tomography Analysis of Plant Golgi Stacks Nanoscale Architecture of Endoplasmic Reticulum Export Sites and of Golgi Membranes as Determined by Electron Tomography1[W] L. Andrew Staehelin and Byung-Ho Kang* Molecular Cellular and Developmental Biology, University of Colorado, Boulder, Colorado 80309 (L.A.S.); and Microbiology and Cell Science Department and Integrated Center for Biotechnology Research, University of Florida, Gainesville, Florida 32611 (B.-H.K.) Modern architecture is guided by the axiom ‘‘form follows function,’’ which emphasizes the need for the shape of a building or an object to reflect its intended function or purpose. Biologists tend to prefer the phrase ‘‘form begets function,’’ because in living organisms, form not only reflects on function but also defines many functional attributes. This relationship between biological structure and function explains why one of the central goals of 21st century cell biologists is to provide a seamless link of structural understanding between the macroscopic level of tissue organization to the molecular and even atomic level organization of the building blocks of cells and tissues. In turn, this structural knowledge provides architectural constraints for developing testable hypotheses for how macromolecular complexes, organelles, cells, and organs operate on a functional level. The goal of this Update is to summarize new information on the three-dimensional (3D) architecture of the membrane systems of the secretory pathway of plant cells produced by dual-axis electron tomography of cells preserved by high-pressure freezing/freeze-substitution techniques. These studies have redefined our understanding of the nanoscale organization of the membranes of endoplasmic reticulum (ER) export sites, Golgi stacks, trans-Golgi network (TGN) cisternae, and associated vesicles and scaffolds, and by doing so have led to many new insights into the functional organization of these membrane systems. In addition, these studies have created a bridge between live cell imaging by confocal microscopy on one hand and biochemical and molecular investigations on the other. 1 This work was supported by the National Institutes of Health (grant no. GM–61306 to L.A.S.). * Corresponding author; e-mail [email protected]. The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantphysiol.org) is: Byung-Ho Kang ([email protected]). [W] The online version of this article contains Web-only data. www.plantphysiol.org/cgi/doi/10.1104/pp.108.120618 1454 HIGH-PRESSURE FREEZING AND LOW-TEMPERATURE PROCESSING METHODS ARE ESSENTIAL TOOLS FOR ELECTRON TOMOGRAPHY STUDIES OF CELLS The quality of a final electron tomogram is determined primarily by the quality of the initial specimen preservation. The membranous organelles of the secretory pathway are highly dynamic cellular structures that undergo changes in organization in the time range of seconds and even fractions of seconds. Thus, to preserve these membrane systems in their natural state for electron microscope (EM) analysis requires fixation methods that can stabilize cellular structures in a fraction of a second and processing techniques that can preserve the stabilized structures until they have been immobilized in polymerized resin. Chemical fixatives such as glutaraldehyde and osmium tetroxide preserve cellular structures by crosslinking cellular molecules, thereby preserving their spatial organization. However, because these chemical reactions are relatively slow and highly selective, immobilization of membranes typically takes minutes, and even then not all molecules are fixed (Buckley, 1973; Mersey and McCully, 1978). In practical terms, considering that plant Golgi stacks can travel at speeds of 3 to 4 mm/s along actin filaments (Boevink et al., 1998; Nebenführ et al., 1999) and that chemical fixatives cross-link and thereby immobilize different cellular structures at different rates, it is easy to imagine how a given Golgi stack that was close to an ER export site at the onset of fixation can be separated from this site by microns of distance by the time the fixation process is completed. Because additional changes in membrane architecture occur during chemical dehydration prior to embedding, it is obvious that the analysis of chemically fixed samples by electron tomography will yield information that is of limited value in terms of defining the macromolecular architecture of dynamic organelles. These limitations of chemical fixatives can be largely overcome with the use of cryofixation techniques, among which high-pressure freezing is the most widely used (Kiss and Staehelin, 1995). Unlike chemical fixatives, cryofixation methods immobilize all cellular molecules Plant Physiology, August 2008, Vol. 147, pp. 1454–1468, www.plantphysiol.org Ó 2008 American Society of Plant Biologists Electron Tomography Analysis of Plant Golgi Stacks (proteins, lipids, carbohydrates, water, salts) simultaneously, and this physical stabilization occurs in approximately 1 ms, which is fast enough to capture most transient membrane events of interest to cell biologists. Samples preserved in this manner can then be freezesubstituted at 280°C to 290°C prior to being infiltrated in resin and fixed in 3D space by resin polymerization (Morphew, 2007). Due to the tight binding of water molecules to plant cell wall polysaccharides, freezesubstitution of plant cells takes longer than that of animal cells. Similarly, optimal preservation of freezesubstituted plant cells requires very slow rates of resin infiltration (see Segui-Simarro et al., 2004; Austin et al., 2005). For immunolabeling experiments, the best results are achieved when the freeze-substituted samples are infiltrated with Lowicryl HM20 at 250°C to 260°C and polymerized by UV light at the same temperature (Donohoe et al., 2007). This specimen preparation method preserves antigenic sites very well as evidenced by the fact that more than 10 antibodies we have tested recently (commercial monoclonal, as well as anti-peptide, anti-protein, and anti-polysaccharide polyclonal antibodies) gave excellent immunolabeling results. Furthermore, considering the excellent structural preservation of the samples, this method is far superior to the Tokyasu technique, which is still widely used (Zeuschner et al., 2006). ELECTRON TOMOGRAPHY CAN GENERATE 3D IMAGES OF PLASTIC EMBEDDED SAMPLES WITH NANOMETER-SCALE RESOLUTION To comprehend the differences in imaging capabilities of different microscope techniques, it is instructive to compare the resolution that can be obtained with different microscopes. At present, confocal light microscopy is the most widely used microscopic technique in cell biology laboratories (Fig. 1A). Typically, the x/y axis resolution of confocal micrographs is approximately 200 nm, and the z axis resolution is 500 to 800 nm (Jakobs, 2006). Classical electron microscopy (Fig. 1B), which is based on the imaging of 60- to 80-nm-thick plastic sections, has an x/y axis resolution of approximately 5 nm and a z axis resolution of 120 to 160 nm (approximately twice the section thickness). Although this z axis resolution is approximately 4-fold better than what can be obtained with confocal microscopy, it still limits the ability of researchers to determine the complete 3D architecture of most cellular organelles and cytoskeletal systems at high resolution. As discussed below, dual-axis electron tomography (McIntosh et al., 2005) provides a means for solving the z axis resolution problem of classical electron microscopy as well as for producing 3D models of cellular structures with nanoscale resolution. Electron tomography generates 3D structures of objects by combining multiple 2D images of the objects as they are systematically tilted from 160° to 260° along two orthogonal axes. The desired 3D structure is computed by back-projecting each 2D image with appropriate weighting (Supplemental Fig. S1; Koster et al., 1997). The tomogram produced in this manner constitutes a 3D block of data points that is represented as an array of volume elements (voxels). Each voxel corresponds to a cube (1–4 nm per side depending on resolution), and each cube is assigned a gray scale that corresponds to the mass density of that region of the specimen. The voxel information can be displayed in the form of 2D images of selected z axis levels of the sample. The resulting voxel images resemble thin section images (compare Figs. 1C and 2A). However, because the displayed voxel layer is only approximately 2 nm thick versus 60 to 80 nm for thin sections, many more details of the cellular structures can be resolved. The data presented in this review are from tomograms generated from 100- to 300-nm-thick plastic sections of high-pressure-frozen/freeze-substituted cells examined in 200- and 300-kV Tecnai intermediate voltage EMs. The z axis resolution of the tomograms is 6 to 8 nm and the x and y axis resolution approxFigure 1. Comparison of the spatial resolution of a confocal and an EM