* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download Gene Regulation Notes
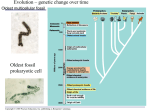
Genome evolution wikipedia , lookup
Epigenetics of neurodegenerative diseases wikipedia , lookup
Gene therapy of the human retina wikipedia , lookup
Gene nomenclature wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Gene expression programming wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Polyadenylation wikipedia , lookup
History of genetic engineering wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
History of RNA biology wikipedia , lookup
Point mutation wikipedia , lookup
RNA interference wikipedia , lookup
Gene expression profiling wikipedia , lookup
Messenger RNA wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Non-coding DNA wikipedia , lookup
Microevolution wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Designer baby wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Transcription factor wikipedia , lookup
Mir-92 microRNA precursor family wikipedia , lookup
RNA silencing wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Epitranscriptome wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Non-coding RNA wikipedia , lookup
Chapter 18 Regulation of Gene Expression Overview: Conducting the Genetic Orchestra • Prokaryotes and eukaryotes alter gene expression in response to their changing environment • Responding through conservation of energy and resources favors these bacteria in terms of natural selection • In other words: Waste not . . . Want not! Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: Bacteria respond to environmental change • One method employed by bacteria we have already discussed - feedback inhibition • Feedback inhibition can control biosynthetic pathways. • The enzyme product inhibits the enzyme activity, turning off production of the product Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: Bacteria respond to environmental change by regulation transcription • A different method is to directly regulate transcription. • Prokaryotic DNA is often under “coordinate control” – meaning multiple polypeptides are made from the same stretch of DNA (makes it easy to turn all the genes on or off). • Gene regulation is how the switch is turned on or off – in prokaryotes the regulatory parts are called an operon. Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: What is an operon? • An operon consists of: – Operator – a segment of DNA within the promotor region that acts as the switch for transcription – Promotor – a segment of DNA where RNA Polymerase II binds and starts transcription – Genes – DNA that codes for polypeptides that form enzymes or other cell products (trp operon – codes for enzymes which make tryptophan) Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: The trp operon Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: How to turn off the trp operon • The operon can be switched off by a protein repressor (ex. trpR repressor) • The repressor prevents gene transcription by binding to the operator and blocking RNA polymerase Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: Where do repressor proteins come from? • The repressor is the product of a separate regulatory gene on the DNA Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: How is the repressor made active? • The repressor is made active by binding with a corepressor (in this case an actual tryptophan molecule). Result = no RNA made. Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: How is repressor made inactive? • The repressor is inactive when no corepressor (tryptophan) is available to bind with it. Result = RNA is made. • trp operon animation Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: Follow up questions 1. In terms of the trp operon what would happen if tryptophan were present at high levels in the cell? 2. Would the trpR be active or inactive? 3. What part of the operon is actually blocked by the repressor? 4. What is the repressor actually blocking? 5. Where does the repressor protein actually come from? 6. Why do cells seek to regulate gene expression? Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Fig. 18-3 trp operon Promoter Promoter Genes of operon DNA trpR Regulatory gene mRNA 5 Protein trpE 3 Operator Start codon mRNA 5 RNA polymerase Inactive repressor E trpD trpB trpA B A Stop codon D C Polypeptide subunits that make up enzymes for tryptophan synthesis (a) Tryptophan absent, repressor inactive, operon on DNA No RNA made mRNA Active repressor Protein trpC Tryptophan (corepressor) (b) Tryptophan present, repressor active, operon off 18.1 Repressible and Inducible Operons: Two Types of Negative Gene Regulation • Repressible operon - one that is usually on; binding of a repressor to the operator shuts off transcription (ex. trp operon) • Inducible operon - one that is usually off; a molecule called an inducer inactivates the repressor and turns on transcription (ex. lac operon) Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1 Repressible and Inducible Operons: Two Types of Negative Gene Regulation Inducible Repressible (Corepressor) (Inducer) Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1 Lac operon - Inducible • The lac operon contains genes that code for enzymes used in the hydrolysis of lactose • The lac repressor is active and keeps the lac operon off (blocks RNA polymerase II) • The inducer (lactose) inactivates the repressor to turn the lac operon on – the repressor leaves the operator • lac operon animation Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Fig. 18-4a Regulatory gene Promoter Operator lacI DNA lacZ No RNA made 3 mRNA 5 Protein RNA polymerase Active repressor (a) Lactose absent, repressor active, operon off Fig. 18-4b lac operon DNA lacZ lacY -Galactosidase Permease lacI 3 mRNA 5 RNA polymerase mRNA 5 Protein Allolactose (inducer) lacA Inactive repressor (b) Lactose present, repressor inactive, operon on Transacetylase 18.1 Inducible vs. Repressible Enzymes and Negative Control of Genes • Inducible enzymes usually function in catabolic pathways; their synthesis is induced by a chemical signal • Repressible enzymes usually function in anabolic pathways; their synthesis is repressed by high levels of the end product • Regulation of the trp and lac operons involves negative control of genes because operons are switched off by the active form of the repressor Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Positive Gene Regulation • Catabolite activator protein (CAP), an activator of transcription • When glucose (a preferred food source of E. coli) is scarce, CAP is activated by binding with cyclic AMP (adenosine monophosphate) Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Positive Gene Regulation • Activated CAP attaches to the promoter of the lac operon and increases the affinity of RNA polymerase, thus accelerating transcription • CAP positive regulation animation Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Follow up questions • What is the difference between repressible and inducible enzymes? • What is the difference between negative and positive gene regulation? Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Concept 18.2: Eukaryotic gene expression can be regulated at any stage • Points at which gene expression can be regulated: chromatin mod., transcription, RNA processing, transport to cytoplasm, translation, protein processing, transport to cell destination • In multicellular organisms gene expression is essential for cell specialization • Although gene expression is regulated at many stages, we will focus on eukaryotic gene regulation at the stage of transcription. Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Differential Gene Expression • Almost all the cells in an organism are genetically identical • Differences between cell types result from differential gene expression, the expression of different genes by cells with the same genome Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.2 Regulation of Transcription Initiation • Chromatin-modifying enzymes provide initial control of gene expression by making a region of DNA either more or less able to bind the transcription machinery (particularly RNA polymerase) Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.2 Organization of a Typical Eukaryotic Gene • Associated with most eukaryotic genes are control elements, segments of noncoding DNA that help regulate transcription by binding certain proteins • Control elements can be proximal (close) to the gene, or distal (far from the gene, thousands of bases away) Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.2 Organization of a Typical Eukaryotic Gene • Other parts of the gene include the promoter, coding sequence (introns and exons), the poly A signal sequence and the termination region Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.2 The Roles of Transcription Factors • Eukaryotic RNA polymerase requires the assistance of proteins called transcription factors • General transcription factors are essential for the transcription of all protein-coding genes Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.2 The Roles of Transcription Factors • An activator is a protein that binds to an enhancer (at the distal control element) and stimulates transcription of a gene • Bound activators interact with mediator proteins and transcription factors at the promoter Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Promoter Activators DNA Enhancer Distal control element Gene TATA box General transcription factors DNA-bending protein Group of mediator proteins 1. Enhancers are distal control elements 2. Activators bind to enhancers 3. Activators on enhancer will interact with transcription factors and mediator proteins Fig. 18-9-3 Promoter Activators DNA Enhancer Distal control element 4. The activators are brought into contact with the proteins by a DNA bending protein 5. The enhancer, activator, transcription factors, mediator proteins, promoter and polymerase all form a transcription initiation complex. Gene TATA box General transcription factors DNA-bending protein Group of mediator proteins RNA polymerase II RNA polymerase II Transcription initiation complex RNA synthesis 18.2 The Roles of Transcription Factors • Some transcription factors function as repressors, inhibiting expression of a particular gene • Some activators and repressors act indirectly by influencing chromatin structure to promote or silence transcription Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.2 Mechanisms of Post-Transcriptional Regulation • Transcription alone does not account for gene expression • Regulatory mechanisms can operate at various stages after transcription to fine-tune gene expression: – Alternative RNA splicing – mRNA degradation – Initiation of translation – Protein processing and degradation Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Concept 18.3: Noncoding RNAs play multiple roles in controlling gene expression • Only a small fraction of DNA codes for proteins, rRNA, and tRNA • A significant amount of the genome may be transcribed into noncoding RNAs • Noncoding RNAs regulate gene expression at two points: mRNA translation and chromatin configuration Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.3 Effects on mRNAs by MicroRNAs and Small Interfering RNAs • MicroRNAs (miRNAs) are small singlestranded RNA molecules that can bind to mRNA • These miRNAs can degrade mRNA or block its translation Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings Fig. 18-13 Hairpin miRNA Hydrogen bond Dicer miRNA 5 3 (a) Primary miRNA transcript mRNA degraded miRNAprotein complex Translation blocked (b) Generation and function of miRNAs 18.3 RNA interference (RNAi) and Small Interfering RNAs (siRNAs) • The phenomenon of inhibition of gene expression by RNA molecules is called RNA interference (RNAi) • RNAi is caused by small interfering RNAs (siRNAs) • siRNAs and miRNAs are similar but form from different RNA precursors Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.3 Chromatin Remodeling and Silencing of Transcription by Small RNAs • siRNAs play a role in heterochromatin formation and can block large regions of the chromosome • siRNAs that convert euchromatin to heterochromatin block transcription of specific genes Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: The trp operon Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: The trp operon – active repressor Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: The lac operon – active repressor Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings 18.1: The lac operon – inactive repressor Copyright © 2008 Pearson Education Inc., publishing as Pearson Benjamin Cummings