* Your assessment is very important for improving the workof artificial intelligence, which forms the content of this project
Download FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE
Long non-coding RNA wikipedia , lookup
Human genome wikipedia , lookup
Genome (book) wikipedia , lookup
Gene expression profiling wikipedia , lookup
History of genetic engineering wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Non-coding DNA wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Frameshift mutation wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
RNA interference wikipedia , lookup
Microevolution wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Point mutation wikipedia , lookup
Deoxyribozyme wikipedia , lookup
RNA silencing wikipedia , lookup
Nucleic acid tertiary structure wikipedia , lookup
Polyadenylation wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Messenger RNA wikipedia , lookup
History of RNA biology wikipedia , lookup
Non-coding RNA wikipedia , lookup
Transfer RNA wikipedia , lookup
Expanded genetic code wikipedia , lookup
Primary transcript wikipedia , lookup
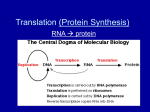
FROM DNA TO PROTEINS: gene expression Chapter 14 LECTURE OBJECTIVES What Is the Evidence that Genes Code for Proteins? How Does Information Flow from Genes to Proteins? How Is the Information Content in DNA Transcribed to Produce RNA? How Is Eukaryotic DNA Transcribed and RNA Processed? How Is RNA Translated into Proteins? What Happens to Polypeptides after Translation? ONE GENE ONE ENZYME IDEA Phenotypic differences due to protein difference Garrod studied ALKAPTONURIA HA accumulated in blood due to an enzyme deficiency MODEL ORGANISMS TO STUDY Drosophila; E. coli; Neurospora crassa (common bread mold) BEADLE & TATUM EXPERIMENT one-gene, one-polypeptide relationship. The gene-enzyme relationship has since been revised to the Example: In hemoglobin, each polypeptide chain is specified by a separate gene. Other genes code for RNA are not translated to polypeptides; some genes are involved in controlling other genes. THE CENTRAL DOGMA The flow of information in cells DNA→ RNA→ PROTEINS REPLICATION Two steps in making proteins: Transcription Translation Remember RNA structure? THREE KINDS OF RNA Messenger RNA (mRNA)—carries copy of a DNA sequence to site of protein synthesis at the ribosome Transfer RNA (tRNA)—carries amino acids for polypeptide assembly Ribosomal RNA (rRNA)—catalyzes peptide bonds and provides structure FROM GENE TO PROTEIN Exception to the central dogma: retroviruses transcription We need: DNA template Nucleoside tri-phosphates (ATP, CTP, GTP, UTP) RNA polymerase RNA POLYMERASE TRANSCRIPTION OCCURS IN 3 PHASES INTITIATION ELONGATION TERMINATION INITIATION ELONGATION & TERMINATION THE GENETIC CODE Codon: A sequence of three bases—each codon specifies a particular amino acid. Start codon: AUG—initiation signal for translation. Stop codons: UAA, UAG, UGA—stop translation and polypeptide is released. How was the code deciphered? 20 “code words” (amino acids) are written with only four “letters.” Triplet code seemed likely: Could account for 4 × 4 × 4 = 64 codons. DECIPHERING THE GENETIC CODE THE GENETIC CODE Is REDUNDANT Not AMBIGUOUS Is UNIVERSAL Codons call for the same amino acids in all species Exception: Chloroplast and Mitochondrion DNA Genetic Code is the common language for evolution Has remained the same Facilitates genetic engineering DIFFERENCES BETWEEN PROKARYOTES & EUKARYOTES EUKARYOTIC GENES Promoters and terminators Non-coding genes: Introns Coding sequences: Exons Introns and exons appear in the primary mRNA transcript—pre-mRNA; introns are removed from the final mRNA EUKARYOTIC GENES HOW INTRONS WERE DISCOVERED in Processing mRNA In the nucleus, pre-mRNA is modified at both ends: G cap is added at the 5′ end (modified guanosine triphosphate)—facilitates mRNA binding to ribosome. G cap protects mRNA from being digested by ribonucleases. Poly A tail added at 3′ end. AAUAAA sequence after last codon is a signal for an enzyme to cut the pre-mRNA; then another enzyme adds 100 to 300 adenines—the “tail.” Adding the g-cap The snrnps (small nuclear ribonucleic particles) & SPLICEOSOME In the disease β-thalassemia, a mutation may occur at an intron consensus sequence in the β-globin gene—the pre-mRNA can not be spliced correctly. Non-functional β-globin mRNA is produced. Messenger rna leaving the nucleus Through nuclear pores Led by TAP proteins attaching to 5’ end Unused mRNA stays in nucleus TRANSLATION Transfer RNA links information to the codons on m-RNA Each t-RNA brings the specific amino acid Transfer RNA must read codons correctly Transfer RNA must bring the correct amino acid to m-RNA Three functions of tRNA: It binds to an amino acid, and is then “charged” It associates with mRNA molecules It interacts with ribosomes Transfer rna 3′ end is the amino acid attachment site—binds covalently. Anticodon: At the midpoint of the tRNA sequence—site of base pairing with mRNA. Unique for each species of tRNA. Wobble: Specificity for the base at the 3′ end of the codon is not always observed. Example: Codons for alanine—GCA, GCC, and GCU—are recognized by the same tRNA. Wobble allows cells to produce fewer tRNA species, but does not allow the genetic code to be ambiguous CHARGING THE TRANSFER RNA MOLECULE RIBOSOMES Ribosomes have two subunits, large and small. In eukaryotes, the large subunit has three molecules of ribosomal RNA (rRNA) and 49 different proteins in a precise pattern. The small subunit has one rRNA and 33 proteins RIBOSOMES PARTS OF A RIBOSOME A (amino acid) site binds with anticodon of charged tRNA P (polypeptide) site is where tRNA adds its amino acid to the growing chain E (exit) site is where tRNA sits before being released from the ribosome Like transcription, translation also occurs in three steps: Initiation Elongation Termination INITIATION An initiation complex forms—a charged tRNA and small ribosomal subunit, both bound to mRNA. In prokaryotes rRNA binds to mRNA recognition site “upstream” from start codon. In eukaryotes the small subunit binds to the 5′ cap on the mRNA and moves until it reaches the start codon. INITIATION (start up codon on mRNA is AUG) Elongation The second charged tRNA enters the A site. Large subunit catalyzes two reactions: It breaks bond between tRNA in P site and its amino acid Peptide bond forms between that amino acid and the amino acid on tRNA in the A site ELONGATION Termination Translation ends when a stop codon enters the A site. Stop codon binds a protein release factor—allows hydrolysis of bond between polypeptide chain and tRNA on the P site. Polypeptide chain separates from the ribosome—C terminus is the last amino acid added. TERNMINATION POLYRIBOSOME OR POLYSOME POSTRANSLATIONAL ASPECTS Protein modifications Proteolysis: Cutting of a long polypeptide chain into final products, by proteases Glycosylation: Addition of sugars to form glycoproteins Phosphorylation: Addition of phosphate groups catalyzed by protein kinases— charged phosphate groups change the conformation POSTRANSLATIONAL MODIFICATIONS TO PROTEINS