image of plant Golgi stacks (A and B). A, Tobacco BY-2 cell expressing a a-1,2 mannosidase I-GFP fusion protein that localizes to Golgi stacks. In this confocal micrograph, the individual Golgi stacks are seen as approximately 1 mm in diameter fluorescent spots. B, Thin section electron micrograph of a Golgi stack in a high-pressure-frozen and freeze-substituted BY-2 cell. Note the differences in architecture and staining of the cis-, medial-, and trans-Golgi cisternae. Ribosomes (approximately 15-nm black dots) and an ER cisterna adjacent to the Golgi stack are also seen (courtesy of A. Nebenführ). C and D, Enhancement of electron microscopy by tomography techniques. Two-nanometer-thick electron tomographic slice image (C) and a 3D reconstruction (D) of an Arabidopsis Golgi stack are shown. The 3D model is generated from tracings of 363 slice images, including the image in C. Plant Physiol. Vol. 147, 2008 1455 Staehelin and Kang Figure 2. Thin section electron micrographs of ER export sites with budding COPII vesicle profiles (arrowheads) in a tobacco BY-2 cell (A; from Ritzenthaler et al., 2002) and in an Arabidopsis root columella cell (B). imately 4 nm. This z axis resolution is approximately 20-fold better than that of classical EM thin sections and approximately 100-fold better than of typical confocal images. To generate a 3D reconstruction of a given cellular structure from a tomographic data set, the structure has to be traced in every serial slice image (typically approximately 2 nm thick) and the tracings combined into a 3D model that can be viewed from all sides with the help of dedicated computer software programs (Fig. 1D; Supplemental Movie S1). The tomographic reconstructions (models) of ER and Golgi membrane systems shown in this review involved the generation of tracings of each membrane compartment in up to 570 sequential slice images! The validity of these models can be verified by comparing the geometries of the different membrane compartments with corresponding freeze-fracture micrographs of high-pressurefrozen cells (Craig and Staehelin, 1988). It is instructive to compare the smoothness of the membranes in the freeze-fracture micrographs and the corresponding membrane smoothness seen in the best tomographic reconstructions. To increase the overall size of a reconstruction, tomograms of serial sections and of montaged images can be joined together (Ladinsky et al., 1999). The IMOD suite of image processing tools employed in all of our studies can be downloaded free of charge at the Web site of the Boulder Laboratory of 3D Electron Microscopy of Cells (University of Colorado, Boulder; http://bio3d.colorado.edu/). A more detailed and practical guide to electron tomography of membranous organelles is presented in a review by Donohoe et al. (2006). CURRENT STATUS OF PLANT SECRETORY PATHWAY RESEARCH During the past 10 years, research on secretory pathway trafficking in plants, i.e. trafficking between ER, Golgi, and TGN cisternae and beyond, has been driven by studies using a combination of molecular biology and confocal microscopy techniques. These investigations have produced three types of advances: (1) they have identified and characterized a multitude of structural and regulatory proteins that operate at different sites in the secretory pathway; (2) they have produced insights into targeting and retention signals; and (3) they have provided information on the dy1456 namic properties of the compartments with which these proteins are associated. A number of excellent reviews summarizing progress in these fields of research have been published in recent years (SaintJore-Dupas et al., 2004; Hawes, 2005; Hawes and Satiat-Jeunemaitre, 2005; Aniento et al., 2006; Hanton et al., 2006; Lam et al., 2007; Moreau et al., 2007; Robinson et al., 2007). The purpose of this review is 2-fold: first, to provide an update of studies of the structural organization of the secretory membrane systems of plants and, second, to explain the implications of this new information in terms of functional organization of these membrane compartments. Research on the Golgi apparatus has a long history of being controversial (see Farquhar and Palade [1998] for a review of early studies and Robinson et al. [2007] for a discussion of more recent controversies in the plant field). Controversies are beneficial for science as long as the focus remains on the facts, because it forces the participants to develop better tests of competing hypotheses. This competition has definitely aided plant secretory pathway research. The following is a current list of agreed-upon facts and controversial features of this field. Agreed-upon facts: 1. The plant Golgi apparatus consists of dispersed, functionally independent Golgi stack units (Moore et al., 1991; Staehelin and Moore, 1995; Boevink et al., 1998; Nebenführ et al., 1999). 2. Each Golgi unit is comprised of cisternae that exhibit a polar organization both with respect to their architecture and to their function, and a TGN type of cisterna on the trans-side of the stack (Staehelin et al., 1990; Zhang et al., 1993). 3. The Golgi units retain their structural integrity and remain functionally active during mitosis (Nebenführ et al., 2000; Segui-Simarro et al., 2004; Reichardt et al., 2007). 4. The Golgi units travel along actin tracks (not microtubules) through the cytoplasm with periods of rapid translocational movements alternating with slower and more wiggling motions (Boevink et al., 1998; Nebenführ et al., 1999). 5. Coat protein complex I (COPI) vesicles bud from the rims of Golgi cisternae, and secretory vesicles and clathrin-coated vesicles from TGN cisternae (Staehelin et al., 1990; Pimpl et al., 2000; Donohoe et al., 2007). 6. Plants produce COPII proteins and these proteins are essential for ER export (Movafeghi et al., 1999; Phillipson et al., 2001). Controversies surround the following features: 1. The mechanism of ER-to-Golgi transport. Is this transport mediated by vesicles formed at ER export sites or by tubules (Hawes and Satiat-Jeunemaitre, 2005; Donohoe et al., 2007; Moreau et al., 2007; Robinson et al., 2007)? 2. The nature of the relationship between ER export sites and Golgi stacks. Are Golgi stacks permaPlant Physiol. Vol. 147, 2008 Electron Tomography Analysis of Plant Golgi Stacks nently attached to ER export sites, or is the association with these sites transient (daSilva et al., 2004; Yang et al., 2005)? 3. The mechanism of trafficking of cargo molecules through Golgi stacks. Do cargo molecules traffic according to the cisternal progression/maturation model or the vesicle shuttle model (Nebenführ, 2003)? 4. The role of the TGN in endocytic trafficking/membrane recycling activities (Geldner, 2004; Lam et al., 2007). In the following, we demonstrate that new, highresolution structural information produced with the help of electron tomography has provided both refining insights into the agreed-upon features of the plant Golgi apparatus as well as clear answers to some of the controversial questions. PLANT ER EXPORT SITES BUD COPII VESICLES BUT DO NOT PRODUCE TUBULES THAT CONNECT DIRECTLY TO GOLGI CISTERNAE One of the longest lasting controversies in plant Golgi research surrounds the mechanism of transport of cargo molecules from ER to Golgi. Do COPII-type vesicles mediate this transport as in other eukaryotic systems (Bonifacino and Glick, 2004), or does this transport occur via tubular connections between ER and Golgi membranes (Hawes and Satiat-Jeunemaitre, 2005; Moreau et al., 2007)? This controversy can be traced to the use of chemical fixation and staining methods, which are unable to preserve ER export sites, termed ERES by some researchers (daSilva et al., 2004), of plants in their natural state. For example, despite 30 years of efforts, no researcher to date has succeeded in producing clear images of budding COPII vesicles in chemically fixed higher plant cells (Robinson, 1980; Aniento et al., 2006). The case for tubular connections between ER and Golgi membranes rests on just two publications (Juniper et al., 1982; Harris and Oparka, 1983), which employed chemical fixation and staining protocols that are now known to produce major changes in the architecture of ER and Golgi membranes. In particular, the harsh chemical reaction conditions used during zinc iodine osmium staining to generate distinctive but membranedistorting stain deposits in ER and Golgi cisternae not only dramatically alter the fine structure of these membrane systems, but also completely destroy all ribosomes and other cytoplasmic proteins. In light of these facts, and because tubular connections between ER and Golgi membranes have never been observed in cells preserved by other specimen preparation methods (chemical fixation, cryofixation/freeze-substitution, or cryofixation/freeze-etching), the heavy-metal-staininginduced tubular precipitates described by Juniper et al. (1982) and Harris and Oparka (1983) should no longer be used as evidence