* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Document
Genealogical DNA test wikipedia , lookup
Genomic imprinting wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Y chromosome wikipedia , lookup
Transposable element wikipedia , lookup
Oncogenomics wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Epigenomics wikipedia , lookup
Point mutation wikipedia , lookup
Neocentromere wikipedia , lookup
Cell-free fetal DNA wikipedia , lookup
DNA vaccination wikipedia , lookup
DNA supercoil wikipedia , lookup
Molecular cloning wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Human genome wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Non-coding DNA wikipedia , lookup
X-inactivation wikipedia , lookup
Designer baby wikipedia , lookup
Genetic engineering wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Genome (book) wikipedia , lookup
Helitron (biology) wikipedia , lookup
Minimal genome wikipedia , lookup
Genome evolution wikipedia , lookup
Genomic library wikipedia , lookup
Genome editing wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Microevolution wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
No-SCAR (Scarless Cas9 Assisted Recombineering) Genome Editing wikipedia , lookup
Cre-Lox recombination wikipedia , lookup
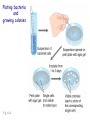
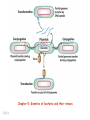
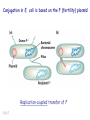
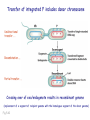
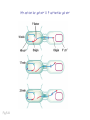
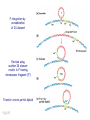
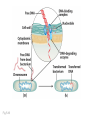
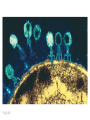
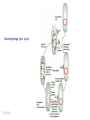
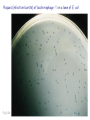
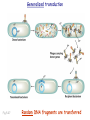
Plating bacteria and growing colonies Fig. 5-2 Commonly used genetic markers • Prototrophic markers: wild-type bacteria are prototrophs (grow on minimal medium) • Auxotrophic markers: mutants that require additional nutrient (fail to grow on minimal medium) • Antibiotic-sensitivity: wild-type bacteria are susceptible (fail to grow on antibiotic-containing medium) • Antibiotic-resistance: mutants that grow in presence of antibiotic (grow on antibiotic-containing medium) Chapter 5: Genetics of bacteria and their viruses Fig. 5-1 Gene transfer mechanisms in bacteria (especially E. coli) Conjugation: orderly, deliberate transfer of DNA from one cell to another; programmed by specialized genes and organelles. Transformation: uptake of environmental DNA into a cell Transduction: transfer of DNA from one cell to another mediated by a virus Properties of gene transfer in bacteria • All are unidirectional (donor – recipient) • Recombination requires two steps: 1. Transfer of DNA into the recipient cell, forming a merozygote (various gene transfer mechanisms) 2. Crossing over that replaces a portion of the recipient genome (endogenote) with the homologous portion of the donor genome (exogenote) • Transfer is always partial Conjugating E. coli pili Fig. 5-6 Conjugation in E. coli is based on the F (fertility) plasmid Replication-coupled transfer of F Fig. 5-7 F can integrate into the bacterial chromosome Hfr: high frequency recombination derivative Fig. 5-8 Transfer of integrated F includes donor chromosome Unidirectional transfer…… Recombination….. Partial transfer….. Crossing over of exo/endogenote results in recombinant genome (replacement of a segment of recipient genome with the homologous segment of the donor genome) Fig. 5-10 DNA transfer during conjugation is time-dependent Transfer of an entire E. coli donor genome requires about 1 hour (F sequence is last to transfer) Therefore, can map the chromosome as a time function: • Mix donor Hfr and recipient F- cells • Interrupt transfer of DNA at various times (violent mixing in a Waring blendor works!) • Plate out cells to determine which genes were transferred within each timeframe Hfr azir tonr lac+ gal+ strs X F- azis tons lac- gal- strr Fig. 5-11 Hfr azir tonr lac+ gal+ strs X F- azis tons lac- gal- strr Fig. 5-11 Genetic map generated by interrupted mating experiment Conjugation map depends upon: • site of F factor insertion within Hfr chromosome (original F insertion can occur at any one of many sites within chromosome) • direction/orientation of the F factor within that Hfr strain (clockwise or counter-clockwise) Mapping using different Hfr strains can provide a map of the entire bacterial chromosome Fig. 5-13 Mapping of small regions by recombination Fig. 5-16 F integration by recombination of IS element Excision using another IS element results in F bearing chromosome fragment (F’) Transfer create partial diploid Fig. 5-17 …at least 10 species ancestors. Fig. 5-18 Transformation: DNA in the environment of a cell is taken into the recipient cell forming a merozygote; then recombination occurs • occurs naturally in some bacteria (e.g., Pneumococcus) • occurs rarely in others, but can be promoted by treating cells to destabilize their membranes (e.g., in recombinant DNA work) • can map genes by co-transformation (frequency with which two genes are simultaneously transferred Fig. 5-19 Transduction: Transfer of DNA from one cell to another mediated by a virus; followed by recombination to integrate the DNA into the recipient cell • can map genes by the frequency of co-transduction (frequency of simultaneous transfer of two genes) Fig. 5-22 Bacteriophage lytic cycle Fig. 5-23 Plaques (infection bursts) of bacteriophage on a lawn of E. coli Fig. 5-24 Generalized transduction Fig. 5-27 Random DNA fragments are transferred Linkage mapping of a segment of the E. coli chromosome by co-transduction experiments with phage P1 Fig. 5-28 Lysogenic infection: integration of a viral genome into one of many sites within the host cell chromosome where it quiescently resides Fig. 5-30 Upon specific cues, the process may be reversed, resulting in lytic infection Specialized transduction (genes nearest the insertion site are most efficiently transferred) Fig. 5-31 Fig. 5- Fig. 5- Fig. 5- Fig. 5-