* Your assessment is very important for improving the work of artificial intelligence, which forms the content of this project
Download this poster
History of RNA biology wikipedia , lookup
Gene expression programming wikipedia , lookup
Genetic engineering wikipedia , lookup
X-inactivation wikipedia , lookup
Metagenomics wikipedia , lookup
Extrachromosomal DNA wikipedia , lookup
Mitochondrial DNA wikipedia , lookup
Deoxyribozyme wikipedia , lookup
Ridge (biology) wikipedia , lookup
Epigenetics of diabetes Type 2 wikipedia , lookup
Non-coding RNA wikipedia , lookup
Epigenomics wikipedia , lookup
Vectors in gene therapy wikipedia , lookup
Genome (book) wikipedia , lookup
Cancer epigenetics wikipedia , lookup
Short interspersed nuclear elements (SINEs) wikipedia , lookup
Pathogenomics wikipedia , lookup
Epigenetics in learning and memory wikipedia , lookup
Genomic imprinting wikipedia , lookup
Long non-coding RNA wikipedia , lookup
Genomic library wikipedia , lookup
Designer baby wikipedia , lookup
Site-specific recombinase technology wikipedia , lookup
Human genome wikipedia , lookup
RNA interference wikipedia , lookup
Gene expression profiling wikipedia , lookup
Primary transcript wikipedia , lookup
Non-coding DNA wikipedia , lookup
Genome editing wikipedia , lookup
Artificial gene synthesis wikipedia , lookup
Polycomb Group Proteins and Cancer wikipedia , lookup
Therapeutic gene modulation wikipedia , lookup
Nutriepigenomics wikipedia , lookup
Epigenetics of human development wikipedia , lookup
Transposable element wikipedia , lookup
Microevolution wikipedia , lookup
History of genetic engineering wikipedia , lookup
Minimal genome wikipedia , lookup
Genome evolution wikipedia , lookup
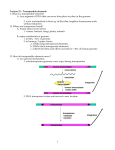
Reproduction Specific Argonaute Genes In Maize and Barley And Their Role In Transposon Silencing Manjit 1 Singh , Daniel 2 Grimanelli and Jaswinder 1 Singh 1Department of Plant Science, McGill University, Ste-Anne de Bellevue, QC Canada H9X 3V9 2Institut de recherché pour le developpement UMR232 Universite de Montpellier, Montpellier, France Abstract RNA-mediated silencing of transposons is an important defence mechanism to suppress the proliferation of transposons in plants and animals. In plants such processes for transposon silencing have been suggested to act in both the female and male gametophytes. Argonaute proteins are key players in RNA dependent silencing mechanism and we are interested in investigating the role of female gametophyte-specific ARGONAUTEs in transposon silencing. Previously a female reproductive tissue-specific ARGONAUTE, AGO104, has been identified in maize. Transcriptional profiling of ovaries from ago104 mutants showed an abundance of transcripts from transposons and repeats compared to the wild type plants suggesting a female gametophytic mechanism for transposon silencing in maize. We are further studying the role of AGO4-like proteins in a large genome cereal, barley, a true diploid grass species with a genome twice the size of maize. Barley has two Ago4-like genes Ago1002 and Ago1003, of which Ago1002 shows a higher homology to Ago104. The comparative expression data of the barley Ago4-like gene will be presented. Mutations in the Ago1002 and Ago1003 genes are also being identified using a TILLING population. A comparative analysis of components of RNA-mediated silencing mechanisms may contribute our understanding of genome expansion and larger genomes in certain species. Maize AGO104 shows reproductive tissue-specific localization •AGO104 protein accumulates during both female and male sporogenesis but not during gametogenesis (when transcription is highest), as weak or no signal was detected. •The gene is broadly transcribed but the AGO104 protein specifically accumulates in the ovaries and the anthers around the time of meiosis, possibly because of posttranscriptional regulation. Ago1002 and Ago1003 are preferentially expressed in reproductive tissue Preliminary gene expression studies show that both Ago1002 and Ago1003 are preferentially expressed in the ovary and anthers compared to leaf tissue. Ago1003 is more strongly expressed in these tissues compared to Ago1002 AGO104 role in transposon silencing Heterochromatin silencing pathway through DNA Methylation Argonautes •AGOs and related PIWI proteins form a large family found in plants, animals, and fungi. They act in transcriptional gene silencing, posttranscriptional gene silencing, or through modification of chromatin by RNA interference. •AGOs are 100-kD proteins that contain characteristic PAZ and PIWI domains. The PAZ domain binds the small RNA, while the PIWI domain is purported to have RNaseH-like endonuclease activity, which is required to cleave bound RNA targets for AGOs that have a slicer function. • Arabidopsis AGO4, AGO5 and AGO9 belong to the AGO4_9 class of Argonaute proteins and are involved in small RNA mediated DNA silencing. AGO4 and possibly AGO9 act via RNA-directed DNA Methylation (RdDM) to silence transposons and heterochromatic regions. Barley genome with a size of 5.3 Mbp is one of the largest in cereal crops and twice the size of the maize genome. Barley is a true diploid and is therefore a model species for genetics and genomics of other important species of the Triticeae tribe including wheat. Survey of the extensive transcriptome data suggests that barley has two Argonaute genes that belong to the Ago4_9 class; Ago1002 and Ago1003. Phylogenetic relationship between maize and barley AGO4_9 class proteins Immunolocalization of AGO104 protein in ovary tissue From Henderson and Jacobson Nature 447, 418-424 Ago4_9 class Argonaute genes in Barley 2 •Transcript profiling of ago104 mutant ovules showed increased transcription of repetitive DNA as compared to wild type ovules. •Increased expression of both the retro- and DNA transposons was observed in ago104 mutant ovules with the strongest effect on DNA transposons. •AGO104 localization pattern along with increased transcripts of transposons in the ovules of mutant suggests a reproductive tissue specific mechanism for transposon and repeat silencing. Species Maize Barley Bread wheat Chromosome number 2n = 2x = 20 2n = 2x = 14 (HH) 2n = 6x = 42 (AABBDD) Genome size of maize and two major Triticeae species 2.5 1.5 Ago1002 Ago1003 1 0.5 0 Ovary Meiosis Ovary Mature Ovary 3DAP Anthers Mixed Leaf Relative transcript abundance of Ago1002 and Ago1003 genes in reproductive tissues Conclusions and future research Monoploid genome size 2,500 Mbp 5,100 Mbp 5,646 Mbp •Ago4_9 Class of genes Ago1002 and Ago1003 are expressed in the reproductive tissues in barley. •Expression pattern suggests that Ago1002 and Ago1003 may have reproduction related function and either of them can be an orthologue of Ago104. •TILLING mutants are being identified to perform functional characterization of Ago1002 and Ago1003 to study their role in reproductive development and transposon silencing in barley and related large genome cereals.