in support of claims that plants employ tubules to transport cargo and membrane Plant Physiol. Vol. 147, 2008 molecules from ER to Golgi cisternae (Brandizzi et al., 2002; Moreau et al., 2007). As reported by Yang et al. (2005), live cell imaging of tobacco (Nicotiana tabacum) BY-2 cells expressing Sec13:GFP, a COPII coat protein and ER export site marker, produced ER-associated fluorescent punctate structures that had an extremely short half-life of only ,10 s and seemed to form randomly over the ER. The number of such structures was also greater than the number of Golgi stacks, and the Golgi stacks were seen to associate intermittently with such sites. These results, which partly contradict the findings of daSilva et al. (2004; see below), suggest that COPII buds produced by plant cells are unstable, transient membrane structures that are resorbed back into the ER unless they are immediately transferred to a Golgi stack. Considering the slow rate of chemical fixation compared to the rate of COPII vesicle assembly/disassembly and the rapid translational movements of the Golgi stacks, it is easy to understand why budding COPII vesicles and their spatial relationship to Golgi stacks have never been preserved in chemically fixed cells. Nevertheless, considering that the first images of high-pressure-frozen and freeze-fractured root tip cells demonstrating ER cisternae budding vesicles in close proximity to Golgi stacks were published 20 years ago (Craig and Staehelin, 1988) and have been confirmed repeatedly since then (Ritzenthaler et al., 2002; Donohoe et al., 2007; Robinson et al., 2007), it is astonishing that the involvement of COPII vesicles in ER-to-Golgi transport is still being questioned. Figure 2 shows two examples of COPII vesicles budding from ER export sites as seen in high-pressure-frozen and freeze-substituted plant cells. One example illustrates a thin section image of budding COPII vesicles in a tobacco BY-2 cell and the other a budding COPII vesicle in a root columella cell of Arabidopsis (Arabidopsis thaliana; Fig. 2B). One problem that has increased the difficulty of obtaining unambiguous images of budding COPII vesicles in high-pressure-frozen plant cells relates to the 3D architecture of the ER export sites in many cell types. The problem is illustrated in Figure 3, which depicts an electron tomography-based 3D reconstruction of an ER export site in a root meristem cell. This ER export site contains four budding COPII vesicles, but the vesicles are seen budding from three different tubular ER domains. This geometry makes it not only difficult to obtain a thin section image of an ER export site that clearly illustrates the budding configuration of a COPII vesicle together with a clear image of an ER membrane tubule (Fig. 3A), but also makes it nearly impossible to show in a single thin section micrograph two or more budding vesicles together with the originating ER membranes. Even in cell types such as columella cells that contain more sheet-like ER cisternae, the budding COPII vesicles at any given ER export site are seen to arise on different tubular membrane domains and not on one contiguous membrane region (Figs. 2B and 3B). The identity of the budding 1457 Staehelin and Kang Figure 3. 3D tomographic model images of ER export sites in a root meristem cell (A) and in a root columella cell (B). A, Four budding COPII vesicles (arrowheads) are located adjacent to the cis-side of a Golgi stack (stippled structure). Note that these COPII buds are forming on three different tubular ER domains and face in different directions. B, Face-on view of an ER export site associated with the cortical ER of a columella cell in which the cisternae are more sheet-like. Here too the COPII buds (arrowheads) are seen budding from different ER domains in close proximity to the cis-most cisterna (C1, stippled structure) of a docked Golgi stack. COPII vesicles seen in high-pressure-frozen cells has been confirmed by EM immunolabeling with antiAtSar1 antibodies (Donohoe et al., 2007). COPII VESICLES ARE BORN WITH A SCAFFOLD LAYER THAT CAPTURES PASSING GOLGI STACKS AND MEDIATES COPII VESICLE TRANSFER TO THE GOLGI Plant Golgi stacks are mobile organelles propelled by molecular motor proteins along actin filaments at speeds of up to 4 mm/s (Boevink et al., 1998; Nebenführ et al., 1999). This mobility raises questions as to how vesicular trafficking between ER and Golgi is organized and regulated. Two models have been proposed to explain this transport. Based on the observation that Golgi stacks alternate between phases of fast (up to 4 mm/s), linear movements and slower (,0.4 mm/s), wiggling-type motions at set locations, the ‘‘stop-and-go’’ model (Nebenführ et al., 1999) postulates that Golgi stacks are induced by a ‘‘signal’’ to stop at active ER export sites to pick up COPII vesicles and then resume their fast movement (go phase) after vesicle transfer. In contrast, and based on live cell images of ER export sites and Golgi stacks moving together, the ‘‘ER-Golgi secretory unit’’ model (daSilva et al., 2004) proposes that ER export sites are stably coupled to Golgi stacks. However, measurements of the rates of movement of the coupled structures shown in their video images indicate that these ER-Golgi secretory units move at the speed of slower, wiggling Golgi and not at the speed of fast Golgi. We have tested these models by determining the percentage of Golgi stacks that are closely associated with an ER export site and the percentage that is located at a distance from the closest ER cisterna. In Arabidopsis root apical meristem cells, approximately 70% of the Golgi were associated with an ER export site, whereas in the gravity-sensing columella cells, only approximately 15% of the Golgi stacks were 1458 located in the vicinity of an ER cisterna (B.-H. Kang and L.A. Staehelin, unpublished data). These results contradict the prediction of the ER-Golgi secretory unit model that close to 100% of the Golgi stacks should be coupled to an ER export site, while supporting the contention of the stop-and-go model that Golgi stacks associate transiently with ER export sites. This begs the question as to the nature of the postulated Golgi ‘‘stop signal’’ produced by ER export sites, and how Golgi stacks become physically coupled to ER export sites as observed by daSilva et al. (2004). Our electron tomograms suggest that both of these activities are mediated by scaffold-type molecules that assemble on COPII vesicles and are transferred to the cis-side of the Golgi matrix together with the vesicles. The COPII-type vesicles that bud from ER export sites in Arabidopsis are approximately 60 nm in diameter. In both thin sections and tomographic slice images, their COPII coat proteins produce a darker staining of the membrane around the buds compared to the adjacent ER membrane (Figs. 2 and 4). In addition to the COPII coat, budding COPII vesicles also assemble a scaffold layer on their outer surface, which is seen as a 40-nm-thick ribosome-excluding zone (Fig. 4, A and B). This COPII scaffold, then, binds directly to the cis-most region of the Golgi matrix, which encompasses each Golgi stack in a cocoon-like fashion (Figs. 4, C and D, and 5). In yeast, COPII vesicles are tethered to the Golgi membrane by a long coiled-coil protein, Uso1p (Cao et al., 1998), and the Arabidopsis genome encodes a protein displaying significant amino acid similarity with Uso1p (At3g27530). Based on the affinity of the COPII scaffold to the cis-side of the Golgi matrix, the COPII scaffold is likely to contain the Arabidopsis Uso1related protein. The COPII scaffold layer linked to the cis-Golgi matrix merges with the matrix structure, transferring enclosed COPII vesicles to the Golgi. (Figs. 4, E–H, and 13A). These observations suggest that the COPII scaffold proteins mediate both the physical coupling between ER export sites and Golgi stacks, noted by daSilva et al. (2004), as well as the ‘‘signaling’’ that has been postulated to induce passing Golgi stacks to stop at active ER export sites (Nebenführ et al., 1999). They also explain why drugs that perturb the actin cytoskeleton do not inhibit ER-to-Golgi transport in plant cells (Brandizzi et al., 2002). The functionality of the scaffold-mediated coupling between ER export sites and Golgi stacks is supported also by the finding that Golgi stacks docked to an ER export site possessed 3 times more free COPII vesicles in the vicinity of their cis-Golgi cisternae than Golgi located .300 nm from an ER cisterna (B.