* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Restriction Enzymes
Epigenetics in learning and memory wikipedia , lookup
DNA barcoding wikipedia , lookup
Zinc finger nuclease wikipedia , lookup
Human genome wikipedia , lookup
DNA profiling wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Cancer epigenetics wikipedia , lookup
DNA polymerase wikipedia , lookup
Primary transcript wikipedia , lookup
DNA damage theory of aging wikipedia , lookup
Point mutation wikipedia , lookup
United Kingdom National DNA Database wikipedia , lookup
SNP genotyping wikipedia , lookup
Genealogical DNA test wikipedia , lookup
Metagenomics wikipedia , lookup
Gel electrophoresis of nucleic acids wikipedia , lookup
Designer baby wikipedia , lookup
DNA vaccination wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Nucleic acid analogue wikipedia , lookup
Nucleic acid double helix wikipedia , lookup
Non-coding DNA wikipedia , lookup
Bisulfite sequencing wikipedia , lookup
DNA supercoil wikipedia , lookup
Epigenomics wikipedia , lookup
Genomic library wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
Molecular cloning wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Microsatellite wikipedia , lookup
Genetic engineering wikipedia , lookup
Microevolution wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
Genome editing wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Helitron (biology) wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
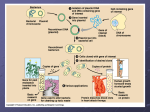
Today: Biotechnology Over 600 recent transposon insertions were identified by examining DNA from 36 genetically diverse humans. Tbl 1 Which transposable elements are active in the human genome? (2007) Ryan E. Mills et al. Trends in Genetics 23: 183-191 DNA fingerprinting using RFLPs Visualizing differences in DNA sequence by using restriction enzymes Sequence 1 Sequence 2 Fig 18.1 Restriction Enzymes cut DNA at specific sequences tbl 18.3 Examples of some restriction enzymes… Recognition Enzyme Sequence EcoRI 5'GAATT C 3'CTTAAG BamHI 5'GGAT CC 3'CCTAGG HindIII 5'AAGCTT 3'TTCGAA TaqI 5'TCGA 3'AGCT AluI 5'AGCT 3'TCGA Cut 5'---G AATT C---3' 3'---CTTAA G--- 5' 5'---G GAT CC---3' 3'---CCTAG G---5' 5'---A AG CTT---3' 3'---TTCGA A--- 5' 5'---T CGA---3' 3'---AGC T---5' 5'---AG CT---3' 3'---TC GA---5' Fig 20.5+.6 Visualizing differences in DNA sequence by using restriction enzymes Sequence 1 Sequence 2 Fig 20.6 Separating DNA on a gel by size • Gel electrophoresis Fig 24.21 The different sized bands can arise from different cut sites and/or different number of nucleotides between the cut sites. Sequence 1 Sequence 2 Sequence 1 Sequence 2 Fig 22.23 DNA fingerprinting DNA fingerprinting DNA fingerprinting Can DNA be obtained from hair? How can DNA be obtained from such a small sample? The inventor of PCR Fig 18.6 Polymerase Chain Reaction: amplifying DNA Polymerase Chain Reaction Fig 18.6 Fig 18.6 Polymerase Chain Reaction: Primers allow specific regions to be amplified. The inventor of PCR PCR animation http://www.dnalc.org/ddnalc/resources/pcr.html Areas of DNA from very small samples can be amplified by PCR, and then cut with restriction enzymes for RFLP analysis. Genetic Engineering: Direct manipulation of DNA Fig 18.2 Bacteria can be modified or serve as intermediates Fig 18.2 a typical bacteria Bacterial DNA plasmid DNA tbl 18.2 A typical bacterial plasmid used for genetic engineering Fig 18.2 Moving a gene into bacteria via a plasmid What problems exist for expressing eukaryotic gene in bacteria? Bacterial DNA plasmid DNA Fig 18.4 Reverse transcriptase can be used to obtain coding regions without introns. Fig 18.6 After RT, PCR will amplify the gene or DNA Fig 18.2 Moving a gene into bacteria via a plasmid RT and PCR Fig 18.1 Restriction Enzymes cut DNA at specific sequences Fig 18.1 Restriction enzymes cut DNA at a specific sequence Fig 18.1 Cutting the plasmid and insert with the same restriction enzyme makes matching sticky ends A typical bacterial plasmid used for genetic engineering Using sticky ends to add DNA to a bacterial plasmid Fig 18.1 tbl 6.1 Transformation of bacteria can happen via several different methods. Bacteria can take up DNA from the environment Fig 9.2 Transformation of bacteria can happen via several different methods all involving perturbing the bacterial membrane: Tbl 6.1 •Electroporation •Heat shock •Osmotic Stress Fig 18.1 How can you know which bacteria have been transformed, and whether they have the insert? Resistance genes allow bacteria with the plasmid to be selected. Bacteria with the resistance gene will survive when grown in the presence of antibiotic Fig 18.1 Fig 20.5 Is the insert present? Plasmids with the MCS in the lacZ gene can be used for blue/white screening… A typical bacterial plasmid used for genetic engineering Intact lacZ makes a blue color when expressed and provided X-galactose When the lacZ gene is disrupted, the bacteria appear white Fig 18.1 Blue/white screening: Transformed bacteria plated on antibiotic and Xgal plates. Each colony represents millions of clones of one transformed cell. Fig 18.1 Successful transformation will grow a colony of genetically modified bacteria RT and/or PCR Fig 18.1 Inserting a gene into a bacterial plasmid Millions of Hectares Bacteria can be used to transform plants Global area planted with GM crops http://www.gmo-compass.org/eng/agri_biotechnology/gmo_planting/257.global_gm_planting_2006.html Texas = 70 ha