-H. Kang and L.A. Staehelin, unpublished data). Based on these and other findings, we have formulated the ‘‘dock, pluck and go’’ model of ER-to-Golgi trafficking shown in Figure 6. The model postulates that the scaffold of budding COPII vesicles captures passing Golgi by binding to the cis-side of the Golgi matrix and that this docking mechanism automatically orients the cis-side Plant Physiol. Vol. 147, 2008 Electron Tomography Analysis of Plant Golgi Stacks of the Golgi toward the ER export site. It also suggests that the wiggling of the connected Golgi stacks provides the energy to pluck the budding COPII vesicles from the ER and that, when the COPII vesicle harvesting is complete, the Golgi is free to resume its translational movement along the actin filaments. This mechanical harvesting mechanism has yet to be tested experimentally, but it would greatly increase the efficiency of ER-to-Golgi vesicle transport by ensuring that COPII vesicles are released from an ER export site only when a Golgi stack is available to pick up the freed vesicles. It also explains why no COPII vesicle fission machinery has been identified in biochemical, molecular, or genetic studies of yeast or mammalian cells (Kirchhausen, 2000). ELECTRON TOMOGRAMS PROVIDE STRUCTURAL EVIDENCE FOR ASSEMBLY INTERMEDIATES OF CIS-GOLGI CISTERNAE The proof for cis-to-trans vectorial transport of cargo molecules through Golgi stacks was provided in the 1960s (for review, see Farquhar and Palade, 1981), but how these molecules are moved from one side of a stack to the other has remained controversial for several decades. The two principal hypotheses are the ‘‘vesicle shuttle’’ and the ‘‘cisternal progression/maturation’’ models of intra-Golgi transport (Farquhar and Palade, 1981; Becker et al., 1995; Glick and Malhotra, 1998; Nebenführ, 2003). The vesicle shuttle model proposes that the stacked Golgi cisternae are stable, long-lived structures that serve as repositories of sets of processing enzymes and that the cargo molecules are transported from one cisterna to the next in an anterior direction by shuttling vesicles. In contrast, the cisternal progression model postulates that the secretory cargo never leaves individual cisternae, and that transport across the stack is brought about by the assembly of new cisternae on the cis-side of the stack and the fragmentation or shedding of old cisternae on the trans-side. According to this model, the function of the COPI-type vesicles that bud from the rims of the cisternae is to transport the cisternal enzymes in a retrograde direction to maintain the characteristic polar distribution of those enzymes across the stack. As discussed by Nebenführ (2003), the preponderance of evidence now supports the cisternal progression/maturation model, but several major predictions of the two models have yet to be tested experimentally. Figure 4. Transfer of COPII vesicles and their scaffolds to the cis-Golgi matrix. A and B, Tomographic slice image (A) and a corresponding model (B) of a COPII vesicle budding from an ER membrane. The COPII bud is surrounded by an approximately 40-nm-wide, ribosome-excluding scaffold. C and D, Tomographic slice image (C) and a corresponding model (D) of a COPII bud (arrowhead), which is still attached to the ER but whose scaffold has become connected to the cis-side of the Golgi Plant Physiol. Vol. 147, 2008 matrix. E and F, Tomographic slice image (E) and a corresponding model (F) of a released COPII vesicle (arrow) that is still close to the ER membrane. The COPII scaffold is linked to the Golgi matrix. G and H, Tomographic slice image (G) and a corresponding model (H) illustrating a COPII vesicle (arrow) that has been transferred to the cis-face of the Golgi stack together with its COPII scaffold. The distal side of the COPII vesicle is completely surrounded by the Golgi matrix. Scale bars 5 300 nm. 1459 Staehelin and Kang Figure 5. Electron tomographic model of a Golgi stack and its encompassing, ribosome-excluding scaffold (Golgi matrix). A TGN cisterna is also contained within the trans-side extension of the Golgi matrix. To date, the four major lines of experimental evidence that support the cisternal progression/maturation model are (1) bulky secretory products such as algal scales (Becker et al., 1995), collagen complexes (Bonfanti et al., 1998), and protein aggregates (Hillmer et al., 2001) never leave the cisternae during intra-Golgi transport; (2) immunolabeling and biochemical fractionation experiments have demonstrated that COPItype vesicles transport primarily Golgi enzymes in a retrograde direction and not cargo molecules in an anterograde direction (Martinez-Menarguez et al., 1999; Lanoix et al., 2001); (3) live cell imaging has demonstrated maturational changes in individual Golgi cisternae in Saccharomyces cerevisiae (Losev et al., 2006; Matsuura-Tokita et al., 2006); and (4) live cell imaging and correlative electron tomography studies of Pichia pastoris cells have shown that trans-Golgi cisternae are shed from Golgi stacks (Bevis et al., 2002; Mogelsvang et al., 2003). One question that has yet to be answered unambiguously is the type of compartment COPII vesicles fuse with upon arriving at the Golgi. According to the cisternal progression model, cis-Golgi cisternae should be highly variable in size and shape, reflecting the de novo assembly of such cisternae from COPII vesicles. In contrast, the vesicle shuttle model predicts cis-Golgi cisternae to possess a more uniform and mature architecture. In thin section electron micrographs of high-pressurefrozen/freeze-substituted plant Golgi stacks, cis-, medial-, and trans-Golgi cisternae can be positively identified by means of three structural criteria: position in a stack, lumenal staining, and cisternal geometry (Staehelin et al., 1990). Golgi stacks in Arabidopsis and tobacco are typically composed of five to seven cisternae, the first two of which are usually classified as cis-type. cis1460 Type cisternae are smaller than medial- and transcisternae, they have a variable shape, and their lumen are not only wider but also very lightly stained (Fig. 1, B and C). The tremendous structural variability of plant cisGolgi cisternae is illustrated in the three tomographic models of Figure 7. These models depict cis-side, faceon views of three different root apical meristem Golgi stacks, with the cis-most (C1) cisterna colored orange and the second cis-cisterna (C2) colored green. The most striking difference between the C1 cisternae shown in Figure 7 is the huge variability in their size and shape. Thus, whereas the first stack displays only one tiny cisternal initial and the second one two branched, tubular cisternal initials, the third stack has a C1 cisterna that is more mature and disc-shaped and has tubular extensions along its margins. The models also suggest that fusion of the COPII vesicles with growing C1 cisternae occurs primarily at the ends of the tubular domains. In all instances, the growing C1 cisternae appear to use the C2 cisternae as a growth template, as evidenced by the fact that C1 cisternae never extend beyond the edge of the C2 cisterna on which they are growing. Furthermore, C2 cisternae are always smaller than the C3 cisternae and associated with both COPII and COPIa-type but not COPIb-type vesicles (see below), consistent with the idea that they too are growing cisternae. Together, these models provide evidence for the de novo assembly of cisGolgi cisternae as postulated by the cisternal progression/maturation model. Figure 6. Dock, pluck, and go model of ER-to-Golgi vesicle trafficking. A, Go-phase: Golgi stacks travel along actin filaments propelled by myosin motors. B, Dock- and Pluck-phase: The COPII scaffold of a budding vesicle attaches to the cis-side of the Golgi matrix and pulls the passing Golgi off the actin track. Once the COPII scaffold binds to the Golgi matrix, the wiggling motion of the Golgi stack facilitates release of the COPII vesicles by plucking. After COPII vesicle harvesting, the Golgi is free to resume its movement along the actin track. Plant Physiol. Vol. 147, 2008 Electron Tomography Analysis of Plant Golgi Stacks Figure 7. Face-on views of three Golgi stack models in which the cismost (C1) cisternae are colored orange (arrows) and the underlying C2 cis-cisternae are colored green. The cis-most cisternae vary in size from a small blob (A), to two intermediate-sized, branched tubules (B), and to a slightly larger disc with tubular extensions around its margins (C). These highly variable shapes are consistent with the de novo assembly of cis-Golgi cisternae from COPII vesicle (yellow spheres) as postulated by the cisternal progression model of Golgi trafficking. CIS-GOLGI CISTERNAE ARE BIOSYNTHETICALLY INACTIVE DURING THEIR ASSEMBLY BUT ENABLE PROTEIN AGGREGATE/COMPLEX FORMATION Another basic question that has not been addressed in the plant Golgi literature is whether cis-Golgi cisternae are functionally active while they are being assembled or whether they only become functionally active after their assembly is complete. One of the characteristic structural features of cis-Golgi cisternae in high-pressure-frozen/freeze-substituted plant cells is the lack of luminal staining of cis-Golgi cisternae (Figs. 1, B and C, and 15D), i.e. they appear devoid of stainable materials except for seed storage protein aggregates in cisternal margins (Otegui et al., 2006) and cell wall scale backbone complexes in Scherffelia dubia (Donohoe et al., 2006, 2007). Indeed, the structural transition between cis- and medial-cisternae is marked by the sudden appearance of stained luminal contents in the medial-cisternae (Fig. 1B; Staehelin et al., 1990; Donohoe et al., 2007). To determine if the lack of luminal staining of cisGolgi cisternae reflects a lack of biosynthetic activities, we have mapped the distribution of a-1,2 mannosidase I, the enzyme that mediates the first reaction of the N-linked glycan-processing pathway in the Golgi (Moremen et al., 1994), using an antibody that detects the two native a-1,2 mannosidase I isotypes of Arabidopsis (At1g51590 and At3g21160). Our data demonstrate (Fig. 8) that native a-1,2 mannosidase I localizes nearly exclusively to medial-Golgi cisternae and is essentially absent from cis-Golgi cisternae and the ER. This labeling pattern contrasts with studies of transgenic cells expressing a-1,2 mannosidase I-GFP constructs in which the labeled protein is seen to accumulate in both cis- and medial-Golgi cisternae (Nebenführ et al., 1999) and even ER cisternae (SaintJore-Dupas et al., 2006), and highlights the pitfalls of using GFP fusion proteins for such localization studies. The abrupt change in luminal staining from cis- to medial-Golgi cisternae suggests that the accumulation Plant Physiol. Vol. 147, 2008 of sugar nucleotides, the activation of biosynthetic enzymes, and the appearance of cargo molecules with modified glycans only begins in medial-Golgi cisternae, i.e. only after the assembly of the cis-Golgi cisterna is complete. This finding is also consistent with the demonstration that the backbone epitopes of pectic polysaccharides and xyloglucans can be detected by immunological means only in medial- and transbut not in cis-Golgi cisternae (Zhang and Staehelin, 1992). In electron micrographs of high-pressure-frozen and freeze-substituted cells of the alga S. dubia (Donohoe et al., 2007) and of the yeast P. pastoris (Mogelsvang et al., 2003), the cis-Golgi cisternae display the same type of unstained lumen as in plants. This suggests that the cis-Golgi cisternae in these organisms are also biosynthetically inactive during their assembly. In support of this hypothesis, the cis-Golgi cisternae of the scale-forming alga Scherffelia only contain wispy cell wall scale complexes, and these complexes only become more heavily glycosylated and stained once they reach the medial-cisternae (L.A. Staehelin, unpublished data). Taken together, these results suggest that in walled cells, i.e. cells that employ H1-gradients to create turgor pressure and to drive their plasma membrane transport systems, cisternal assembly and biosynthesis of secretory products are mutually incompatible Golgi activities and therefore occur sequentially during cisternal maturation. Figure 8. Immunoelectron tomography localization of native a-1,2 mannosidase I (ManI) in an Arabidopsis meristem Golgi stack. A to C, Electron micrographs of three serial thin sections labeled with antiManI 15-nm immunogold particles. D, 3D tomographic model of the Golgi stack and immunogold particles seen in A to C. Note that nearly all of the immunogold particles are associated with the C3 and C4 medial-cisternae (arrowheads). The Golgi cisternae models were rendered semitransparent to make all of the gold particles visible. Scale bars 5 100 nm. 1461 Staehelin and Kang Figure 9. COPII-, COPIa-, and COPIb-type vesicles of S. dubia. A to C, A gallery of tomographic slice images showing COPII-type vesicles. D to F, A gallery of tomographic slice images showing COPIa-type vesicles. G to I, A gallery of tomographic slice images depicting COPIb-type vesicles. In C, F, and I, models of COPII, COPIa, and COPIb vesicles are overlaid on tomographic slices, respectively. J, 3D model of Golgi associated vesicles superimposed on a single slice of an S. dubia Golgi stack illustrating the nonoverlapping distribution of COPIa-type (light green spheres) and COPIb-type vesicles (light purple spheres). COPII-type vesicles (gold spheres) colocalize with COPIatype vesicles, though. Light blue and pink spheres in the trans-side of the Golgi represent secretory and clathrin-coated vesicles (Donohoe et al., 2007). Scale bar 5 100 nm. GOLGI STACKS PRODUCE TWO TYPES OF COPI VESICLES, COPIA AND COPIB As already discussed in preceding sections, COPIItype vesicles transport membrane and cargo molecules from ER to Golgi. In contrast, COPI vesicles originate on Golgi cisternae and transport cisternal membrane 1462 proteins in a retrograde direction (Malhotra and Mayor, 2006). The Golgi-to-ER, COPI vesicle-mediated transport is responsible for returning escaped ER proteins to the ER, whereas the COPI vesicles involved in retrograde intra-Golgi transport help maintain the steady state of enzyme distribution within Golgi stacks as the individual cisternae traverse the stacks in an anterograde direction. This latter type of retrograde transport has been postulated to operate between the TGN and cis-Golgi cisternae and to possibly extend back to the ER. Here we address the question whether COPI vesicles derived from medial- and trans-Golgi cisternae can recycle molecules directly back to the ER, or if they are limited to intra-Golgi transport activities. To answer this question, we took advantage of the increased resolution of electron tomographic images of high-pressure-frozen/freeze-substituted Golgi stacks in which COPII and COPI vesicles can be identified by means of six structural criteria: coat architecture, coat thickness, membrane structure, cargo staining, cisternal origin, and spatial distribution (Donohoe et al., 2007). In addition, COPI and COPII buds and vesicles were positively identified by immunolabeling techniques. Using this multi-parameter approach and the ability to map the distribution of all vesicles in 3D space, we demonstrated that COPII and COPI vesicles are confined to well-defined regions around ER export sites and Golgi cisternae and that Golgi stacks produce two types of COPI vesicles, COPIa and COPIb. COPII vesicles bud from ER export sites and localize to the space between the ER and cis-Golgi cisternae (Fig. 9, A–C). COPIa vesicles bud exclusively from cisGolgi cisternae and, like these cisternae, they have lightly stained contents (Fig. 9, D–F). They are also confined to the ER-cis-Golgi interface, colocalizing with COPII vesicles. In contrast, the COPIb vesicles bud exclusively from medial, trans, and early TGN cisternae, have darkly stained luminal contents like the cisternae from which they originate (Fig. 9, G–I), and are confined to the space around these latter cisternae. To determine the generality of this distribution pattern, we compared the distribution of COPIa and COPIb vesicle types in Arabidopsis and in Scherffelia, which contains two Golgi stacks, each of which generates a new cis-Golgi cisterna every 20 to 30 s (McFadden et al., 1986). Due to the large size of those Golgi stacks and the high rate of cisternal turnover, the number of COPII and COPI vesicles produced by Scherffelia is very large, which allows for the clear demarcation of the domains in which the COPIa and COPIb operate (Fig. 9J). In Arabidopsis, the distribution patterns of COPIa- and COPIb-type vesicles are the same as in Scherffelia, but due to the much smaller number of vesicles, the patterns can be confirmed only by means of statistical analysis of the vesicle distribution around many Golgi stacks. A model summarizing the trafficking patterns of COPII, COPIa, and COPIb, as well as of clathrin-coated and secretory vesicles around ER, Golgi, and TGN cisternae (Fig. 10), is illustrated in Figure 10. Based on these vesicle distribution Plant Physiol. Vol. 147, 2008 Electron Tomography Analysis of Plant Golgi Stacks Figure 10. Schematic diagram illustrating the sites of origin and the trafficking routes of five types of ER/Golgi/TGN-associated vesicles. COPIa-type vesicles bud from assembling cis-cisternae and recycle molecules back to the ER, whereas the COPIb-type vesicles are produced by medial, trans, and early TGN cisternae and recycle molecules between these cisternae. patterns, the COPIa vesicles are postulated to recycle escaped ER proteins from forming cis-Golgi cisternae back to the ER, and the COPIb vesicles to mediate retrograde transport of Golgi enzymes between early TGN, trans-, medial-, and late-stage cis-Golgi cisternae. PLANT GOLGI STACKS DIVIDE BY FISSION Three mechanisms of Golgi division have been reported in the literature (Munro, 2002): (1) the ‘‘disintegration and reassembly model,’’ which appears to be the principal mechanism in mammalian cells; (2) the ‘‘de novo construction model’’ employed by the yeast P. pastoris (Bevis et al., 2002) and by mammalian cells treated with microtubule-disrupting drugs such as nocodazole (Cole et al., 1996); and (3) the ‘‘fission model’’ used in plants, algae, and protozoa (Morre et al., 1971; Benchimol et al., 2001; Pelletier et al., 2002). In Arabidopsis shoot meristem cells, the number of Golgi stacks has been shown to double from approximately 34 to approximately 66 during the G2 stage of the cell cycle, i.e. just prior to mitosis (Segui-Simarro and Staehelin, 2006). Golgi stack duplication can also occur in interphase cells during cell growth. The fission model of Golgi multiplication arose in the 1960s based on electron micrographs of Golgi stacks that exhibited two partial stacks attached to one or several larger trans-cisternae (for review, see Morre et al., 1971), but the mechanism of division has remained a mystery. Assuming that Golgi stacks traffic according to the cisternal progression/maturation model discussed above, the fission model predicts that duplication starts with the assembly of two halfsized cis-cisternae instead of just one on the surface of Plant Physiol. Vol. 147, 2008 a laterally expanded C2 cisterna. Each of these new, separate cisternae then serves as an independent template for the assembly of the next cisterna. As this process is repeated and as the older trans-side cisternae are shed, the two new stacks increase in size, eventually giving rise to two complete and separate daughter Golgi stacks when the last joining transcisterna is released. The tomographic model of a dividing Golgi stack illustrated in Figure 11 provides several additional lines of evidence for the fission model. First, the dividing Golgi is seen to be docked to an active ER export site, and several released COPII vesicles are located adjacent to the cis-Golgi cisternae. Second, the two cis-most Golgi cisternae (C1, C2) are also smaller than the underlying older cisternae and display morphological features of assembling cis-Golgi cisternae (compare Fig. 11B with Fig. 7). Finally, the model shows three TGN cisternae at different stages of maturation and moving away from the Golgi stack, which supports the interpretation that this Golgi stack was actively shedding trans-cisternae at the time of cryofixation. Together, these structural features provide strong direct support for the fission model of plant Golgi multiplication. TRANSFORMATION OF A TRANS-MOST GOLGI CISTERNA INTO A TGN CISTERNA INVOLVES A REDUCTION IN SURFACE AREA AND RECRUITMENT OF THE GTPASE RABA4B By definition, the TGN sorts and packages Golgi products into secretory and clathrin-coated vesicles for transport to the cell surface and to vacuoles/lysosomes, respectively (Griffiths and Simons, 1986). In plants, TGN cisternae arise from the trans-most Golgi cisternae by means of a cisternal peeling process that leads to the separation of the TGN cisterna from the stack and to the formation of independent TGN compartments (Figs. 11C, 12, 13, and 14). These independent TGN compartments continue to undergo maturational changes until the entire cisterna fragments into vesicles and small, residual membrane fragments (Figs. 12 and 14). TGN cisternae that are in the process of peeling away from a Golgi stack are referred to as early TGN cisternae (Fig. 12B), and those that have completed this separation are referred to as late TGN cisternae (Fig. 12C). In small meristem cells, where Golgi movements and cytoplasmic streaming are restricted during interphase (B.-H. Kang and L.A. Staehelin, unpublished data) and stop during cell division (Nebenführ et al., 2000), electron tomographic reconstructions often show multiple TGN cisternae at different distances from the trans-side of a Golgi stack (Figs. 11C and 13). In contrast, in large interphase suspension-cultured cells that exhibit fast Golgi movements and vigorous cytoplasmic streaming (Nebenführ et al., 1999), very few Golgi stacks possess a late TGN cisterna near their trans-face, and many stacks even seem to lack a 1463 Staehelin and Kang originating Golgi stacks. Free-floating TGN cisternae have been observed in high-pressure-frozen and freeze-substituted pine cambial cells, which also exhibit rigorous streaming (Rensing et al., 2002). Transformation of a trans-Golgi cisterna into an early TGN cisterna starts with cisternal peeling, a reduction in surface area (approximately 30% in Arabidopsis root meristem cells) by COPI vesicle-mediated recycling, the formation of secretory vesicles outside the plane of the cisterna, and the binding of antiRabA4b and anti-phosphatidyl inositol-4-kinase b antibodies (Preuss et al., 2006; B.-H. Kang and L.A. Staehelin, unpublished data). In contrast, clathrincoated vesicle formation is not a reliable early TGN marker, because the production of such vesicles can be delayed until most secretory vesicles have formed on late TGN cisternae (Figs. 11 and 13). Both RabA4b and phosphatidyl inositol-4-kinase b appear to be components of the coat protein complexes of budding secretory vesicles that contain mostly complex carbohydrate-type cell wall matrix polysaccharides (B.-H. Kang and L.A. Staehelin, unpublished data). LATE TGN CISTERNAE CONTINUE TO MATURE AS INDEPENDENT ORGANELLES AND RELEASE THEIR VESICLES BY MEANS OF FRAGMENTATION The maturational steps of TGN cisternae can be deduced from the sequence of structural and compositional changes seen in reconstructed meristem GolgiTGN complexes in which the older TGN cisternae are displaced further from the trans-side of the Golgi stacks (Figs. 11C and 13). The sequence of morphological changes associated with TGN maturation shown in Figure 14 is typical for meristematic cells, except that clathrin-coated buds often form already on early TGN cisternae (Fig. 13, D–E). Early TGN cisternae display a central, flat domain where they were attached to the Golgi stack. After their release from the stack, the late-type TGN cisternae continue to generate budding secretory and clathrincoated vesicles, yielding mature cisternae with a Figure 11. 3D tomographic model of a dividing Golgi stack docked to an ER export site in an Arabidopsis root meristem cell. A, The two sets of cisand medial-cisternae are held together by a larger trans-Golgi cisterna (pink). The Golgi stack is connected to budding COPII vesicles via interactions between the COPII scaffolds and the cis-Golgi matrix (arrowheads). B, Face-on view of the dividing Golgi stack shown in A. The C1 (orange) and C2 (green) cisternae of the two cis-side stacks (C1, C2, C1’, and C2’ marked with arrows) display varying shapes resembling the assembling cis-most cisternae in Figure 7. C, On the trans-side, three TGN cisternae (TGN1–TGN3) at different stages of maturation are seen. distinct TGN cisterna (Zhang and Staehelin, 1992). These observations suggest that shearing forces generated by Golgi movements and cytoplasmic streaming can dislodge and then quickly disperse late and sometimes even early TGN cisternae away from the 1464 Figure 12. 3D models (face-on views) of a trans-most Golgi, an early TGN, and a late TGN cisterna of meristem cell Golgi stacks illustrating the TGN maturation process. CCV, Clathrin-coated vesicle; SV, secretory vesicle. Plant Physiol. Vol. 147, 2008 Electron Tomography Analysis of Plant Golgi Stacks different ratios of these proteins will be locally produced and then delivered to individual Golgi stacks, thereby leading to constantly changing mixtures of cargo molecules in Golgi and TGN cisternae. These findings highlight the inherent plasticity of the TGN compartments of plant cells and their ability to quickly adapt their sorting and packaging systems to the type of cargo molecules present. Electron tomography has also enabled us to address the question as to what happens to TGN cisternae at the end of their life. Do they release completed secretory and clathrin-coated vesicles one at a time, withering away as this loss of membrane and contents occurs, or do they release all of their vesicles simultaneously by means of cisternal fragmentation? Based on a quantitative analysis of large cytoplasmic volumes surrounding Golgi stacks, we estimate that ,10% of the secretory and clathrin-coated vesicles are released prior to the fragmentation of late TGN cisternae. Thus, cisternal fragmentation appears to the principal mechanism for releasing TGN vesicles (Fig. 14A). Mollenhauer (1971) was the first to describe the fragmentation of what he called mature dictyosome cisternae. In the model depicted in Figure 14, 20 secretory vesicles and eight clathrin-coated vesicles together with four membrane fragments of variable size can be discerned. Yet to be determined is the fate of the residual membrane fragments. CONCLUDING REMARKS Figure 13. A, Tomographic slice image of an Arabidopsis meristem cell with two Golgi stacks (Golgi 1, Golgi 2). Golgi stack 1 (B and C) is associated with a late TGN cisterna that is budding mostly secretory vesicles (SV). In contrast, the late TGN cisterna associated with Golgi 2 (D and E) is producing mostly clathrin-coated buds (CCV). This latter type of TGN is also known as partially coated reticulum (Pesacreta and Lucas, 1985). Scale bar in A 5 250 nm, scale bars in B to E 5 200 nm. grape-like architecture (Figs. 12C and 13, B and C). Interestingly, the ratio of complex carbohydratecontaining secretory vesicles to clathrin-coated vesicles on TGN cisternae can vary from 5:1 to 1:3 within a single cell (Fig. 13, A–E). The clathrin-coated, bud-rich TGN cisternae appear to correspond to membrane compartments known as partially coated reticulum (Pesacreta and Lucas, 1985). The variation in vesicle type ratios is most likely due to the presence of varying ratios of vacuolar to secretory cargo molecules within individual TGN cisternae. Obviously, depending on when and where the translation of mRNAs coding for vacuolar and secretory proteins occurs on the ER, Plant Physiol. Vol. 147, 2008 The principal goals of this Update were to develop a coherent framework of nanoscale structural information on ER export sites, Golgi stacks, and TGN cisternae of higher plants and to demonstrate how this information can be used to address questions related to Golgi trafficking and Golgi function. The review started out by highlighting the importance of integrating several new, cutting-edge research tools to gain paradigm-shifting insights into these dynamic membrane systems. It then followed the path taken by Figure 14. 3D model images of a Golgi stack and its TGN cisternae. A, The late TGN cisterna has fragmented into a cluster of secretory vesicles (SV) that surround small, residual membrane fragments. CCV, Clathrincoated vesicle. B, Face-on view of the late TGN model shown in A in which the secretory and clathrin-coated vesicles have been omitted to illustrate the morphologies of the residual fragments of the cisternal membrane (arrowheads). 1465 Staehelin and Kang cargo molecules though these membrane systems, describing the 3D architecture of ER export sites, the mechanism of ER-to-Golgi transport, the assembly of new cis-Golgi cisternae, the immunolocalization of native Golgi enzymes, trafficking of cargo and membrane enzymes, the importance of the Golgi matrix, the mechanism of Golgi stack division, and the structural and functional plasticity of TGN cisternae. Together, this new information should aid other cell biologists in not only interpreting their confocal microscope images of secretory pathway organelles but also appreciating the limits of the confocal microscopy technique. In addition, the new nanoscale database should be a valuable resource for helping molecular biologists, biochemists, and geneticists plan novel types of studies of the plant secretory pathway. Where might these advanced EM and EM tomography techniques be employed next? The greatest need would appear to be in the field of plant cell endocytosis, where confusion reigns supreme with respect to the role of the TGN in endocytosis (Geldner, 2004; Lam et al., 2007). In that field, confocal microscopy studies have reached a resolution-based ceiling, and major progress will require comprehensive, correlative confocal and EM studies of entire cells or large cellular volumes together with the quantitative accounting of all of the membrane compartments, including vesicles, that traffic endocytosed and secretory molecules. A critical challenge of these studies will be to distinguish compartments that are involved in transporting the actual endocytic cargo molecules to the lytic vacuoles from those that serve as recipients of recycled molecules from early endosomal compartments. This analysis will also require a much more detailed characterization of the physicochemical properties and uptake mechanisms of the tracer molecules used in endocytosis experiments such as cationized ferritin and the FM4-64-type dye molecules. Last but not least, these next generation studies will have to distinguish within the TGN which proteins and polysaccharides are new and being packaged for secretion and which molecules have been brought to this and other compartments via endocytosis. The following articles provide examples of the type of comprehensive 3D EM/EM tomography studies needed in the field of endocytosis research: Golgi region of a liver cell (amazing models!; Marsh et al., 2001); membranes and microtubules involved in syncytial-type cytokinesis (Otegui et al., 2001); membranes and microtubules involved in somatic cell cytokinesis (Segui-Simarro et al., 2004); and cell cycle-dependent changes in Golgi, TGN, vacuole, and multivesicular body compartments in entire cells (Segui-Simarro and Staehelin, 2006). Supplemental Data The following materials are available in the online version of this article. Supplemental Figure S1. Principles of electron tomography. 1466 Supplemental Movie S1. 3D model of the Arabidopsis Golgi stack in Figure 1. Received April 4, 2008; accepted May 22, 2008; published August 6, 2008. LITERATURE CITED Aniento F, Matsuoka K, Robinson DG (2006) ER-to-Golgi transport: the COPII pathway. In DG Robinson, ed, The Plant Endoplasmic Reticulum. Springer Verlag, New York, pp 99–124 Austin JR II, Segui-Simarro JM, Staehelin LA (2005) Quantitative analysis of changes in spatial distribution and plus-end geometry of microtubules involved in plant-cell cytokinesis. J Cell Sci 118: 3895–3903 Becker B, Bolinger B, Melkonian M (1995) Anterograde transport of algal scales through the Golgi complex is not mediated by vesicles. Trends Cell Biol 5: 305–307 Benchimol M, Ribeiro KC, Mariante RM, Alderete JF (2001) Structure and division of the Golgi complex in Trichomonas vaginalis and Tritrichomonas foetus. Eur J Cell Biol 80: 593–607 Bevis BJ, Hammond AT, Reinke CA, Glick BS (2002) De novo formation of transitional ER sites and Golgi structures in Pichia pastoris. Nat Cell Biol 4: 750–756 Boevink P, Oparka K, Santa Cruz S, Martin B, Betteridge A, Hawes C (1998) Stacks on tracks: the plant Golgi apparatus traffics on an actin/ER network. Plant J 15: 441–447 Bonfanti L, Mironov AA Jr, Martinez-Menarguez JA, Martella O, Fusella A, Baldassarre M, Buccione R, Geuze HJ, Mironov AA, Luini A (1998) Procollagen traverses the Golgi stack without leaving the lumen of cisternae: evidence for cisternal maturation. Cell 95: 993–1003 Bonifacino JS, Glick BS (2004) The mechanisms of vesicle budding and fusion. Cell 116: 153–166 Brandizzi F, Snapp EL, Roberts AG, Lippincott-Schwartz J, Hawes C (2002) Membrane protein transport between the endoplasmic reticulum and the Golgi in tobacco leaves is energy dependent but cytoskeleton independent: evidence from selective photobleaching. Plant Cell 14: 1293–1309 Buckley IK (1973) Studies in fixation for electron microscopy using cultured cells. Lab Invest 29: 398–410 Cao X, Ballew N, Barlowe C (1998) Initial docking of ER-derived vesicles requires Uso1p and Ypt1p but is independent of SNARE proteins. EMBO J 17: 2156–2165 Cole NB, Sciaky N, Marotta A, Song J, Lippincott-Schwartz J (1996) Golgi dispersal during microtubule disruption: regeneration of Golgi stacks at peripheral endoplasmic reticulum exit sites. Mol Biol Cell 7: 631–650 Craig S, Staehelin LA (1988) High pressure freezing of intact plant tissues. Evaluation and characterization of novel features of the endoplasmic reticulum and associated membrane systems. Eur J Cell Biol 46: 81–93 daSilva LL, Snapp EL, Denecke J, Lippincott-Schwartz J, Hawes C, Brandizzi F (2004) Endoplasmic reticulum export sites and Golgi bodies behave as single mobile secretory units in plant cells. Plant Cell 16: 1753–1771 Donohoe BS, Kang BH, Staehelin LA (2007) Identification and characterization of COPIa- and COPIb-type vesicle classes associated with plant and algal Golgi. Proc Natl Acad Sci USA 104: 163–168 Donohoe BS, Mogelsvang S, Staehelin LA (2006) Electron tomography of ER, Golgi and related membrane systems. Methods 39: 154–162 Farquhar MG, Palade GE (1981) The Golgi apparatus (complex)-(19541981)-from artifact to center stage. J Cell Biol 91: 77s–103s Farquhar MG, Palade GE (1998) The Golgi apparatus: 100 years of progress and controversy. Trends Cell Biol 8: 2–10 Geldner N (2004) The plant endosomal system: its structure and role in signal transduction and plant development. Planta 219: 547–560 Glick BS, Malhotra V (1998) The curious status of the Golgi apparatus. Cell 95: 883–889 Griffiths G, Simons K (1986) The trans Golgi network: sorting at the exit site of the Golgi complex. Science 234: 438–443 Hanton SL, Matheson LA, Brandizzi F (2006) Seeking a way out: export of proteins from the plant endoplasmic reticulum. Trends Plant Sci 11: 335–343 Plant Physiol. Vol. 147, 2008 Electron Tomography Analysis of Plant Golgi Stacks Harris N, Oparka K (1983) Connections between dictyosomes, ER and GERL in cotyledons of mung bean. Protoplasma 114: 93–102 Hawes C (2005) Cell biology of the plant Golgi apparatus. New Phytol 165: 29–44 Hawes C, Satiat-Jeunemaitre B (2005) The plant Golgi apparatus: going with the flow. Biochim Biophys Acta 1744: 93–107 Hillmer S, Movafeghi A, Robinson DG, Hinz G (2001) Vacuolar storage proteins are sorted in the cis-cisternae of the pea cotyledon Golgi apparatus. J Cell Biol 152: 41–50 Jakobs S (2006) High resolution imaging of live mitochondria. Biochim Biophys Acta 1763: 561–575 Juniper GE, Hawes C, Horne JC (1982) The relationships between the dictyosomes and the forms of endoplasmic reticulum in plant cells with different export programs. Bot Gaz 143: 135–145 Kirchhausen T (2000) Three ways to make a vesicle. Nat Rev Mol Cell Biol 1: 187–198 Kiss JZ, Staehelin LA (1995) High pressure freezing. In DM Shotton, ed, Rapid Freezing, Freeze Fracture and Deep Etching. Wiley-Liss, New York, pp 89–104 Koster AJ, Grimm R, Typke D, Hegerl R, Stoschek A, Walz J, Baumeister W (1997) Perspectives of molecular and cellular electron tomography. J Struct Biol 120: 276–308 Ladinsky MS, Mastronarde DN, McIntosh JR, Howell KE, Staehelin LA (1999) Golgi structure in three dimensions: functional insights from the normal rat kidney cell. J Cell Biol 144: 1135–1149 Lam SK, Tse YC, Robinson DG, Jiang L (2007) Tracking down the elusive early endosome. Trends Plant Sci 12: 497–505 Lanoix J, Ouwendijk J, Stark A, Szafer E, Cassel D, Dejgaard K, Weiss M, Nilsson T (2001) Sorting of Golgi resident proteins into different subpopulations of COPI vesicles: a role for ArfGAP1. J Cell Biol 155: 1199–1212 Losev E, Reinke CA, Jellen J, Strongin DE, Bevis BJ, Glick BS (2006) Golgi maturation visualized in living yeast. Nature 441: 1002–1006 Malhotra V, Mayor S (2006) Cell biology: the Golgi grows up. Nature 441: 939–940 Marsh BJ, Mastronarde DN, Buttle KF, Howell KE, McIntosh JR (2001) Organellar relationships in the Golgi region of the pancreatic beta cell line, HIT-T15, visualized by high resolution electron tomography. Proc Natl Acad Sci USA 98: 2399–2406 Martinez-Menarguez JA, Geuze HJ, Slot JW, Klumperman J (1999) Vesicular tubular clusters between the ER and Golgi mediate concentration of soluble secretory proteins by exclusion from COPI-coated vesicles. Cell 98: 81–90 Matsuura-Tokita K, Takeuchi M, Ichihara A, Mikuriya K, Nakano A (2006) Live imaging of yeast Golgi cisternal maturation. Nature 441: 1007–1010 McFadden GI, Preisig HR, Melkonian M (1986) Golgi apparatus activity and membrane flow during scale biogenesis in the green flagellate Scherffelia dubia (Prasinophyceae). 2. Cell wall secretion and assembly. Protoplasma 131: 174–184 McIntosh R, Nicastro D, Mastronarde D (2005) New views of cells in 3D: an introduction to electron tomography. Trends Cell Biol 15: 43–51 Mersey B, McCully M (1978) Monitoring the course of fixation of plant cells. J Microsc 166: 43–56 Mogelsvang S, Gomez-Ospina N, Soderholm J, Glick BS, Staehelin LA (2003) Tomographic evidence for continuous turnover of Golgi cisternae in Pichia pastoris. Mol Biol Cell 14: 2277–2291 Mollenhauer HH (1971) Fragmentation of mature dictyosome cisternae. J Cell Biol 49: 212–214 Moore PJ, Swords KM, Lynch MA, Staehelin LA (1991) Spatial organization of the assembly pathways of glycoproteins and complex polysaccharides in the Golgi apparatus of plants. J Cell Biol 112: 589–602 Moreau P, Brandizzi F, Hanton S, Chatre L, Melser S, Hawes C, SatiatJeunemaitre B (2007) The plant ER-Golgi interface: a highly structured and dynamic membrane complex. J Exp Bot 58: 49–64 Moremen KW, Trimble RB, Herscovics A (1994) Glycosidases of the asparagine-linked oligosaccharide processing pathway. Glycobiology 4: 113–125 Morphew MK (2007) 3D immunolocalization with plastic sections. In JR McIntosh, ed, Cellular Electron Microscopy, Vol 79. Elsevier, San Diego, pp 494–511 Morre JD, Mollenhauer HH, Bracker CE (1971) Origin and Continuity of Golgi Apparatus. Springer-Verlag, New York, pp 82–126 Plant Physiol. Vol. 147, 2008 Movafeghi A, Happel N, Pimpl P, Tai GH, Robinson DG (1999) Arabidopsis Sec21p and Sec23p homologs. Probable coat proteins of plant COP-coated vesicles. Plant Physiol 119: 1437–1446 Munro S (2002) More than one way to replicate the Golgi apparatus. Nat Cell Biol 4: E223–224 Nebenführ A (2003) Intra-Golgi transport: escalator or bucket brigade? In DG Robinson, ed, The Golgi Apparatus and the Plant Secretory Pathway, Vol 9. Blackwell Publishing, Oxford, pp 76–89 Nebenführ A, Frohlick JA, Staehelin LA (2000) Redistribution of Golgi stacks and other organelles during mitosis and cytokinesis in plant cells. Plant Physiol 124: 135–151 Nebenführ A, Gallagher LA, Dunahay TG, Frohlick JA, Mazurkiewicz AM, Meehl JB, Staehelin LA (1999) Stop-and-go movements of plant Golgi stacks are mediated by the acto-myosin system. Plant Physiol 121: 1127–1142 Otegui MS, Herder R, Schulze J, Jung R, Staehelin LA (2006) The proteolytic processing of seed storage proteins in Arabidopsis embryo cells starts in the multivesicular bodies. Plant Cell 18: 2567–2581 Otegui MS, Mastronarde DN, Kang BH, Bednarek SY, Staehelin LA (2001) Three-dimensional analysis of syncytial-type cell plates during endosperm cellularization visualized by high resolution electron tomography. Plant Cell 13: 2033–2051 Pelletier L, Stern CA, Pypaert M, Sheff D, Ngo HM, Roper N, He CY, Hu K, Toomre D, Coppens I, et al (2002) Golgi biogenesis in Toxoplasma gondii. Nature 418: 548–552 Pesacreta TC, Lucas WJ (1985) Presence of a partially-coated reticulum in angiosperms. Protoplasma 123: 173–184 Phillipson BA, Pimpl P, daSilva LL, Crofts AJ, Taylor JP, Movafeghi A, Robinson DG, Denecke J (2001) Secretory bulk flow of soluble proteins is efficient and COPII dependent. Plant Cell 13: 2005–2020 Pimpl P, Movafeghi A, Coughlan S, Denecke J, Hillmer S, Robinson DG (2000) In situ localization and in vitro induction of plant COPI-coated vesicles. Plant Cell 12: 2219–2236 Preuss ML, Schmitz AJ, Thole JM, Bonner HK, Otegui MS, Nielsen E (2006) A role for the RabA4b effector protein PI-4Kb1 in polarized expansion of root hair cells in Arabidopsis thaliana. J Cell Biol 172: 991–998 Reichardt I, Stierhof YD, Mayer U, Richter S, Schwarz H, Schumacher K, Jürgens G (2007) Plant cytokinesis requires de novo secretory trafficking but not endocytosis. Curr Biol 17: 2047–2053 Rensing KH, Samuels AL, Savidge RA (2002) Ultrastructure of vascular cambial cell cytokinesis in pine seedlings preserved by cryofixation and substitution. Protoplasma 220: 39–49 Ritzenthaler C, Nebenführ A, Movafeghi A, Stussi-Garaud C, Behnia L, Pimpl P, Staehelin LA, Robinson DG (2002) Reevaluation of the effects of brefeldin A on plant cells using tobacco Bright Yellow 2 cells expressing Golgi-targeted green fluorescent protein and COPI antisera. Plant Cell 14: 237–261 Robinson DG (1980) Dictyosome-endoplasmic reticulum associations in higher plant cells? A serial-section analysis. Eur J Cell Biol 23: 22–36 Robinson DG, Herranz MC, Bubeck J, Pepperkok R, Ritzenthaler C (2007) Membrane dynamics in the early secretory pathway. Crit Rev Plant Sci 26: 199–225 Saint-Jore-Dupas C, Gomord V, Paris N (2004) Protein localization in the plant Golgi apparatus and the trans-Golgi network. Cell Mol Life Sci 61: 159–171 Saint-Jore-Dupas C, Nebenführ A, Boulaflous A, Follet-Gueye ML, Plasson C, Hawes C, Driouich A, Faye L, Gomord V (2006) Plant N-glycan processing enzymes employ different targeting mechanisms for their spatial arrangement along the secretory pathway. Plant Cell 18: 3182–200 Segui-Simarro JM, Austin JR II, White EA, Staehelin LA (2004) Electron tomographic analysis of somatic cell plate formation in meristematic cells of Arabidopsis preserved by high-pressure freezing. Plant Cell 16: 836–856 Segui-Simarro JM, Staehelin LA (2006) Cell cycle-dependent changes in Golgi stacks, vacuoles, clathrin-coated vesicles and multivesicular bodies in meristematic cells of Arabidopsis thaliana: a quantitative and spatial analysis. Planta 223: 223–236 Staehelin LA, Giddings TH Jr, Kiss JZ, Sack FD (1990) Macromolecular differentiation of Golgi stacks in root tips of Arabidopsis and Nicotiana seedlings as visualized in high pressure frozen and freeze-substituted samples. Protoplasma 157: 75–91 1467 Staehelin and Kang Staehelin LA, Moore I (1995) The plant golgi appratus: structure, functional organization and trafficking mechanisms. Annu Rev Plant Physiol Plant Mol Biol 46: 261–288 Yang YD, Elamawi R, Bubeck J, Pepperkok R, Ritzenthaler C, Robinson DG (2005) Dynamics of COPII vesicles and the Golgi apparatus in cultured Nicotiana tabacum BY-2 cells provides evidence for transient association of Golgi stacks with endoplasmic reticulum exit sites. Plant Cell 17: 1513–1531 Zeuschner D, Geerts WJ, van Donselaar E, Humbel BM, Slot JW, Koster AJ, Klumperman J (2006) Immuno-electron tomography of ER exit sites 1468 reveals the existence of free COPII-coated transport carriers. Nat Cell Biol 8: 377–383 Zhang GF, Driouich A, Staehelin LA (1993) Effect of monensin on plant Golgi: re-examination of the monensin-induced changes in cisternal architecture and functional activities of the Golgi apparatus of sycamore suspension-cultured cells. J Cell Sci 104: 819–831 Zhang GF, Staehelin LA (1992) Functional compartmentation of the Golgi apparatus of plant cells: immunocytochemical analysis of high-pressure frozen- and freeze-substituted sycamore maple suspension culture cells. Plant Physiol 99: 1070–1083 Plant Physiol. Vol. 147, 2008